Abstract
Autosomal recessive spastic ataxia of Charlevoix–Saguenay is a distinct form of hereditary early-onset spastic ataxia caused by cerebellum and spinal cord degeneration. The SACS gene has been demonstrated to be responsible for the disease through worldwide description of different mutations. We report here a computational analysis of a novel SACS gene mutation identified in a Tunisian family, using workflow implemented on the BioExtract Server. Several online computational tools are currently available to explore the effect of novel identified mutations in human and other organisms. Such analysis is time-consuming and generates a batch of files that researchers need to extract and save. The BioExtract Server workflow described here offers an easy way to execute the required tools together, avoiding entering queries independently in each web tool or service.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Bouchard J-P, Barbeau A, Bouchard R, Bouchard RW (1978) Autosomal recessive spastic ataxia of Charlevoix-Saguenay. Can J Neurol Sci 5:61–69
Bouchard JP, Richter A, Mathieu J et al (1998) Autosomal recessive spastic ataxia of Charlevoix-Saguenay. Neuromuscul Dis 8:474–479
Bouhlal Y, Zouari M, Kefi M, Ben Hamida C, Hentati F, Amouri R (2008) Autosomal recessive ataxia caused by three distinct gene defects in a single consanguineous family. J Neurogenet 22(2):139–148
Bouhlal Y, El Euch-Fayeche G, Hentati F, Amouri R (2009) A novel SACS gene mutation in a Tunisian family. J Mol Neurosci 39(3):333–336
Carson SD, Baggenstoss B (1991) Identification of peptides within a known protein sequence using COMSEQ analysis of data containing multiple sequences. Comput Methods Programs Biomed 35(1):35–42
Eisenberg D, Weiss RM, Terwilliger TC (1984) The hydrophobic moment detects periodicity in protein hydrophobicity. Proc Natl Acad Sci 81(1):140–144
El Euch-Fayache G, Lalani I, Amouri R et al (2003) Phenotypic features and genetic findings in sacsin-related autosomal recessive ataxia in Tunisia. Arch Neurol 60:982–988
Engert JC, Bérubé P, Mercier J et al (2000) ARSACS, a spastic ataxia common in northeastern Québec, is caused by mutations in a new gene encoding an 11.5-kb ORF. Nat Genet 24:120–125
Garnier J, Gibrat JF, Robson B (1996) GOR method for predicting protein secondary structure from amino acid sequence. Methods Enzymol 266:540–553
Ouyang Y, Takiyama Y, Sakoe K et al (2006) Sacsin-related ataxia (ARSACS): expanding the genotype upstream from the gigantic exon. Neurology 66:1103–1104
Parfitt DA, Michael GJ, Vermeulen EG et al (2009) The ataxia protein sacsin is a functional co-chaperone that protects against polyglutamine-expanded ataxin-1. Hum Mol Genet 18(9):1556–1565
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16(6):276–277
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Vermeer S, Meijer RP, Pijl BJ (2008) ARSACS in the Dutch population: a frequent cause of early-onset cerebellar ataxia. Neurogenetics 10(1):87
Acknowledgments
The authors would like to thank Dr. Volker Brendel for the helpful comments and suggestions on the earlier versions of their manuscript.
Funding
The BioExtract Server is funded through the National Science Foundation grant: DBI-0606906. V. Brendel, PI, C. Lushbough (U. South Dakota), and Carolyn Lawrence (ARS-USDA), co-PIs; Cyberinfrastructure for (Comparative) Plant Genome Research through PlantGDB.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplemental material 1
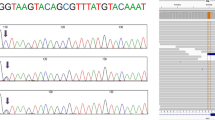
Supermatcher tool result showing the p.D4464Y substitution. (DOC 457 kb)
Supplemental material 2
Multiple alignment result showing that the D residue is highly conserved within species (DOC 602 kb)
Supplemental material 3
Pepstat analysis results: replacement of a polar residue in the wild-type protein (right) by a nonpolar one in the mutated protein (left). (DOC 422 kb)
Supplemental material 4
The predicted effect of p.D4464Y substitution on the secondary structure using GOR method. (DOC 420 kb)
Supplemental material 5
Hmoment analysis results showing modifications in the hydrophobic moment between the wild-type and the mutated protein (highlighted region). (DOC 115 kb)
Supplemental material 6
Comparison of hydrophobicity between the wild-type (left) and the mutated (right) protein with pep info tool using Kyte & Doolittle, OHM, and consensus parameters. Kyte & Doolittle methods do not predict any obvious change between the two plots, whereas hydrophobicity decreasing could be observed by the two remaining methods between residues 4450 and 4470. (DOC 1343 kb)
Rights and permissions
About this article
Cite this article
Bouhlal, Y., Jennewein, D.M., Anderson, B. et al. Computational Analysis of a Novel SACS Gene Mutation with BioExtract Server. J Mol Neurosci 44, 53–58 (2011). https://doi.org/10.1007/s12031-011-9512-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-011-9512-8