Abstract
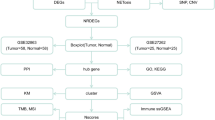
Atypical teratoid/rhabdoid tumor (ATRT) is a devastating intracranial tumor in children. Currently, its molecular mechanisms cannot be studied effectively because patient samples are limited, and many factors are involved in its pathogenesis. In this study, we analyzed three gene expression profile data sets obtained from the Gene Expression Omnibus (GEO) database to identify genes that participate in ATRT. The datasets were integrated and analyzed using the RobustRankAggreg method to screen for differentially expressed genes (DEGs). We identified 197 DEGs, including 94 downregulated and 103 upregulated genes which were then used for gene set enrichment analysis. The results showed that the downregulated genes were mainly enriched in synaptic vesicle cycle, nicotine addiction, and GABAergic synapse, whereas the upregulated genes were enriched in the cell cycle, p53 signaling pathway, and cellular senescence. Consistent with these results, gene set enrichment analysis showed that E2F targets, G2M checkpoints, and MYC targets were significantly enriched in datasets. Protein-protein interaction (PPI) network revealed that CDK1, CCNA2, BUB1B, CDC20, KIF11, KIF20A, KIF2C, NCAPG, NDC80, NUSAP1, PBK, RRM2, TPX2, TOP2A, and TTK were hub genes. NetworkAnalyst algorithm was used to predict the transcription factor (TF), and the results showed that MYC, SOX2, and KDM5B could regulate these hub genes. In conclusion, the present study brings a new perspective of ATRT pathogenesis and the strategy targeted to cell cycle related gene may be promising treatments for the disease.






Similar content being viewed by others
References
Alonso MM, Fueyo J, Gomez-Manzano C (2011) Malignant Gliomas: role of E2F1 transcription factor. In: Tumors of the Central Nervous System, Volume 1. pp 89–97. doi:https://doi.org/10.1007/978-94-007-0344-5_10
Bie L, Zhao G, Wang YP, Zhang B (2012) Kinesin family member 2C (KIF2C/MCAK) is a novel marker for prognosis in human gliomas clinical neurology and neurosurgery 114:356-360 doi:https://doi.org/10.1016/j.clineuro.2011.11.005
Brooks TA, Hurley LH (2009) The role of supercoiling in transcriptional control of MYC and its importance in molecular therapeutics. Nat Rev Cancer 9:849–861. https://doi.org/10.1038/nrc2733
Cancer Genome Atlas Research N et al (2017) Integrated genomic characterization of oesophageal carcinoma. Nature 541:169–175. https://doi.org/10.1038/nature20805
Chen W et al (2014) MiR-136 targets E2F1 to reverse cisplatin chemosensitivity in glioma cells. J Neuro-Oncol 120:43–53. https://doi.org/10.1007/s11060-014-1535-x
Cole MD, Cowling VH (2008) Transcription-independent functions of MYC: regulation of translation and DNA replication. Nat Rev Mol Cell Biol 9:810–815. https://doi.org/10.1038/nrm2467
Duan J, Huang W, Shi H (2016) Positive expression of KIF20A indicates poor prognosis of glioma patients. OncoTargets and therapy 9:6741–6749. https://doi.org/10.2147/ott.S115974
Feng Y, Li L, Du Y, Peng X, Chen F (2020) E2F4 functions as a tumour suppressor in acute myeloid leukaemia via inhibition of the MAPK signalling pathway by binding to EZH2. J Cell Mol Med 24:2157–2168. https://doi.org/10.1111/jcmm.14853
Fruhwald MC, Biegel JA, Bourdeaut F, Roberts CW, Chi SN (2016) Atypical teratoid/rhabdoid tumors-current concepts, advances in biology, and potential future therapies. Neuro-Oncology 18:764–778. https://doi.org/10.1093/neuonc/nov264
Gao X, Chen Y, Chen M, Wang S, Wen X, Zhang S (2018) Identification of key candidate genes and biological pathways in bladder cancer. PeerJ 6:e6036. https://doi.org/10.7717/peerj.6036
Harmeyer KM, Facompre ND, Herlyn M, Basu D (2017) JARID1 histone Demethylases: emerging targets in cancer. Trends Cancer 3:713–725. https://doi.org/10.1016/j.trecan.2017.08.004
Hnit SST et al (2019) Transcriptional regulation of G2/M regulatory proteins and perturbation of G2/M cell cycle transition by a traditional Chinese medicine recipe. J Ethnopharmacol 251:112526. https://doi.org/10.1016/j.jep.2019.112526
Huang P, Guo YD, Zhang HW (2019) Identification of Hub Genes in Pediatric Medulloblastoma by Multiple-Microarray Analysis Journal of molecular neuroscience : MN doi:https://doi.org/10.1007/s12031-019-01451-4
Jackson EM et al (2009) Genomic analysis using high-density single nucleotide polymorphism-based oligonucleotide arrays and multiplex ligation-dependent probe amplification provides a comprehensive analysis of INI1/SMARCB1 in malignant rhabdoid tumors. Clin Cancer Res 15:1923–1930. https://doi.org/10.1158/1078-0432.Ccr-08-2091
Jin Q, Dai Y, Wang Y, Zhang S, Liu G (2019) High kinesin family member 11 expression predicts poor prognosis in patients with clear cell renal cell carcinoma. J Clin Pathol 72:354–362. https://doi.org/10.1136/jclinpath-2018-205390
Johann PD et al (2016) Atypical Teratoid/Rhabdoid tumors are comprised of three epigenetic subgroups with distinct enhancer landscapes. Cancer Cell 29:379–393. https://doi.org/10.1016/j.ccell.2016.02.001
Ma Q, Liu Y, Shang L, Yu J, Qu Q (2017) The FOXM1/BUB1B signaling pathway is essential for the tumorigenicity and radioresistance of glioblastoma. Oncol Rep 38:3367–3375. https://doi.org/10.3892/or.2017.6032
Mercurio S, Serra L, Nicolis SK (2019) More than just stem cells: functional roles of the transcription factor Sox2 in differentiated glia and neurons. International journal of molecular sciences 20 doi:https://doi.org/10.3390/ijms20184540
Mollaoglu G et al. (2018) The lineage-defining transcription factors SOX2 and NKX2-1 determine lung Cancer cell fate and shape the tumor immune microenvironment Immunity 49:764-779 e769 doi:https://doi.org/10.1016/j.immuni.2018.09.020
Mondal G, Sengupta S, Panda CK, Gollin SM, Saunders WS, Roychoudhury S (2007) Overexpression of Cdc20 leads to impairment of the spindle assembly checkpoint and aneuploidization in oral cancer. Carcinogenesis 28:81–92. https://doi.org/10.1093/carcin/bgl100
Novak D, Huser L, Elton JJ, Umansky V, Altevogt P, Utikal J (2019) SOX2 in development and cancer biology Semin Cancer Biol doi:https://doi.org/10.1016/j.semcancer.2019.08.007
Phi JH et al (2019) NPM1 as a potential therapeutic target for atypical teratoid/rhabdoid tumors. BMC Cancer 19:848. https://doi.org/10.1186/s12885-019-6044-z
Singh AM, Dalton S (2009) The cell cycle and Myc intersect with mechanisms that regulate pluripotency and reprogramming. Cell Stem Cell 5:141–149. https://doi.org/10.1016/j.stem.2009.07.003
Suzuki M, Kondo A, Ogino I, Arai H, Tomita T, Sredni ST (2017) Overexpression of TEAD4 in atypical teratoid/rhabdoid tumor: new insight to the pathophysiology of an aggressive brain tumor. Pediatric blood & cancer 64 doi:https://doi.org/10.1002/pbc.26398
Szmyd R et al (2019) Premature activation of Cdk1 leads to mitotic events in S phase and embryonic lethality. Oncogene 38:998–1018. https://doi.org/10.1038/s41388-018-0464-0
Torchia J, Golbourn B, Feng S, Ho KC, Huang AJCC (2016) Integrated (epi)-genomic analyses identify subgroup-specific therapeutic targets in CNS. Rhabdoid Tumors 30:891–908
Vogel FCE et al (2019) Targeting the H3K4 Demethylase KDM5B reprograms the Metabolome and phenotype of melanoma cells. J Invest Dermatol 139:2506–2516 e2510. https://doi.org/10.1016/j.jid.2019.06.124
Wang Z et al (2014) Cyclin B1/Cdk1 coordinates mitochondrial respiration for cell-cycle G2/M progression. Dev Cell 29:217–232. https://doi.org/10.1016/j.devcel.2014.03.012
Wang J et al (2019) Invasion of white matter tracts by glioma stem cells is regulated by a NOTCH1-SOX2 positive-feedback loop. Nat Neurosci 22:91–105. https://doi.org/10.1038/s41593-018-0285-z
Yang R, Du Y, Wang L, Chen Z, Liu X (2020) Weighted gene co-expression network analysis identifies CCNA2 as a treatment target of prostate cancer through inhibiting cell cycle. J Cancer 11:1203–1211. https://doi.org/10.7150/jca.38173
Yuan J, Levitin HM, Frattini V, Bush EC, Boyett DM, Samanamud J, Ceccarelli M, Dovas A, Zanazzi G, Canoll P, Bruce JN, Lasorella A, Iavarone A, Sims PA (2018) Single-cell transcriptome analysis of lineage diversity in high-grade glioma. Genome Med 10:57. https://doi.org/10.1186/s13073-018-0567-9
Zou MR, Cao J, Liu Z, Huh SJ, Polyak K, Yan Q (2014) Histone demethylase jumonji AT-rich interactive domain 1B (JARID1B) controls mammary gland development by regulating key developmental and lineage specification genes. J Biol Chem 289:17620–17633. https://doi.org/10.1074/jbc.M114.570853
Acknowledgments
We thank Dr. Chris Lou (Blogger, WeChat Official Accounts) and Xiaoyahuatu for the bioinformatics technology support, Figure Ya (Blogger, WeChat Official Accounts) for the technical support in figure drafting and revision and Dr. Jerry, for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pan, X., Liu, W., Chai, Y. et al. Identification of Hub Genes in Atypical Teratoid/Rhabdoid Tumor by Bioinformatics Analyses. J Mol Neurosci 70, 1906–1913 (2020). https://doi.org/10.1007/s12031-020-01587-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-020-01587-8