Abstract
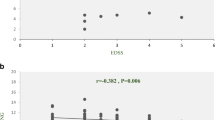
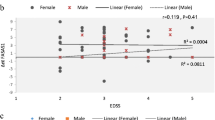
Multiple sclerosis is a disease that affects male and female patients differently. Several studies have been performed to explain the gender differences in MS susceptibility, but the genetic causes underlying gender differences remain unknown. The association between multiple sclerosis and the HLA-DRB1*15:01 haplotype has been confirmed to be female-specific. We hypothesized other immunological components such as lnc-DC may be gender-specific among multiple sclerosis patients, especially when MS patients are negative for the HLA-DRB1*15:01 allele. Therefore, the current study, considering the results of previous studies, aimed to evaluate the expression level of the lnc-DC gene in HLA-DRB1*15:01-negative female patients with relapsing remitting MS (RRMS). A total of 50 MS female patients and 50 female healthy controls were enrolled in this observational case-control study. HLA-DRB1*15:01, as a critical risk factor for MS, was ruled out in all patients. The peripheral blood mononuclear cells were obtained from all patients and total RNA was isolated and cDNA synthesis was carried out. The gene expression of lnc-DC was evaluated by real-time quantitative PCR. Our results have shown that lnc-DC expression level was significantly higher in total MS female patients compared with female controls (P = 0.0044). In addition, the correlation between lnc-DC with disease duration, EDSS, and age at onset did not reach a statistical significance in our study (r = 0.0336, P = 0.817; r = 0.0914, P = 0.5278 and r = 0.0743, P = 0.6083, respectively). Our results give further evidence that lnc-DC may play a gender-dependent role in MS pathogenesis.


Similar content being viewed by others
References
Alonso A, Hernán MA (2008) Temporal trends in the incidence of multiple sclerosis: a systematic review. Neurology 71:129–135
Bahreini SA, Jabalameli MR, Saadatnia M, Zahednasab H (2010) The role of non-HLA single nucleotide polymorphisms in multiple sclerosis susceptibility. J Neuroimmunol 229:5–15
Celius E, Harbo H, Egeland T, Vartdal F, Vandvik B, Spurkiand A (2000) Sex and age at diagnosis are correlated with the HLA-DR2, DQ6 haplotype in multiple sclerosis. J Neurol Sci 178:132–135
Compston A, Coles A (2008) Multiple sclerosis. Lancet (British edition) 372:1502–1517
Dastmalchi R, Ghafouri-Fard S, Omrani MD, Mazdeh M, Sayad A, Taheri M (2018) Dysregulation of long non-coding RNA profile in peripheral blood of multiple sclerosis patients. Multiple Sclerosis Related Dis 25:219–226
Debouverie M, Pittion-Vouyovitch S, Louis S, Guillemin F, Group L. (2008) Natural history of multiple sclerosis in a population-based cohort. Eur J Neurol 15:916–921
Eftekharian MM, Ghannad MS, Taheri M, Roshanaei G, Mazdeh M, Musavi M, Hormoz MB (2016) Frequency of viral infections and environmental factors in multiple sclerosis. Human Antibodies 24:17–23
Eikelenboom M, Killestein J, Kragt J, Uitdehaag B, Polman C (2009) Gender differences in multiple sclerosis: cytokines and vitamin D. J Neurol Sci 286:40–42
Fukazawa T, Yamasaki K, Ito H, Kikuchi S, Minohara M, Horiuchi I, Tsukishima E, Sasaki H, Hamada T, Nishimura Y (2000) Both the HLA-DPB1 and-DRB1 alleles correlate with risk for multiple sclerosis in Japanese: clinical phenotypes and gender as important factors. Tissue Antigens 55:199–205
Gharesouran J, Taheri M, Sayad A, Ghafouri-Fard S, Mazdeh M, Omrani MD (2018) The growth arrest-specific transcript 5 (GAS5) and nuclear receptor subfamily 3 group C member 1 (NR3C1): novel markers involved in multiple sclerosis. Int J Mol Cell Med 7:102
Irizar H, Muñoz-Culla M, Zuriarrain O, Goyenechea E, Castillo-Triviño T, Prada A, Saenz-Cuesta M, De Juan D, Lopez De Munain A, Olascoaga J (2012) HLA-DRB1* 15: 01 and multiple sclerosis: a female association? Mult Scler J 18:569–577
Kakhki MP, Rakhshi N, Aleagha MSE, Abdari M, Alikhah A, Safarian G, Behmanesh M, Nikravesh A (2019) Differential expression of STAT3 gene and its regulatory long non-coding RNAs, namely lnc-DC and THRIL, in two eastern Iranian ethnicities with multiple sclerosis. Neurol Sci 41:561–568.
Mazdeh M, Taheri M, Sayad A, Bahram S, Omrani MD, Movafagh A, Inoko H, Akbari MT, Noroozi R, Hajilooi M (2016) HLA genes as modifiers of response to IFN-β-1a therapy in relapsing-remitting multiple sclerosis. Pharmacogenomics 17:489–498
Münz C, Lünemann JD, Getts MT, Miller SD (2009) Antiviral immune responses: triggers of or triggered by autoimmunity? Nat Rev Immunol 9:246
Polman CH, Reingold SC, Banwell B, Clanet M, Cohen JA, Filippi M, Fujihara K, Havrdova E, Hutchinson M, Kappos L (2011) Diagnostic criteria for multiple sclerosis: 2010 revisions to the McDonald criteria. Ann Neurol 69:292–302
Pozzilli C, Tomassini V, Marinelli F, Paolillo A, Gasperini C, Bastianello S (2003) ‘Gender gap’in multiple sclerosis: magnetic resonance imaging evidence. Eur J Neurol 10:95–97
Quan Z, Zheng D, Qing H (2017) Regulatory roles of long non-coding RNAs in the central nervous system and associated neurodegenerative diseases. Front Cell Neurosci 11:175
Shaker OG, Mahmoud RH, Abdelaleem OO, Ibrahem EG, Mohamed AA, Zaki OM, Abdelghaffar NK, Ahmed TI, Hemeda NF, Ahmed NA (2019) LncRNAs, MALAT1 and lnc-DC as potential biomarkers for multiple sclerosis diagnosis. Biosci Rep 39:BSR20181335
Sigdel KR, Cheng A, Wang Y, Duan L, Zhang Y (2015) The emerging functions of long noncoding RNA in immune cells: autoimmune diseases. J Immunol Res. https://doi.org/10.1155/2015/848790
Spach KM, Hayes CE (2005) Vitamin D3 confers protection from autoimmune encephalomyelitis only in female mice. J Immunol 175:4119–4126
Spielman RS, Bastone LA, Burdick JT, Morley M, Ewens WJ, Cheung VG (2007) Common genetic variants account for differences in gene expression among ethnic groups. Nat Genet 39:226
Stürner KH, Siembab I, Schön G, Stellmann J-P, Heidari N, Fehse B, Heesen C, Eiermann TH, Martin R, Binder TM (2019) Is multiple sclerosis progression associated with the HLA-DR15 haplotype? Mult Scler J Exp Transl Clin 5:2055217319894615
Taheri M, Noroozi R, Sarrafzadeh M, Sayad A 2016 HLA genes as modifiers of response to IFN-β-1a therapy in relapsing-remitting multiple sclerosis. Pharmacogenomics17:489–98
Tomassini V, Onesti E, Mainero C, Giugni E, Paolillo A, Salvetti M, Nicoletti F, Pozzilli C (2005) Sex hormones modulate brain damage in multiple sclerosis: MRI evidence. J Neurol Neurosurg Psychiatry 76:272–275
Valceckiene V, Kontenyte R, Jakubauskas A, Griskevicius L (2010) Selection of reference genes for quantitative polymerase chain reaction studies in purified B cells from B cell chronic lymphocytic leukaemia patients. Br J Haematol 151:232–238
Vandesompele J, de Preter K, Pattyn F, Poppe B, van Roy N, de Paepe A, Speleman F (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:research0034.1
Wang P, Xue Y, Han Y, Lin L, Wu C, Xu S, Jiang Z, Xu J, Liu Q, Cao X (2014) The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 344:310–313
Wang F, Ying H-Q, He B-S, Pan Y-Q, Deng Q-W, Sun H-L, Chen J, Liu X, Wang S-K (2015) Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget 6:7899
Yuan JS, Reed A, Chen F, Stewart CN (2006) Statistical analysis of real-time PCR data. BMC bioinformatics 7:85
Acknowledgments
The current study was supported by grant number (25341) from Shahid Beheshti University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declared no potential conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bahrami, T., Taheri, M., Javadi, S. et al. Expression Analysis of Long Non-coding RNA Lnc-DC in HLA-DRB1*15:01-Negative Patients with Multiple Sclerosis: A Probable Cause for Gender Differences in Multiple Sclerosis Susceptibility?. J Mol Neurosci 71, 821–825 (2021). https://doi.org/10.1007/s12031-020-01704-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-020-01704-7