Abstract
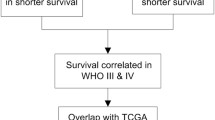
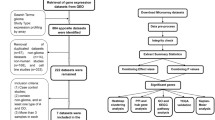
Glioma is one of the most common neurological malignancies worldwide. Delta-like ligand 3 (DLL3), an inhibitory ligand-driven activation of the Notch pathway, has been shown to be significantly associated with overall survival in patients with glioma. Therefore, the purpose of this study was to determine whether DLL3 as a biomarker in glioma is associated with patients' clinicopathological features and prognosis. We identified differences in transcriptome and promoter methylation in the Chinese Glioma Genome Atlas (CGGA) in patients with malignant glioma with shorter (less than 1 year) and longer (greater than 3 years) survival time. Further analysis of The Cancer Genome Atlas (TCGA) revealed that four genes (DLL3, TSPAN15, RTN1, PAK7) are highly associated with patient prognosis and play an indispensable role in evolution. We chose the expression level of DLL3 in glioma patients for our study. Patients were divided into groups with low and high expression of DLL3 according to the cutoff values obtained, and Kaplan–Meier and Cox analysis were used to examine the correlation between DLL3 gene expression and patient survival. We then performed a gene set enrichment analysis (GSEA) to identify significantly enriched signaling pathways. Our results confirmed that the overall survival of patients with low DLL3 expression was significantly shorter than that of patients with high DLL3 expression. GSEA showed that the signaling pathways of the immune process and immune response, among others, were enhanced with the DLL3 low-expression phenotype. Collectively, our findings signify that DLL3 is a potent prognostic factor for glioma, which can provide a viable approach for glioma prognostic assessment and valuable insights for anti-tumor immune-targeted therapies.






Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in the published article.
Abbreviations
- DLL3:
-
Delta-like canonical notch ligand 3
- CGGA:
-
Chinese Glioma Genome Atlas
- TCGA:
-
The Cancer Genome Atlas
- GSEA:
-
Gene set enrichment analysis
- GBM:
-
Glioblastoma
- LGG:
-
Low-grade glioma
- MRI:
-
Magnetic resonance imaging
- CT:
-
Computed tomography
- IDH:
-
Isocitrate dehydrogenase
- 2HG:
-
2-Hydroxyglutarate
- DEGs:
-
Differentially expressed genes
- DMGs:
-
Differentially methylated genes
- SAM:
-
Significance analysis of microarrays
- HR:
-
Hazard ratio
- 95% CI:
-
95% Confidence interval
- FDR:
-
False discovery rate
- ROC:
-
Characteristic curve
References
Adamski V, Schmitt A, Flüh C, Synowitz M, Hattermann K, Held-Feindt J (2017) Isolation and characterization of fast-migrating human glioma cells in the progression of malignant gliomas. Oncol Res 25:341–353. https://doi.org/10.3727/096504016x14737243054982
Akyerli C et al (2018) Use of telomerase promoter mutations to mark specific molecular subsets with reciprocal clinical behavior in IDH mutant and IDH wild-type diffuse gliomas. J Neurosurg 128:1102–1114. https://doi.org/10.3171/2016.11.Jns16973
Borromeo M et al (2016) ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep 16:1259–1272. https://doi.org/10.1016/j.celrep.2016.06.081
Chen X et al (2020) Combined DLL3-targeted bispecific antibody with PD-1 inhibition is efficient to suppress small cell lung cancer growth. J Immunother Cancer 8. https://doi.org/10.1136/jitc-2020-000785
Cooper L et al (2010) The proneural molecular signature is enriched in oligodendrogliomas and predicts improved survival among diffuse gliomas. PloS One 5:e12548. https://doi.org/10.1371/journal.pone.0012548
Dao Trong P, Rösch S, Mairbäurl H, Pusch S, Unterberg A, Herold-Mende C, Warta R (2018) Identification of a Prognostic Hypoxia-Associated Gene Set in IDH-Mutant Glioma. Int J Mol Sci 19. https://doi.org/10.3390/ijms19102903
Darwiche N (2020) Epigenetic mechanisms and the hallmarks of cancer: an intimate affair. Am J Cancer Res 10:1954–1978
Fang S et al (2017) Anatomic location of tumor predicts the accuracy of motor function localization in diffuse lower-grade gliomas involving the hand knob area AJNR. Am J Neuroradiol 38:1990–1997. https://doi.org/10.3174/ajnr.A5342
Furuta M et al (2019) DLL3 regulates the migration and invasion of small cell lung cancer by modulating Snail. Cancer Sci 110:1599–1608. https://doi.org/10.1111/cas.13997
Gentleman R, Ihaka R (2011) R: A Language and Environment for Statistical Computing. Computing 1:12–21
Hendriks L, Menis J, Reck M (2019) Prospects of targeted and immune therapies in SCLC. Expert Rev Anticancer Ther 19:151–167. https://doi.org/10.1080/14737140.2019.1559057
Henke R, Meredith D, Borromeo M, Savage T, Johnson J (2009) Ascl1 and Neurog2 form novel complexes and regulate Delta-like3 (Dll3) expression in the neural tube. Dev Biol 328:529–540. https://doi.org/10.1016/j.ydbio.2009.01.007
Hermans B et al (2019) DLL3 expression in large cell neuroendocrine carcinoma (LCNEC) and association with molecular subtypes and neuroendocrine profile. Lung Cancer (Amsterdam, Netherlands) 138:102–108. https://doi.org/10.1016/j.lungcan.2019.10.010
Hipp S et al (2020) A bispecific DLL3/CD3 IgG-Like T-Cell engaging antibody induces antitumor responses in small cell lung cancer. Clin Cancer Res 26:5258–5268. https://doi.org/10.1158/1078-0432.Ccr-20-0926
Hoyne G, Chapman G, Sontani Y, Pursglove S, Dunwoodie S (2011) A cell autonomous role for the Notch ligand Delta-like 3 in αβ T-cell development. Immunol Cell Biol 89:696–705. https://doi.org/10.1038/icb.2010.154
Hu B et al (2012) Fibulin-3 promotes glioma growth and resistance through a novel paracrine regulation of Notch signaling. Cancer Res 72:3873–3885. https://doi.org/10.1158/0008-5472.Can-12-1060
Hu H et al (2017) Gene Expression and Methylation Analyses Suggest DCTD as a Prognostic Factor in Malignant Glioma. Sci Rep 7:11568. https://doi.org/10.1038/s41598-017-11962-y
Huang J, Cao D, Sha J, Zhu X, Han S (2019) DLL3 is regulated by LIN28B and miR-518d-5p and regulates cell proliferation, migration and chemotherapy response in advanced small cell lung cancer. Biochem Biophys Res Commun 514:853–860. https://doi.org/10.1016/j.bbrc.2019.04.130
Karsy M, Guan J, Cohen A, Jensen R, Colman H (2017) New Molecular Considerations for Glioma: IDH, ATRX, BRAF, TERT, H3 K27M. Curr Neurol Neurosci Rep 17:19. https://doi.org/10.1007/s11910-017-0722-5
Lashari B, Vallatharasu Y, Kolandra L, Hamid M, Uprety D (2018) Rovalpituzumab tesirine: a novel DLL3-targeting antibody-drug conjugate. Drugs R&D 18:255–258. https://doi.org/10.1007/s40268-018-0247-7
Liu Z, Han H, He X, Li S, Wu C, Yu C, Wang S (2016) Expression of the galectin-9-Tim-3 pathway in glioma tissues is associated with the clinical manifestations of glioma. Oncol Lett 11:1829–1834. https://doi.org/10.3892/ol.2016.4142
Louis D et al (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109. https://doi.org/10.1007/s00401-007-0243-4
Louis D et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820. https://doi.org/10.1007/s00401-016-1545-1
Lu W, Xie H, Yuan C, Li J, Li Z, Wu A (2020) Identification of potential biomarkers and candidate small molecule drugs in glioblastoma. Cancer Cell Int 20:419. https://doi.org/10.1186/s12935-020-01515-1
Luo L et al (2020) Blockade of cell volume regulatory protein NKCC1 increases TMZ-induced glioma apoptosis and reduces astrogliosis. Mol Cancer Ther 19:1550–1561. https://doi.org/10.1158/1535-7163.Mct-19-0910
Mellai M, Annovazzi L, Senetta R, Dell'Aglio C, Mazzucco M, Cassoni P, Schiffer D (2017) Diagnostic revision of 206 adult gliomas (including 40 oligoastrocytomas) based on ATRX, IDH1/2 and 1p/19q status. J Neuro-Oncol 131:213–222. https://doi.org/10.1007/s11060-016-2296-5
Narayanappa R, Rout P, Aithal M, Chand A (2016) Aberrant expression of Notch1, HES1, and DTX1 genes in glioblastoma formalin-fixed paraffin-embedded tissues. Tumour Biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 37:6935–6942. https://doi.org/10.1007/s13277-015-4592-7
Ohba S, Hirose Y (2016) Biological Significance of Mutant Isocitrate Dehydrogenase 1 and 2 in Gliomagenesis. Neurol Med Chir 56:170–179. https://doi.org/10.2176/nmc.ra.2015-0322
Phillips H et al (2006) Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9:157–173. https://doi.org/10.1016/j.ccr.2006.02.019
Puca L et al (2019) Delta-like protein 3 expression and therapeutic targeting in neuroendocrine prostate cancer. Sci Transl Med 11. https://doi.org/10.1126/scitranslmed.aav0891
Regzedmaa O et al (2019) Prevalence of DLL3, CTLA-4 and MSTN expression in patients with small cell lung cancer. OncoTargets and Therapy 12:10043–10055. https://doi.org/10.2147/ott.S216362
Sabari J, Lok B, Laird J, Poirier J, Rudin C (2017) Unravelling the biology of SCLC: implications for therapy Nature reviews. Clin Oncol 14:549–561. https://doi.org/10.1038/nrclinonc.2017.71
Saito N et al (2014) A high Notch pathway activation predicts response to γ secretase inhibitors in proneural subtype of glioma tumor-initiating cells. Stem Cells (Dayton, Ohio) 32:301–312. https://doi.org/10.1002/stem.1528
Shahjaman M, Kumar N, Mollah M, Ahmed M, Ara Begum A, Shahinul Islam S, Mollah M (2017) βRobust significance analysis of microarrays by minimum -divergence method. Biomed Res Int 2017:5310198. https://doi.org/10.1155/2017/5310198
Spino M et al (2019) Cell surface notch ligand DLL3 is a therapeutic target in isocitrate dehydrogenase-mutant glioma. Clin Cancer Res 25:1261–1271. https://doi.org/10.1158/1078-0432.Ccr-18-2312
Stiles C, Rowitch D (2008) Glioma stem cells: a midterm exam. Neuron 58:832–846. https://doi.org/10.1016/j.neuron.2008.05.031
Tang F, Liang S, Zhong T, Huang X, Deng X, Zhang Y, Zhou L (2020) Postoperative glioma segmentation in CT image using deep feature fusion model guided by multi-sequence MRIs. Eur Radiol 30:823–832. https://doi.org/10.1007/s00330-019-06441-z
Tateishi K et al (2015) Extreme Vulnerability of IDH1 Mutant Cancers to NAD+ Depletion. Cancer Cell 28:773–784. https://doi.org/10.1016/j.ccell.2015.11.006
Turchi L et al (2013) Tumorigenic potential of miR-18A* in glioma initiating cells requires NOTCH-1 signaling. Stem Cells (Dayton, Ohio) 31:1252–1265. https://doi.org/10.1002/stem.1373
Verckist L, Lembrechts R, Thys S, Pintelon I, Timmermans J, Brouns I, Adriaensen D (2017) Selective gene expression analysis of the neuroepithelial body microenvironment in postnatal lungs with special interest for potential stem cell characteristics. Respir Res 18:87. https://doi.org/10.1186/s12931-017-0571-4
Verhaak R et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17:98–110. https://doi.org/10.1016/j.ccr.2009.12.020
Wu C, Song H, Fu X, Li S, Jiang T (2020) Transcriptomic analysis of glioma based on IDH status identifies ACAA2 as a prognostic factor in lower grade glioma. Biomed Res Int 2020:1086792. https://doi.org/10.1155/2020/1086792
Yan L et al (2019) Prognostic value of delta-like protein 3 combined with thyroid transcription factor-1 in small-cell lung cancer. Oncol Lett 18:2254–2261. https://doi.org/10.3892/ol.2019.10538
Zhao Z, Meng F, Wang W, Wang Z, Zhang C, Jiang T (2017) Comprehensive RNA-seq transcriptomic profiling in the malignant progression of gliomas. Scientific Data 4:170024. https://doi.org/10.1038/sdata.2017.24
Zheng S, Tao W (2020) Identification of novel transcriptome signature as a potential prognostic biomarker for anti-angiogenic therapy in glioblastoma multiforme. Cancers 12. https://doi.org/10.3390/cancers12092368
Funding
This study received no funding.
Author information
Authors and Affiliations
Contributions
All authors contributed substantially to the preparation of this manuscript and approved its final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Maimaiti, A., Wang, X., Hao, Y. et al. Integrated Gene Expression and Methylation Analyses Identify DLL3 as a Biomarker for Prognosis of Malignant Glioma. J Mol Neurosci 71, 1622–1635 (2021). https://doi.org/10.1007/s12031-021-01817-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-021-01817-7