Abstract
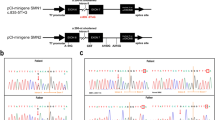
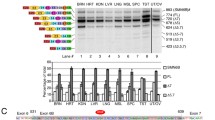
Spinal muscular atrophy (SMA) is an autosomal recessive neuromuscular disorder caused by deletion or subtle variant of survival motor neuron 1 (SMN1) gene. By multiplex ligation-dependent probe amplification, genomic sequencing, and T-A cloning on cDNA level, we identified one novel SMN1 subtle variant c.835G>C (p.Gly279Arg) in a non-homozygous patient with type 1 SMA. Full-length SMN1 (fl-SMN1) transcripts in the peripheral bloods of the patient were significantly decreased compared with those in healthy individuals and the carries (p < 0.05). And two fragments of SMN1 transcripts including fl-SMN1 and △7-SMN1 were observed by RT-PCR, which indicated Exon 7 skipping of SMN1 gene. To further evaluate its splicing effects on Exon 7, we performed ex vivo splicing analysis, which showed that the mutant mini gene with c.835G>C reduced Exon 7 inclusion to 54%. In addition, self-oligomerization between mutant SMN protein with the c.835G>C (p.Gly279Arg) and wild SMN was decreased in self-interaction assays. Our study clearly demonstrates that the c.835G>C (p.Gly279Arg) variant can lead to a decrease in fl-SMN1 transcripts by interrupting correct splicing of SMN1. What is more, the variant also affects SMN self-oligomerization via amino acid substitution from Gly to Arg at amino acid position of 279. This work presents the first evidence that it does exit double-hit events for the novel variant, which is crucial to understanding a severe SMA phenotype (type 1).




Similar content being viewed by others
References
Alias L, Bernal S, Fuentes-Prior P, Barcelo MJ, Also E, Martinez-Hernandez R, Rodriguez-Alvarez FJ, Martin Y, Aller E, Grau E, Pecina A, Antinolo G, Galan E, Rosa AL, Fernandez-Burriel M, Borrego S, Millan JM, Hernandez-Chico C, Baiget M, Tizzano EF (2009) Mutation update of spinal muscular atrophy in Spain: molecular characterization of 745 unrelated patients and identification of four novel mutations in the SMN1 gene. Hum Genet 125:29–39
Bai J, Qu Y, Cao Y, Yang L, Ge L, Jin Y, Wang H, Song F (2018) The SMN1 common variant c.22 dupA in Chinese patients causes spinal muscular atrophy by nonsense-mediated mRNA decay in humans. Gene 644:49–55
Brichta L, Garbes L, Jedrzejowska M, Grellscheid SN, Holker I, Zimmermann K, Wirth B (2008) Nonsense-mediated messenger RNA decay of survival motor neuron 1 causes spinal muscular atrophy. Hum Genet 123:141–153
Bürglen L, Amiel J, Viollet L, Lefebvre S, Burlet P, Clermont O, Raclin V, Landrieu P, Verloes A, Munnich A, Melki J (1996) Survival motor neuron gene deletion in the arthrogryposis multiplex congenita-spinal muscular atrophy association. J Clin Invest 98(5):1130–1132
Clermont O, Burlet P, Benit P, Chanterau D, Saugier-Veber P, Munnich A, Cusin V (2004) Molecular analysis of SMA patients without homozygous SMN1 deletions using a new strategy for identification of SMN1 subtle mutations. Hum Mutat 24:417–427
Eggermann T, Eggermann K, Elbracht M, Zerres K, Rudnik-Schoneborn S (2008) A new splice site mutation in the SMN1 gene causes discrepant results in SMN1 deletion screening approaches. Neuromuscul Disord 18(2):146–149
Hahnen E, Schönling J, Rudnik-Schöneborn S, Raschke H, Zerres K, Wirth B (1997) Missense mutations in exon 6 of the survival motor neuron gene in patients with spinal muscular atrophy (SMA). Hum Mol Genet 6(5):821–825
He J, Zhang QJ, Lin QF, Chen YF, Lin XZ, Lin MT, Murong SX, Wang N, Chen WJ (2013) Molecular analysis of SMN1, SMN2, NAIP, GTF2H2, and H4F5 genes in 157 Chinese patients with spinal muscular atrophy. Gene 518(2):325–329. https://doi.org/10.1016/j.gene.2012.12.109
Jedrzejowska M, Gos M, Zimowski JG, Kostera-Pruszczyk A, Ryniewicz B, Hausmanowa-Petrusewicz I (2014) Novel point mutations in survival motor neuron 1 gene expand the spectrum of phenotypes observed in spinal muscular atrophy patients. Neuromuscul Disord 24:617–623
Kang SH, Cho SI, Chae JH, Chung KN, Ra EK, Kim SY, Seong MW, Kim JY, Park SS (2009) False homozygous deletions of SMN1 exon 7 using Dra I PCR-RFLP caused by a novel mutation in spinal muscular atrophy. Genet Test Mol Biomarkers 13(4):511–513
Kolb SJ, Kissel JT (2011) Spinal muscular atrophy: a timely review. Arch Neurol 68:979–984
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L, Benichou B, Cruaud C, Millasseau P, Zeviani M et al (1995) Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80:155–165
Lorson CL, Strasswimmer J, Yao JM, Baleja JD, Hahnen E, Wirth B, Le T, Burghes AH, Androphy EJ (1998) SMN oligomerization defect correlates with spinal muscular atrophy severity. Nat Genet 19:63–66
Lorson CL, Hahnen E, Androphy EJ, Wirth B (1999) A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc Natl Acad Sci U S A 96:6307–6311
Lunn MR, Wang CH (2008) Spinal muscular atrophy. Lancet 371:2120–2133
Mailman MD, Heinz JW, Papp AC, Snyder PJ, Sedra MS, Wirth B, Burghes AH, Prior TW (2002) Molecular analysis of spinal muscular atrophy and modification of the phenotype by SMN2. Genet Med 4(1):20–26
Martin R, Gupta K, Ninan NS, Perry K, Van Duyne GD (2012) The survival motor neuron protein forms soluble glycine zipper oligomers. Structure 20:1929–1939
McAndrew PE, Parsons DW, Simard LR, Rochette C, Ray PN, Mendell JR, Prior TW, Burghes AH (1997) Identification of proximal spinal muscular atrophy carriers and patients by analysis of SMNT and SMNC gene copy number. Am J Hum Genet 60:1411–1422
Monani UR, Lorson CL, Parsons DW, Prior TW, Androphy EJ, Burghes AH, McPherson JD (1999) A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum Mol Genet 8:1177–1183
Parsons DW, McAndrew PE, Iannaccone ST, Mendell JR, Burghes AH, Prior TW (1998) Intragenic telSMN mutations: frequency, distribution, evidence of a founder effect, and modification of the spinal muscular atrophy phenotype by cenSMN copy number. Am J Hum Genet 63(6):1712–1723
Pearn J (1978) Incidence, prevalence, and gene frequency studies of chronic childhood spinal muscular atrophy. J Med Genet 15:409–413
Praveen K, Wen Y, Gray KM, Noto JJ, Patlolla AR, Van Duyne GD, Matera AG (2014) SMA-causing missense mutations in survival motor neuron (Smn) display a wide range of phenotypes when modeled in Drosophila. PLoS Genet 10:e1004489
Prior TW (2007) Spinal muscular atrophy diagnostics. J Child Neurol 22:952–956
Qu YJ, Bai JL, Cao YY, Wang H, Jin YW, Du J, Ge XS, Zhang WH, Li Y, He SX, Song F (2016a) Mutation spectrum of the survival of motor neuron 1 and functional analysis of variants in Chinese spinal muscular atrophy. J Mol Diagn 18:741–752
Qu YJ, Bai JL, Cao YY, Zhang WH, Wang H, Jin YW, Song F (2016b) A rare variant (c.863G>T) in exon 7 of SMN1 disrupts mRNA splicing and is responsible for spinal muscular atrophy. Eur J Hum Genet 24:864–870
Rochette CF, Surh LC, Ray PN, McAndrew PE, Prior TW, Burghes AH, Vanasse M, Simard LR (1997) Molecular diagnosis of non-deletion SMA patients using quantitative PCR of SMN exon 7. Neurogenetics 1(2):141–147
Ronchi D, Previtali SC, Sora MG, Barera G, Del Menico B, Corti S, Bresolin N, Comi GP (2015) Novel splice-site mutation in SMN1 associated with a very severe SMA-I phenotype. J Mol Neurosci 56(1):212–215
Sheng-Yuan Z, Xiong F, Chen YJ, Yan TZ, Zeng J, Li L, Zhang YN, Chen WQ, Bao XH, Zhang C, Xu XM (2010) Molecular characterization of SMN copy number derived from carrier screening and from core families with SMA in a Chinese population. Eur J Hum Genet 18(9):978–984
Singh RN, Howell MD, Ottesen EW, Singh NN (2017) Diverse role of survival motor neuron protein. Biochim Biophys Acta Gene Regul Mech 1860:299–315
Skordis LA, Dunckley MG, Burglen L, Campbell L, Talbot K, Patel S, Melki J, Davies KE, Dubowitz V, Muntoni F (2001) Characterisation of novel point mutations in the survival motor neuron gene SMN, in three patients with SMA. Hum Genet 108(4):356–357
Sun Y, Grimmler M, Schwarzer V, Schoenen F, Fischer U, Wirth B (2005) Molecular and functional analysis of intragenic SMN1 mutations in patients with spinal muscular atrophy. Hum Mutat 25(1):64–71
Talbot K, Ponting CP, Theodosiou AM, Rodrigues NR, Surtees R, Mountford R, Davies KE (1997) Missense mutation clustering in the survival motor neuron gene: a role for a conserved tyrosine and glycine rich region of the protein in RNA metabolism? Hum Mol Genet 6(3):497–500
Thauvin-Robinet C, Drunat S, Saugier Veber P, Chantereau D, Cossee M, Cassini C, Soichot P, Masurel-Paulet A, De Monleon JV, Sagot P, Huet F, Antin M, Calmels N, Faivre L, Gerard B (2012) Homozygous SMN1 exons 1-6 deletion: pitfalls in genetic counseling and general recommendations for spinal muscular atrophy molecular diagnosis. Am J Med Genet A 158a(7):1735–1741
Thieme A, Mitulla B, Schulze F, Spiegler AW (1993) Epidemiological data on Werdnig-Hoffmann disease in Germany (West-Thuringen). Hum Genet 91:295–297
van der Steege G, Grootscholten PM, van der Vlies P, Draaijers TG, Osinga J, Cobben JM, Scheffer H, Buys CH (1995) PCR-based DNA test to confirm clinical diagnosis of autosomal recessive spinal muscular atrophy. Lancet (London, England) 345:985–986
Vezain M, Gerard B, Drunat S, Funalot B, Fehrenbach S, N’Guyen-Viet V, Vallat JM, Frebourg T, Tosi M, Martins A, Saugier-Veber P (2011) A leaky splicing mutation affecting SMN1 exon 7 inclusion explains an unexpected mild case of spinal muscular atrophy. Hum Mutat 32(9):989–994
Wang CH, Papendick BD, Bruinsma P, Day JK (1998) Identification of a novel missense mutation of the SMN(T) gene in two siblings with spinal muscular atrophy. Neurogenetics 1(4):273–276
Wirth B (2000) An update of the mutation spectrum of the survival motor neuron gene (SMN1) in autosomal recessive spinal muscular atrophy (SMA). Hum Mutat 15:228–237
Wirth B, Herz M, Wetter A, Moskau S, Hahnen E, Rudnik-Schoneborn S, Wienker T, Zerres K (1999) Quantitative analysis of survival motor neuron copies: identification of subtle SMN1 mutations in patients with spinal muscular atrophy, genotype-phenotype correlation, and implications for genetic counseling. Am J Hum Genet 64:1340–1356
Wu S, Li YL, Cheng NY, Wang C, Dong EL, Lu YQ, Li JJ, Guo XX, Lin X, Lai LL, Liu ZW, Wang N, Chen WJ (2018) c.835-5T>G variant in SMN1 gene causes transcript exclusion of exon 7 and spinal muscular atrophy. J Mol Neurosci 65:196–202
Yu-Jin Q, Juan D, Er-zhen L, Jin-li B, Yu-wei J, Hong W, Fang S (2012) Subtle mutations in the SMN1 gene in Chinese patients with SMA: p.Arg288Met mutation causing SMN1 transcript exclusion of exon7. BMC Med Genet 13:86. https://doi.org/10.1186/1471-2350-13-86
Acknowledgments
We are grateful to the SMA family for their cooperation in our study.
Funding
This work was supported by the National Natural Science Foundation of China (Project Nos. 81501850, 81470056 and 81509719), the Beijing Municipal Administration of Hospitals’ Youth Program (QML20161303), the CAMS Initiative for Innovative Medicine (CAMS-I2M), the National Key Research and Development Program of China (No. 2016YFC0901505), and the Research Foundation of Capital Institute of Pediatrics (No. FX-2020-01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the Capital Institute of Pediatric Ethics Committee.
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bai, J., Qu, Y., Song, F. et al. Dual Mechanism of a New SMN1 Variant (c.835G>C, p.Gly279Arg) by Interrupting Exon 7 Skipping and YG Oligomerization in Causation of Spinal Muscular Atrophy. J Mol Neurosci 71, 112–121 (2021). https://doi.org/10.1007/s12031-020-01631-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-020-01631-7