Abstract
Purpose
Metabolomic analysis in colorectal cancer (CRC) is an emerging research area with both prognostic and therapeutic targeting potential. We aimed to identify metabolomic pathway activity prognostic for CRC recurrence and overall survival and cross-reference such metabolomic data with prognostic genomic single-nucleotide polymorphisms (SNPs).
Methods
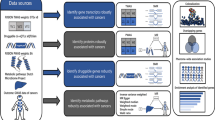
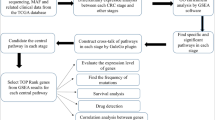
A systematic search of PubMed, Embase and Cochrane Library was performed for studies reporting prognostic metabolomic pathway activity in CRC in keeping with PRISMA guidelines. The QUADOMICS tool was used to assess study quality. MetaboAnalyst software (version4.0) was used to map metabolites that were associated with recurrence and survival in CRC to recognise metabolic pathways and identify genomic SNPs associated with CRC prognosis, referencing the following databases: Human Metabolome Database (HMDB), the Small Molecule Pathway Database (SMPDB), PubChem and Kyoto Encyclopaedia of Genes and Genomes (KEGG) Pathway Database.
Results
Nine studies met the inclusion criteria, reporting on 1117 patients. Increased metabolic activity in the urea cycle (p = 0.002, FDR = 0.198), ammonia recycling (p = 0.004, FDR = 0.359) and glycine and serine metabolism (p = 0.004, FDR = 0.374) was prognostic of CRC recurrence. Increased activity in aspartate metabolism (p < 0.001, FDR = 0.079) and ammonia recycling (p = 0.004, FDR = 0.345) was prognostic of survival. Eight resulting SNPs were prognostic for CRC recurrence (rs2194980, rs1392880, rs2567397, rs715, rs169712, rs2300701, rs313408, rs7018169) and three for survival (rs2194980, rs169712, rs12106698) of which two overlapped with recurrence (rs2194980, rs169712).
Conclusions
With a caveat on study heterogeneity, specific metabolites and metabolic pathway activity appear evident in the setting of poor prognostic colorectal cancers and such metabolic signatures are associated with specific genomic SNPs.



Similar content being viewed by others
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Dunn WB, Broadhurst DI, Atherton HJ, Goodacre R, Griffin JL. Systems level studies of mammalian metabolomes: the roles of mass spectrometry and nuclear magnetic resonance spectroscopy. Chem Soc Rev. 2011;40(1):387–426.
Romero-Garcia S, Lopez-Gonzalez JS, B´ez-Viveros JL, Aguilar-Cazares D, Prado-Garcia H. Tumor cell metabolism. Cancer Biol Ther. 2014;12(11):939–948.
Janke R, Dodson A, Rine J. Metabolism and epigenetics. Annu Rev Cell Dev Biol. 2015;31:473e496.
Santhanama S, Alvaradoa DM, Ciorba MA. Therapeutic targeting of inflammation and tryptophan metabolism in colon and gastrointestinal cancer. Transl Res. 2016;167(1):67–9.
Bettencourt IA, Powell JD. Targeting metabolism as a novel therapeutic approach to autoimmunity, inflammation, and transplantation. J Immunol. 2017;198(3):999–1005.
Phua LC, et al. Non-invasive fecal metabonomic detection of colorectal cancer. Cancer Biol Ther. 2014;15(4):389–397.
Bertini I, et al. Metabolomic NMR fingerprinting to identify and predict survival of patients with metastatic colorectal cancer. Cancer Res. 2012;72(37):356–65.
Long Z, et al. Metabolomic markers of colorectal tumor with different clinicopathological features. Frony Oncol. 2020;17(10):981.
Wang Z, et al. Development of a correlative strategy to discover colorectal tumor tissue derived metabolite biomarkers in plasma using untargeted metabolomics. Anal Chem. 2019;91(3):2401–8.
Zhang A, Sun H, Yan G, Wang P, Han Y, Wang X. Metabolomics in diagnosis and biomarker discovery of colorectal cancer. Cancer Lett. 2014;345(1):17–20.
Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2002;3(6):453–8.
O’Leary DP, O’Leary E, Foley N, Cotter TG, Wang JH, Redmond HP. Effects of surgery on the cancer stem cell niche. Eur J Surg Oncol. 2016;42(3):319–25.
Zhang F, et al. Metabolomics for biomarker discovery in the diagnosis, prognosis, survival and recurrence of colorectal cancer: a systematic review. Oncotarget. 2017;8(21):35460–72.
Yoo BC, Kim KH, Woo SM, Myung JK. Clinical multi-omics strategies for the effective cancer management. J Proteomics. 2017;0–1.
Vucic EA, et al. Translating cancer ‘omics’ to improved outcomes paradigm Translating cancer ‘omics’ to improved outcomes. Genome Research. 2012;188–195.
Liao Y. Akt and glucose metabolism in cancer (invited review). J Metab Nutr Cancer. 2014;1(2):80e88.
Dang L, White DW, Gross S, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2009;462(7274):739e744.
Page MJ, Matthew J, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ (Clinical research ed.) 2021;372:71.
Reitsma JB, Leeflang MMG, Sterne JAC, Bossuyt PMM. Research and reporting methods accuracy studies. 2011;4.
Lumbreras B, Porta M, Márquez S, Pollán M, Parker LA, Hernández-Aguado I. QUADOMICS: an adaptation of the quality assessment of diagnostic accuracy assessment (QUADAS) for the evaluation of the methodological quality of studies on the diagnostic accuracy of ‘-omics’-based technologies. Clin Biochem. 2008;41(16–17):1316–25.
Chong J, et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. 2018;0–9.
Sherry ST, Ward MH, Kholodov M, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001;29(1):308–311. https://doi.org/10.1093/nar/29.1.308.
Cai Y, et al. Sex differences in colon cancer metabolism reveal a novel subphenotype. Sci Rep. 2020;17(1):4905.
Zaimenko I, et al. Non-invasive metastasis prognosis from plasma metabolites in stage II colorectal cancer patients: the DACHS study. Int J Cancer. 2019;145(1):221–31.
De Vroome SW, et al. Serum N-glycome alterations in colorectal cancer associate with survival. Oncotarget. 2018;9(55):30610–23.
Pacholczyk-sienicka B, Fabia A. Prediction of survival for patients with advanced colorectal cancer using 1 H high-resolution magic angle spinning nuclear MR spectroscopy. J Magn Reson Imaging. 2015;1674:1669–74.
Qiu Y, et al. A distinct metabolic signature of human colorectal cancer with prognostic potential. Clin Cancer Res. 2014;20(8):2136–47.
Jimenez B, et al. H HR-MAS NMR spectroscopy of tumor-induced local metabolic ‘field-effects’ enables colorectal cancer staging and prognostication. J Proteome Res. 2013;12(2):959–68.
Farshidfar F, et al. Serum metabolomic profile as a means to distinguish stage of colorectal cancer. Genome Med. 2012;4(5):42.
Beger RD, et al. Metabolomics enables precision medicine: a white paper, community perspective. Metabolomics. 2016;12:10.
Spratlin JL, Serkova NJ, Eckhardt SG. Clinical applications of metabolomics in oncology: a review. Clin Cancer Res. 2009;15(2):431–40.
Zhang F, et al. Applications of metabolomics in cancer studies. Oncotarget. 2017;345(1):552–74.
Morris E, et al. Surgical management and outcomes of colorectal cancer liver metastases. Br J Surg. 2010;97(7):1110.
Väyrynen V, et al. Incidence and management of patients with colorectal cancer and synchronous and metachronous colorectal metastases: a population-based study. BJS Open. 2020;4(4):685–92.
Shinkins B, et al. Serum carcinoembryonic antigen trends for diagnosing colorectal cancer recurrence in the FACS randomized clinical trial. Br J Surg. 2018;105(6):658–62.
Leong K, Hartley J, Karandikar S. Association of Coloproctology of Great Britain & Ireland (ACPGBI): guidelines for the management of cancer of the colon, rectum and anus (2017) – follow up, lifestyle and survivorship. Color Dis. 2017;19:67–70.
Ryuk JP, et al. Predictive factors and the prognosis of recurrence of colorectal cancer within 2 years after curative resection. Ann Surg Treat Res. 2014;86(3):143–51.
Van der Geest LGM, Lam-Boer J, Koopman M, Verhoef C, Elferink MAG, de Wilt JHW. Nationwide trends in incidence, treatment and survival of colorectal cancer patients with synchronous metastases. Clin Exp Metastasis. 2015;32(5):457–65.
Keshet R, Szlosarek P, Carracedo A, Erez A. Rewiring urea cycle metabolism in cancer to support anabolism. Nat Rev Cancer. 2018;18.
Adeva M, Souto G, Blanco N, Donapetry C. Ammonia metabolism in humans. Metabolism. 2012;61:1495–511.
DeBerardinis R, Cheng T. The diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene. 2010;29:313–324.
Coloff J, et al. Differential glutamate metabolism in proliferating and quiescent mammary epithelial cells. Cell Metab. 2016;23:867–880.
Haigis MC. Metabolic recycling of ammonia via glutamate dehydrogenase supports breast cancer biomass. Science. 2018;358(6365):941–6.
Gu Y, et al. Perioperative dynamics and significance of amino acid profiles in patients with cancer. J Transl Med. 2015;13(1):1–14.
Antonov A, Amelio I, Cutruzzola F, Agostini M, Melino G. Serine and glycine metabolism in cancer. Trends Biochem Sci. 2014;39(4):191–8.
Potter M, Newport E, Morten K. The Warburg effect: 80 years on. Biochem Soc Trans. 2016;44(5)1499e1505.
Warburg O, Wind F, Negelein E. The metabolism of tumors in the body. J Gen Physiol. 1927;8:519–30.
Shyh-Chang N, et al. Influence of threonine metabolism on S-adenosylmethionine and histone methylation. Science. 2013;339:222–22638.
Maddocks OD, et al. Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature. 2013;493:542–546.
Han T, et al. How does cancer cell metabolism affect tumor migration and invasion? Cell Adhes Migr. 2013;7(5):395–403.
Mayers J, Vander Heiden M. Famine versus feast: understanding the metabolism of tumors in vivo. Trends Biochem Sci. 2015;40:130–140.
Batool T, Makky E, Jalal M, Yusoff M. A comprehensive review on l-asparaginase and its applications. Appl Biochem Biotechnol. 2016;178:900–23.
Wu X, Zhao J, Ruan Y, Sun L, Xu C, Jiang H. Sialyltransferase ST3GAL1 promotes cell migration, invasion, and TGF-β1-induced EMT and confers paclitaxel resistance in ovarian cancer. Cell Death Dis. 2018;9(11).
Zhang L, Al E. Effects of Kras activation and Pten deletion alone or in combination on MUC1 biology and epithelial-to-mesenchymal transition in ovarian cancer. Oncogene. 2016;35:5010–20.
Canel M, Serrels A, Frame M, Brunton V. E-cadherin-integrin crosstalk in cancer invasion and metastasis. J Cell Sci. 2013;126:393–401.
Acknowledgements
All studies from which data was extracted are accurately referenced.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Previous communication: Awarded the British Association of Surgical Oncology (BASO)-Association of Cancer Surgeons (ACS) Proffered Paper Prize, November 2020
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fleming, C.A., Mohan, H.M., O’Leary, D.P. et al. Metabolomic Pathway Activity with Genomic Single-Nucleotide Polymorphisms Associated with Colorectal Cancer Recurrence and 5-Year Overall Survival. J Gastrointest Canc 54, 247–258 (2023). https://doi.org/10.1007/s12029-022-00813-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-022-00813-3