Abstract
Summary
It has been demonstrated that epigenetic modifications of histone (acetylation/deacetylation) participate in a critical role in cancer progression by the regulation of gene expression. Several processes could be regulated by deacetylation of histone and non-histone proteins such as apoptosis, proliferation, cell metabolism, differentiation, and DNA repair. Hence, histone deacetylase inhibitors (HDACis) are employed as a hopeful group of anti-cancer drugs that could inhibit tumor cell proliferation or apoptosis. The elimination of the acetylation marks that take place as an essential epigenetic change in cancer cells is associated to HDAC expression and activity. In this regard, it has been reported that class I HDACs have a vital role in the regulation of tumor cell proliferation.
Objectives
In this review, we discuss whether gut origin microorganisms could promote cancer or tumor resistance and explain mechanisms of these processes.
Conclusions
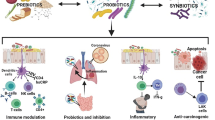
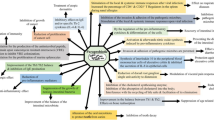
According to the enormous capacity of the metabolism of the intestine microbiota, bacteria are likely to convert nutrients and digestive compounds into metabolites that regulate epigenetic in cancer. The effect of the food is of interest on epigenetic changes in the intestinal mucosa and colonocytes, as misleading nucleotide methylation may be a prognostic marker for colorectal cancer (CRC). Since epigenetic changes are potentially reversible, they can serve as therapeutic targets for preventing CRC. However, various mechanisms have been identified in the field of prevention, treatment, and progression of cancer by probiotics, which include intestinal microbiota modulation, increased intestinal barrier function, degradation of potential carcinogens, protective effect on intestinal epithelial damage, and increased immune function.


Similar content being viewed by others
References
Eslami M, et al. Importance of probiotics in the prevention and treatment of colorectal cancer. J Cell Physiol. 2019.
Bultman SJ. Interplay between diet, gut microbiota, epigenetic events, and colorectal cancer. Mol Nutri Food Res. 2017;61(1):1500902.
Vaissière T, Sawan C, Herceg Z. Epigenetic interplay between histone modifications and DNA methylation in gene silencing. Mutat Res/Reviews in Mutation Research. 2008;659(1-2):40–8.
Guerra A, et al. Relevance and challenges in modeling human gastric and small intestinal digestion. Trends biotechnol. 2012;30(11):591–600.
Peterson LW, Artis D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat Rev Immunol. 2014;14(3):141.
McCracken VJ, Lorenz RG. The gastrointestinal ecosystem: a precarious alliance among epithelium, immunity and microbiota: Microreview. Cell Microbiol. 2001;3(1):1–11.
Groh V, et al. Cell stress-regulated human major histocompatibility complex class I gene expressed in gastrointestinal epithelium. Proc Natl Acad Sc. 1996;93(22):12445–50.
Marks PA, et al. Histone deacetylases and cancer: causes and therapies. Nat Rev Cancer. 2001;1(3):194.
Marks PA, Richon VM, Rifkind RA. Histone deacetylase inhibitors: inducers of differentiation or apoptosis of transformed cells. J Natil Cancer Inst. 2000;92(15):1210–6.
Grunstein M. Histone acetylation in chromatin structure and transcription. Nature. 1997;389(6649):349.
Wolffe AP. Histone Deacetylase--A Regulator of Transcription. Science. 1996;272(5260):371–2.
Zentner GE, Henikoff S. Regulation of nucleosome dynamics by histone modifications. Nat Struct Mol Biol. 2013;20(3):259.
Madrigal P, Krajewski P. Uncovering correlated variability in epigenomic datasets using the Karhunen-Loeve transform. BioData mining. 2015;8(1):20.
Spange S, et al. Acetylation of non-histone proteins modulates cellular signalling at multiple levels. Int J Biochem Cell Biol. 2009;41(1):185–98.
Cheung P, Allis CD, Sassone-Corsi P. Signaling to chromatin through histone modifications. Cell. 2000;103(2):263–71.
Winston F, Allis CD. The bromodomain: a chromatin-targeting module? Nat Struct Mol Biol. 1999;6(7):601.
Ng HH, Bird A. Histone deacetylases: silencers for hire. Trends Biochem Sci. 2000;25(3):121–6.
Lusser A, Kölle D, Loidl P. Histone acetylation: lessons from the plant kingdom. Trends Plant Sci. 2001;6(2):59–65.
Gregoretti I, Lee Y-M, Goodson HV. Molecular evolution of the histone deacetylase family: functional implications of phylogenetic analysis. J Mol Biol. 2004;338(1):17–31.
Glozak M, Seto E. Histone deacetylases and cancer. Oncogene. 2007;26(37):5420.
Witt O, et al. HDAC family: What are the cancer relevant targets? Cancer Lett. 2009;277(1):8–21.
Dokmanovic M, Clarke C, Marks PA. Histone deacetylase inhibitors: overview and perspectives. Mol Cancer Res. 2007;5(10):981–9.
Minucci S, Pelicci PG. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer. 2006;6(1):38.
Marks PA, Miller T, Richon VM. Histone deacetylases. Curr Opin Pharmacol. 2003;3(4):344–51.
De Ruijter AJ, et al. Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J. 2003;370(3):737–49.
Bolden JE, Peart MJ, Johnstone RW. Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov. 2006;5(9):769.
Glaser KB, et al. Gene expression profiling of multiple histone deacetylase (HDAC) inhibitors: defining a common gene set produced by HDAC inhibition in T24 and MDA carcinoma cell lines. Mol Cancer Ther. 2003;2(2):151–63.
Gump JM, Thorburn A. Autophagy and apoptosis: what is the connection? Trends Cell Biol. 2011;21(7):387–92.
Eslami M, et al. Current information on the association of Helicobacter pylori with autophagy and gastric cancer. J Cell Physiol. 2019.
Yousefi B, et al. Role of autophagy associated with Helicobacter pylori CagA and VacA toxins in gastric cancer. Koomesh. 2019;21(2):205–14.
West AC, Johnstone RW. New and emerging HDAC inhibitors for cancer treatment. J Clin Invest. 2014;124(1):30–9.
Sayers TJ. Targeting the extrinsic apoptosis signaling pathway for cancer therapy. Cancer Immunol Immunother. 2011;60(8):1173–80.
Rahmani M, et al. Coadministration of histone deacetylase inhibitors and perifosine synergistically induces apoptosis in human leukemia cells through Akt and ERK1/2 inactivation and the generation of ceramide and reactive oxygen species. Cancer Res. 2005;65(6):2422–32.
Yang X, Seto E. HATs and HDACs: from structure, function and regulation to novel strategies for therapy and prevention. Oncogene. 2007;26(37):5310.
Shin J, et al. The intestinal epithelial cell differentiation marker intestinal alkaline phosphatase (ALPi) is selectively induced by histone deacetylase inhibitors (HDACi) in colon cancer cells in a Kruppel-like factor 5 (KLF5)-dependent manner. J Biol Chem. 2014;289(36):25306–16.
Felekkis K, et al. microRNAs: a newly described class of encoded molecules that play a role in health and disease. Hippokratia. 2010;14(4):236.
Bandres E, et al. MicroRNAs as cancer players: potential clinical and biological effects. DNA Cell Biol. 2007;26(5):273–82.
Zhu P, et al. Induction of HDAC2 expression upon loss of APC in colorectal tumorigenesis. Cancer Cell. 2004;5(5):455–63.
Mao Q, et al. MicroRNA-455 suppresses the oncogenic function of HDAC2 in human colorectal cancer. Braz J Med Biol Res. 2017:50(6).
De Palma M, Biziato D, Petrova TV. Microenvironmental regulation of tumour angiogenesis. Nat Rev Cancer. 2017;17(8):457.
Chou C-W, et al. HDAC inhibition decreases the expression of EGFR in colorectal cancer cells. PloS one. 2011;6(3):e18087.
Buckley DL, et al. Targeting the von Hippel–Lindau E3 ubiquitin ligase using small molecules to disrupt the VHL/HIF-1α interaction. J Am Chem Soc. 2012;134(10):4465–8.
Hagelkruys A, et al. The biology of HDAC in cancer: the nuclear and epigenetic components, in Histone Deacetylases: the Biology and Clinical Implication. In: Springer; 2011. p. 13–37.
Xu W, Parmigiani R, Marks P. Histone deacetylase inhibitors: molecular mechanisms of action. Oncogene. 2007;26(37):5541.
Carew JS, Giles FJ, Nawrocki ST. Histone deacetylase inhibitors: mechanisms of cell death and promise in combination cancer therapy. Cancer Lett. 2008;269(1):7–17.
Andresen L, et al. Molecular regulation of MHC class I chain-related protein A expression after HDAC-inhibitor treatment of Jurkat T cells. J Immunol. 2007;179(12):8235–42.
Shen L, Pili R. Class I histone deacetylase inhibition is a novel mechanism to target regulatory T cells in immunotherapy. Oncoimmunology. 2012;1(6):948–50.
Hull EE, Montgomery MR, Leyva KJ. HDAC inhibitors as epigenetic regulators of the immune system: impacts on cancer therapy and inflammatory diseases. BioMed Res Int. 2016;2016.
Wang JC, Dick JE. Cancer stem cells: lessons from leukemia. Trends Cell Biol. 2005;15(9):494–501.
Debeb BG, et al. Histone deacetylase inhibitor-induced cancer stem cells exhibit high pentose phosphate pathway metabolism. Oncotarget. 2016;7(19):28329.
Pantic I. Cancer stem cell hypotheses: impact on modern molecular physiology and pharmacology research. J Biosci. 2011;36(5):957–61.
Rotili D, et al. Simplification of the tetracyclic SIRT1-selective inhibitor MC2141: coumarin-and pyrimidine-based SIRT1/2 inhibitors with different selectivity profile. Bioorg Med Chem. 2011;19(12):3659–68.
Salek Farrokhi A, et al. Is it true that gut microbiota is considered as panacea in cancer therapy? J Cell Physiol. 2019.
Burns MB, et al. Virulence genes are a signature of the microbiome in the colorectal tumor microenvironment. Genome Med. 2015;7(1):55.
Greiner T, Bäckhed F. Effects of the gut microbiota on obesity and glucose homeostasis. Trends Endocrinol Metab. 2011;22(4):117–23.
Yousefi B, et al. Probiotics importance and their immunomodulatory properties. J Cell Physiol. 2019;234(6):8008–18.
Aoyama M, Kotani J, Usami M. Butyrate and propionate induced activated or non-activated neutrophil apoptosis via HDAC inhibitor activity but without activating GPR-41/GPR-43 pathways. Nutrition. 2010;26(6):653–61.
Meeran SM, Ahmed A, Tollefsbol TO. Epigenetic targets of bioactive dietary components for cancer prevention and therapy. Clin Epigenetics. 2010;1(3):101.
Meijer K, et al. Cell-based screening assay for anti-inflammatory activity of bioactive compounds. Food Chem. 2015;166:158–64.
Ghasemian A, et al. Probiotics and their increasing importance in human health and infection control. Rev Med Microbiol. 2018;29(4):153–8.
Abel T, Zukin RS. Epigenetic targets of HDAC inhibition in neurodegenerative and psychiatric disorders. Curr Opin Pharmacol. 2008;8(1):57–64.
Bourassa MW, et al. Butyrate, neuroepigenetics and the gut microbiome: can a high fiber diet improve brain health? Neurosci Lett. 2016;625:56–63.
Davie JR. Inhibition of histone deacetylase activity by butyrate. J Nutr. 2003;133(7):2485S–93S.
Ali SR, et al. Impact of histone deacetylase inhibitors on microRNA expression and cancer therapy: a review. Drug Dev Res. 2015;76(6):296–317.
Hamer HM, et al. The role of butyrate on colonic function. Aliment Pharmacol Ther. 2008;27(2):104–19.
Mei S, Ho AD, Mahlknecht U. Role of histone deacetylase inhibitors in the treatment of cancer. Int J Oncol. 2004;25(6):1509–19.
Bonci D, et al. The miR-15a–miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med. 2008;14(11):1271.
Farazi TA, et al. MicroRNAs in human cancer, in MicroRNA cancer regulation. In: Springer; 2013. p. 1–20.
Li Y, Seto E. HDACs and HDAC inhibitors in cancer development and therapy. Cold Spring Harb Perspect Med. 2016;6(10):a026831.
Lauffer BE, et al. Histone deacetylase (HDAC) inhibitor kinetic rate constants correlate with cellular histone acetylation but not transcription and cell viability. J Biol Chem. 2013;288(37):26926–43.
Codd R, et al. Zn (II)-dependent histone deacetylase inhibitors: suberoylanilide hydroxamic acid and trichostatin A. Int J Biochem Cell Biol. 2009;41(4):736–9.
Nalls D, et al. Targeting epigenetic regulation of miR-34a for treatment of pancreatic cancer by inhibition of pancreatic cancer stem cells. PloS one. 2011;6(8):e24099.
Mann BS, et al. FDA approval summary: vorinostat for treatment of advanced primary cutaneous T-cell lymphoma. Oncologist. 2007;12(10):1247–52.
Beckers T, et al. Distinct pharmacological properties of second generation HDAC inhibitors with the benzamide or hydroxamate head group. Int J Cancer. 2007;121(5):1138–48.
Zhao Z-N, et al. TSA suppresses miR-106b-93-25 cluster expression through downregulation of MYC and inhibits proliferation and induces apoptosis in human EMC. PLoS One. 2012;7(9):e45133.
Yousefi B, et al. Probiotics can really cure an autoimmune disease? Gene Rep. 2019:100364.
Roller M, Rechkemmer G, Watzl B. Prebiotic inulin enriched with oligofructose in combination with the probiotics Lactobacillus rhamnosus and Bifidobacterium lactis modulates intestinal immune functions in rats. J Nutr. 2004;134(1):153–6.
Meurman JH. Probiotics: do they have a role in oral medicine and dentistry? Eur J Oral Sci. 2005;113(3):188–96.
Tuddenham S, Sears CL. The intestinal microbiome and health. Curr Opin Infect Dis. 2015;28(5):464.
Gholizadeh P, et al. Role of oral microbiome on oral cancers, a review. Biomed Pharmacother. 2016;84:552–8.
Rooks MG, Garrett WS. Gut microbiota, metabolites and host immunity. Nat Rev Immunol. 2016;16(6):341.
Lebeer S, Vanderleyden J, De Keersmaecker SC. Host interactions of probiotic bacterial surface molecules: comparison with commensals and pathogens. Nat Revi Microbiol. 2010;8(3):171.
Kumar M, et al. Cancer-preventing attributes of probiotics: an update. Int J Food Scie Nutr. 2010;61(5):473–96.
Pandey KR, Naik SR, Vakil BV. Probiotics, prebiotics and synbiotics-a review. J Food Sci Technol. 2015;52(12):7577–87.
Chakraborti C. The status of synbiotics in colorectal cancer. Life Sci Med Res. 2011;2011:1–13.
Acknowledgments
The authors thank Semnan University of Medical Sciences and Cancer Research Center of Semnan University of Medical Sciences.
Author information
Authors and Affiliations
Contributions
Amir Salek Farrokhi investigated and supervised the findings of this work, wrote the article, provided critical feedback, and helped the analysis of manuscript; Majid Eslami designed the study, helped supervise the project, and conceived the original idea, discussed the results, and commented on the manuscript; Bahman Yousefi contributed to the final version of the manuscript, supervised the project, and contributed to the interpretation of the results; Maryam Abdollahi developed the theoretical framework and processed the experimental data; Maryam Mohammadlou designed the model and the computational framework and analyzed the data.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Salek Farrokhi, A., Mohammadlou, M., Abdollahi, M. et al. Histone Deacetylase Modifications by Probiotics in Colorectal Cancer. J Gastrointest Canc 51, 754–764 (2020). https://doi.org/10.1007/s12029-019-00338-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-019-00338-2