Abstract
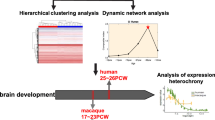
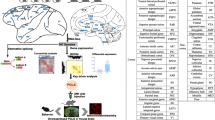
As the key organ that separates humans from nonhuman primates, the brain has continuously evolved to adapt to environmental and climatic changes. Although humans share most genetic, molecular, and cellular features with other primates such as macaques, there are significant differences in the structure and function of the brain between humans and these species. Thus, exploring the differences between the brains of human and nonhuman primates in the context of evolution will provide insights into the development, functionality, and diseases of the human central nervous system (CNS). Since the genes involved in many aspects of the human brain are under common pressures of natural selection, their evolutionary features can be analyzed collectively at the pathway level. In this study, the molecular mechanisms underlying human brain capabilities were explored by comparing the evolution features of pathways enriched in genes expressed in the human brain and the macaque brain. We identified 31 pathways with differential evolutionary properties, including those related to neurological diseases, signal transduction, immunological response, and metabolic processes. By analyzing genes differentially expressed in brain regions or development stages between humans and macaques, 9 and 4 pathways with differential evolutionary properties were detected, respectively. We further performed crosstalk analysis on the pathways to obtain an intuitive correlation between the pathways, which is helpful in understanding the mechanisms of interaction between pathways. Our results provide on a comprehensive view of the evolutionary pathways of the human CNS and can serve as a reference for the study of human brain development.



Similar content being viewed by others
Availability of Data and Material
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Abe C, Yamaoka Y et al (2019) Hypergravity-induced plastic alteration of the vestibulo-sympathetic reflex involves decrease in responsiveness of CAMK2-expressing neurons in the vestibular nuclear complex. J Physiol Sci 69(6):903–917
Adams JB, Johansen LJ et al (2011) Gastrointestinal flora and gastrointestinal status in children with autism–comparisons to typical children and correlation with autism severity. BMC Gastroenterol 11:22
Akiyama H, Barger S et al (2000) Inflammation and Alzheimer’s disease. Neurobiol Aging 21(3):383–421
Ardesch DJ, Scholtens LH et al (2019) The human connectome from an evolutionary perspective. Prog Brain Res 250:129–151
Berg JJ, Coop G (2014) A population genetic signal of polygenic adaptation. PLoS Genet 10(8):e1004412
Bergey CM, Lopez M et al (2018) Polygenic adaptation and convergent evolution on growth and cardiac genetic pathways in African and Asian rainforest hunter-gatherers. Proc Natl Acad Sci U S A 115(48):E11256–E11263
Brandolini L, d'Angelo M et al (2019) Chemokine Signaling in Chemotherapy-Induced Neuropathic Pain. Int J Mol Sci 20(12)
Buck MD, Sowell RT et al (2017) Metabolic Instruction of Immunity. Cell 169(4):570–586
Buckner RL, DiNicola LM (2019) The brain’s default network: updated anatomy, physiology and evolving insights. Nat Rev Neurosci 20(10):593–608
Buse E, Markert UR (2019) The immunology of the macaque placenta: A detailed analysis and critical comparison with the human placenta. Crit Rev Clin Lab Sci 56(2):118–145
Chen YH, Lu CW et al (2017) Revealing the Saline Adaptation Strategies of the Halophilic Bacterium Halomonas beimenensis through High-throughput Omics and Transposon Mutagenesis Approaches. Sci Rep 7(1):13037
Chikina M, Robinson JD et al (2016) Hundreds of Genes Experienced Convergent Shifts in Selective Pressure in Marine Mammals. Mol Biol Evol 33(9):2182–2192
Childs BG, Gluscevic M et al (2017) Senescent cells: an emerging target for diseases of ageing. Nat Rev Drug Discov 16(10):718–735
Daub JT, Dupanloup I et al (2015) Inference of Evolutionary Forces Acting on Human Biological Pathways. Genome Biol Evol 7(6):1546–1558
David MM, Enard D et al (2016) Comorbid Analysis of Genes Associated with Autism Spectrum Disorders Reveals Differential Evolutionary Constraints. PLoS ONE 11(7):e0157937
de Juan D, Pazos F et al (2013) Emerging methods in protein co-evolution. Nat Rev Genet 14(4):249–261
Deverman BE, Patterson PH (2009) Cytokines and CNS development. Neuron 64(1):61–78
Eidem HR, Rinker DC et al (2016) Comparing human and macaque placental transcriptomes to disentangle preterm birth pathology from gestational age effects. Placenta 41:74–82
Galatro TF, Holtman IR et al (2017) Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat Neurosci 20(8):1162–1171
Hensman Moss DJ, Flower MD et al (2017) Huntington’s disease blood and brain show a common gene expression pattern and share an immune signature with Alzheimer’s disease. Sci Rep 7:44849
Hu YS, Xin J et al (2017) Analyzing the genes related to Alzheimer’s disease via a network and pathway-based approach. Alzheimers Res Ther 9(1):29
Kong Y, Liang X et al (2015) High Throughput Sequencing Identifies MicroRNAs Mediating alpha-Synuclein Toxicity by Targeting Neuroactive-Ligand Receptor Interaction Pathway in Early Stage of Drosophila Parkinson’s Disease Model. PLoS ONE 10(9):e0137432
Kurochkin I, Khrameeva E et al (2019) Metabolome signature of autism in the human prefrontal cortex. Commun Biol 2:234
Li M, Santpere G et al (2018) Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362(6420)
Li Y, Li Q et al (2019) Long Noncoding RNA Expression Profile in BV2 Microglial Cells Exposed to Lipopolysaccharide. Biomed Res Int 2019:5387407
Lopez-Otin C, Blasco MA et al (2013) The hallmarks of aging. Cell 153(6):1194–1217
Louveau A, Smirnov I et al (2015) Structural and functional features of central nervous system lymphatic vessels. Nature 523(7560):337–341
Martinez G, Duran-Aniotz C et al (2017) Endoplasmic reticulum proteostasis impairment in aging. Aging Cell 16(4):615–623
Maulik U, Sen S et al (2018) Detecting TF-miRNA-gene network based modules for 5hmC and 5mC brain samples: a intra- and inter-species case-study between human and rhesus. BMC Genet 19(1):9
Muchnik SK, Lorente-Galdos B et al (2019) Modeling the Evolution of Human Brain Development Using Organoids. Cell 179(6):1250–1253
O’Neill LA, Pearce EJ (2016) Immunometabolism governs dendritic cell and macrophage function. J Exp Med 213(1):15–23
Palomero-Gallagher N, Zilles K (2019) Differences in cytoarchitecture of Broca’s region between human, ape and macaque brains. Cortex 118:132–153
Pattabiraman K, Muchnik SK et al (2020) The evolution of the human brain and disease susceptibility. Curr Opin Genet Dev 65:91–97
Pearce EL, Poffenberger MC et al (2013) Fueling immunity: insights into metabolism and lymphocyte function. Science 342(6155):1242454
Petrides M, Cadoret G et al (2005) Orofacial somatomotor responses in the macaque monkey homologue of Broca’s area. Nature 435(7046):1235–1238
Pollen AA, Bhaduri A et al (2019) Establishing Cerebral Organoids as Models of Human-Specific Brain Evolution. Cell 176(4):743-756e717
Preuss TM, Caceres M et al (2004) Human brain evolution: insights from microarrays. Nat Rev Genet 5(11):850–860
Prokop S, Miller KR et al (2019) Impact of TREM2 risk variants on brain region-specific immune activation and plaque microenvironment in Alzheimer’s disease patient brain samples. Acta Neuropathol 138(4):613–630
Rilling JK (2014) Comparative primate neuroimaging: insights into human brain evolution. Trends Cogn Sci 18(1):46–55
Rilling JK, Glasser MF et al (2008) The evolution of the arcuate fasciculus revealed with comparative DTI. Nat Neurosci 11(4):426–428
Sakai T, Hirata S et al (2012) Fetal brain development in chimpanzees versus humans. Curr Biol 22(18):R791-792
Schafer DP, Stevens B (2015) Microglia Function in Central Nervous System Development and Plasticity. Cold Spring Harb Perspect Biol 7(10):a020545
Shiina T, Blancher A et al (2017) Comparative genomics of the human, macaque and mouse major histocompatibility complex. Immunology 150(2):127–138
Silbereis JC, Pochareddy S et al (2016) The Cellular and Molecular Landscapes of the Developing Human Central Nervous System. Neuron 89(2):248–268
Simon AK, Hollander GA et al (2015) Evolution of the immune system in humans from infancy to old age. Proc Biol Sci 282(1821):20143085
Singh A, Kukreti R et al (2019) Oxidative Stress: A Key Modulator in Neurodegenerative Diseases. Molecules 24(8)
Snyder B, Shell B et al (2017) Chronic intermittent hypoxia induces oxidative stress and inflammation in brain regions associated with early-stage neurodegeneration. Physiol Rep 5(9)
Song H, Li H et al (2018) Targeting Gpr52 lowers mutant HTT levels and rescues Huntington’s disease-associated phenotypes. Brain 141(6):1782–1798
Sousa AMM, Meyer KA et al (2017a) Evolution of the Human Nervous System Function, Structure, and Development. Cell 170(2):226–247
Sousa AMM, Zhu Y et al (2017b) Molecular and cellular reorganization of neural circuits in the human lineage. Science 358(6366):1027–1032
Srivastava M, Simakov O et al (2010) The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466(7307):720–726
Stephenson J, Nutma E et al (2018) Inflammation in CNS neurodegenerative diseases. Immunology 154(2):204–219
Stranger BE, Stahl EA et al (2011) Progress and promise of genome-wide association studies for human complex trait genetics. Genetics 187(2):367–383
Sudmant PH, Rausch T et al (2015) An integrated map of structural variation in 2,504 human genomes. Nature 526(7571):75–81
Tansey MG, Goldberg MS (2010) Neuroinflammation in Parkinson’s disease: its role in neuronal death and implications for therapeutic intervention. Neurobiol Dis 37(3):510–518
Verendeev A, Sherwood CC (2017) Human Brain Evolution. Curr Opin Behav Sci 16:41–45
Wang K, Li M et al (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164
Wei Y, de Lange SC et al (2019) Genetic mapping and evolutionary analysis of human-expanded cognitive networks. Nat Commun 10(1):4839
Wen MC, Chan LL et al (2016) Depression, anxiety, and apathy in Parkinson’s disease: insights from neuroimaging studies. Eur J Neurol 23(6):1001–1019
Winiarska-Mieczan A, Baranowska-Wojcik E et al (2020) The Role of Dietary Antioxidants in the Pathogenesis of Neurodegenerative Diseases and Their Impact on Cerebral Oxidoreductive Balance. Nutrients 12(2)
Xicoy H, Wieringa B et al (2019) The Role of Lipids in Parkinson's Disease. Cells 8(1)
Yin S, Lu K et al (2020) Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain. Nat Commun 11(1):474
Zhou Y, Qiao H et al (2019) Immune and cytokine/chemokine responses of PBMCs in rotavirus-infected rhesus infants and their significance in viral pathogenesis. J Med Virol 91(8):1448–1469
Zhu Y, Sousa AMM et al (2018). Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362(6420)
Acknowledgements
The authors thank Mr. Yonglin Peng and Ms. Meng Yuan for helpful discussions in preparation of the manuscript.
Funding
This work was supported by grants from the National Key Research and Development Program of China (no. 2016YFC0906300), and the National Natural Science Foundation of China (no. 91746205 and 31271411).
Author information
Authors and Affiliations
Contributions
YM, FC and JW designed the experiments. YM, CC, MZ and JW performed experiments and data analysis. YM, FC and JW wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ma, Y., Cao, C., Zhao, M. et al. Evolutionary Changes in Pathways and Networks of Genes Expressed in the Brains of Humans and Macaques. J Mol Neurosci 71, 1825–1837 (2021). https://doi.org/10.1007/s12031-021-01874-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-021-01874-y