Abstract
Background
Molecular switches in phosphatidylinositol 3-kinase (PI3K)-AKT signaling pathway may serve as potential targets for the treatment of colorectal cancer (CRC). This study aims to profile the gene alterations involved in PI3K-AKT signaling pathway in patients with CRC.
Methods
Tumoral and matched peritumoral tissues were collected from 15 CRC patients who went routine surgery. A human PI3K-AKT signaling pathway polymerase chain reaction (PCR) array, which profiled the transcriptional changes of a total number of 84 genes involved in the PI3K-AKT pathway, was then applied to determine the gene alterations in CRC tumoral tissue with matched peritumoral tissue as a healthy control. Subsequent real-time reverse transcription PCR and western blot (WB) with different subgroups of CRC patients were then performed to further validate the array findings.
Results
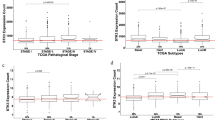
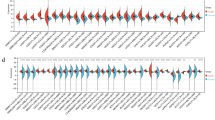
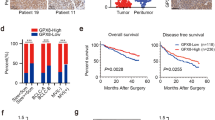
The PCR array identified 14 aberrantly expressed genes involved in the PI3K-AKT signaling pathway in CRC tumoral tissue, among which 12 genes, CCND1, CSNK2A1, EIF4E, EIF4EBP1, EIF4G1, FOS, GRB10, GSK3B, ILK, PTK2, PTPN11, and PHEB were significantly up-modulated (> two fold) while the remaining two, PDK1 and PIK3CG, were down-regulated (> two fold). These genes involve in the regulation of gene transcription and translation, cell cycle, and cell growth, proliferation, and differentiation. The real-time reverse transcription PCR validation agreed with the array data towards the tested genes, CCND1, EIF4E, FOS, and PIK3CG, while it failed to obtain similar result for PDK1. Interestingly, the WB analyses were further consistent with the PCR results that the protein levels of CCND1, EIF4E, and FOS were apparently up-regulated and that protein PIK3CG was down-modulated.
Conclusion
Taken together, the present study identified a deregulated PI3K-AKT signaling pathway in CRC patients, which might serve as therapeutic target(s).


Similar content being viewed by others
References
Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2016;0:1–9.
Weitz J, Koch M, Debus J, Hohler T, Galle PR, Buchler MW. Colorectal cancer. Lancet. 2005;365(9454):153–65.
Gout S, Huot J. Role of cancer microenvironment in metastasis: focus on colon cancer. Cancer Microenviron. 2008;1(1):69–83.
Whitman M, Kaplan DR, Schaffhausen B, Cantley L, Roberts TM. Association of phosphatidylinositol kinase activity with polyoma middle-T competent for transformation. Nature. 1985;315(6016):239–42.
Engelman JA, Luo J, Cantley LC. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet. 2006;7(8):606–19.
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304(5670):554.
Vivanco I, Sawyers CL. The phosphatidylinositol 3-kinase AKT pathway in human cancer. Nat Rev Cancer. 2002;2(7):489–501.
Papadatos-Pastos D, Rabbie R, Ross P, Sarker D. The role of the PI3K pathway in colorectal cancer. Crit Rev Oncol Hematol. 2015;94(1):18–30.
Baldin V, Lukas J, Marcote MJ, Pagano M, Draetta G. Cyclin D1 is a nuclear protein required for cell cycle progression in G1. Genes Dev. 1993;7(5):812–21.
Lewis RC, Bostick RM, Xie D, Deng Z, Wargovich MJ, Fina MF, et al. Polymorphism of the cyclin D1 gene, CCND1, and risk for incident sporadic colorectal adenomas. Cancer Res. 2003;63(23):8549–53.
Lee E, Jin D, Lee BB, Kim Y, Han J, Shim YM, et al. Negative effect of cyclin D1 overexpression on recurrence-free survival in stage II-IIIA lung adenocarcinoma and its expression modulation by vorinostat in vitro. BMC Cancer. 2015;15:982.
Burandt E, Grunert M, Lebeau A, Choschzick M, Quaas A, Janicke F, et al. Cyclin D1 gene amplification is highly homogeneous in breast cancer. Breast Cancer. 2016;23(1):111–9.
Bae JS, Park SH, Jamiyandorj U, Kim KM, Noh SJ, Kim JR, et al. CK2alpha/CSNK2A1 phosphorylates SIRT6 and is involved in the progression of breast carcinoma and predicts shorter survival of breast carcinoma patients. Am J Pathol. 2016;186(12):3297–315.
Wennerberg K, Rossman KL, Der CJ. The Ras superfamily at a glance. J Cell Sci. 2005;118(Pt 5):843–6.
Groenewoud MJ, Zwartkruis FJ. Rheb and Rags come together at the lysosome to activate mTORC1. Biochem Soc Trans. 2013;41(4):951–5.
Heard JJ, Fong V, Bathaie SZ, Tamanoi F. Recent progress in the study of the Rheb family GTPases. Cell Signal. 2014;26(9):1950–7.
Lu ZH, Shvartsman MB, Lee AY, Shao JM, Murray MM, Kladney RD, et al. Mammalian target of rapamycin activator RHEB is frequently overexpressed in human carcinomas and is critical and sufficient for skin epithelial carcinogenesis. Cancer Res. 2010;70(8):3287–98.
Jiang H, Vogt PK. Constitutively active Rheb induces oncogenic transformation. Oncogene. 2008;27(43):5729–40.
Durchdewald M, Angel P, Hess J. The transcription factor Fos: a Janus-type regulator in health and disease. Histol Histopathol. 2009;24(11):1451–61.
Gamberi G, Benassi MS, Bohling T, Ragazzini P, Molendini L, Sollazzo MR, et al. C-myc and c-fos in human osteosarcoma: prognostic value of mRNA and protein expression. Oncology. 1998;55(6):556–63.
Volm M, Koomagi R, Mattern J, Efferth T. Expression profile of genes in non-small cell lung carcinomas from long-term surviving patients. Clin Cancer Res. 2002;8(6):1843–8.
Silvestre DC, Gil GA, Tomasini N, Bussolino DF, Caputto BL. Growth of peripheral and central nervous system tumors is supported by cytoplasmic c-Fos in humans and mice. PLoS One. 2010;5(3):e9544.
Guinea-Viniegra J, Zenz R, Scheuch H, Jimenez M, Bakiri L, Petzelbauer P, et al. Differentiation-induced skin cancer suppression by FOS, p53, and TACE/ADAM17. J Clin Invest. 2012;122(8):2898–910.
Siddiqui N, Sonenberg N. Signalling to eIF4E in cancer. Biochem Soc Trans. 2015;43(5):763–72.
Rhoads RE. eIF4E: new family members, new binding partners, new roles. J Biol Chem. 2009;284(25):16711–5.
De Benedetti A, Graff JR. eIF-4E expression and its role in malignancies and metastases. Oncogene. 2004;23(18):3189–99.
Karaki S, Andrieu C, Ziouziou H, Rocchi P. The eukaryotic translation initiation factor 4E (eIF4E) as a therapeutic target for cancer. Adv Protein Chem Struct Biol. 2015;101:1–26.
Heesom KJ, Gampel A, Mellor H, Denton RM. Cell cycle-dependent phosphorylation of the translational repressor eIF-4E binding protein-1 (4E-BP1). Curr Biol. 2001;11(17):1374–9.
Topisirovic I, Ruiz-Gutierrez M, Borden KL. Phosphorylation of the eukaryotic translation initiation factor eIF4E contributes to its transformation and mRNA transport activities. Cancer Res. 2004;64(23):8639–42.
Chao MW, Wang LT, Lai CY, Yang XM, Cheng YW, Lee KH, et al. eIF4E binding protein 1 expression is associated with clinical survival outcomes in colorectal cancer. Oncotarget. 2015;6(27):24092–104.
Gingras AC, Raught B, Sonenberg N. mTOR signaling to translation. Curr Top Microbiol Immunol. 2004;279:169–97.
Cromer A, Carles A, Millon R, Ganguli G, Chalmel F, Lemaire F, et al. Identification of genes associated with tumorigenesis and metastatic potential of hypopharyngeal cancer by microarray analysis. Oncogene. 2004;23(14):2484–98.
Comtesse N, Keller A, Diesinger I, Bauer C, Kayser K, Huwer H, et al. Frequent overexpression of the genes FXR1, CLAPM1 and EIF4G located on amplicon 3q26-27 in squamous cell carcinoma of the lung. Int J Cancer. 2007;120(12):2538–44.
Silvera D, Arju R, Darvishian F, Levine PH, Zolfaghari L, Goldberg J, et al. Essential role for eIF4GI overexpression in the pathogenesis of inflammatory breast cancer. Nat Cell Biol. 2009;11(7):903–8.
Jiang X, Wang J, Zhang K, Tang S, Ren C, Chen Y. The role of CD29-ILK-Akt signaling-mediated epithelial-mesenchymal transition of liver epithelial cells and chemoresistance and radioresistance in hepatocellular carcinoma cells. Med Oncol. 2015;32(5):141.
Han KS, Li N, Raven PA, Fazli L, Ettinger S, Hong SJ, et al. Targeting integrin-linked kinase suppresses invasion and metastasis through downregulation of epithelial-to-mesenchymal transition in renal cell carcinoma. Mol Cancer Ther. 2015;14(4):1024–34.
Hannigan GE, McDonald PC, Walsh MP, Dedhar S. Integrin-linked kinase: not so ‘pseudo’ after all. Oncogene. 2011;30(43):4375–85.
Mohi MG, Neel BG. The role of Shp2 (PTPN11) in cancer. Curr Opin Genet Dev. 2007;17(1):23–30.
Gagliardi PA, di Blasio L, Primo L. PDK1: a signaling hub for cell migration and tumor invasion. Biochim Biophys Acta. 2015;1856(2):178–88.
Yoon S, Kim JG, Seo AN, Park SY, Kim HJ, Park JS, et al. Clinical implication of serine metabolism-associated enzymes in colon cancer. Oncology. 2015;89(6):351–9.
Hur H, Xuan Y, Kim YB, Lee G, Shim W, Yun J, et al. Expression of pyruvate dehydrogenase kinase-1 in gastric cancer as a potential therapeutic target. Int J Oncol. 2013;42(1):44–54.
Franke TF, Kaplan DR, Cantley LC. PI3K: downstream AKTion blocks apoptosis. Cell. 1997;88(4):435–7.
Cantley LC, Neel BG. New insights into tumor suppression: PTEN suppresses tumor formation by restraining the phosphoinositide 3-kinase/AKT pathway. Proc Natl Acad Sci U S A. 1999;96(8):4240–5.
Stoyanov B, Volinia S, Hanck T, Rubio I, Loubtchenkov M, Malek D, et al. Cloning and characterization of a G protein-activated human phosphoinositide-3 kinase. Science. 1995;269(5224):690–3.
Stephens LR, Eguinoa A, Erdjument-Bromage H, Lui M, Cooke F, Coadwell J, et al. The G beta gamma sensitivity of a PI3K is dependent upon a tightly associated adaptor, p101. Cell. 1997;89(1):105–14.
Semba S, Itoh N, Ito M, Youssef EM, Harada M, Moriya T, et al. Down-regulation of PIK3CG, a catalytic subunit of phosphatidylinositol 3-OH kinase, by CpG hypermethylation in human colorectal carcinoma. Clin Cancer Res. 2002;8(12):3824–31.
Lu CW, Lin SC, Chien CW, Lin SC, Lee CT, Lin BW, et al. Overexpression of pyruvate dehydrogenase kinase 3 increases drug resistance and early recurrence in colon cancer. Am J Pathol. 2011;179(3):1405–14.
Korotchkina LG, Patel MS. Site specificity of four pyruvate dehydrogenase kinase isoenzymes toward the three phosphorylation sites of human pyruvate dehydrogenase. J Biol Chem. 2001;276(40):37223–9.
Acknowledgements
We would like to thank Guo-Kuan Chen, Shanghai KangChen Bio-tech (China), for technical assistance for the PCR array experiments.
Funding
The current study was supported by the National Natural Science Foundation of China (grant no. 81260536).
Author information
Authors and Affiliations
Contributions
TZ designed the study, analyzed the data, and wrote the paper; CL, JSF, YPM, and LRC collected the specimens, performed the experiments, and collected the data. All authors have read and approved the final version to be submitted.
Corresponding author
Ethics declarations
Ethical Approval
The study was approved by the Ethics Committee of Ruikang Hospital of Guangxi Traditional Chinese Medical University (Guangxi, China). All procedures were performed according to the principles expressed in the Declaration of Helsinki.
Consent
All participants were explained their rights and signed the written informed consent before participation.
Conflict of Interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Zhang, T., Ma, Y., Fang, J. et al. A Deregulated PI3K-AKT Signaling Pathway in Patients with Colorectal Cancer. J Gastrointest Canc 50, 35–41 (2019). https://doi.org/10.1007/s12029-017-0024-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-017-0024-9