Abstract
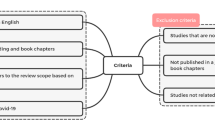
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a known virus that leads to a respiratory disease called coronavirus disease 19 (COVID-19). Natural killer (NK) cells, as members of innate immunity, possess crucial roles in restricting viral infections, including COVID-19. Their functions and development depend on receiving signals through various receptors, of which killer cell immunoglobulin-like receptors (KIRs) belong to the most effective ones. Different studies investigated the association between KIR gene content and susceptibility to COVID-19. Since previous studies have yielded contradictory results, we designed this meta-analysis study to draw comprehensive conclusions about COVID-19 risk and KIR gene association. According to PRISMA guidelines, a systematic search was performed in the electronic databases to find all studies investigating KIR gene contents in COVID-19 patients before March 2023. Any association between KIR genes and COVID-19 risk was determined by calculating pooled odds ratio (OR) and 95% confidence interval (CI). After applying the inclusion and exclusion criteria, 1673 COVID-19 patients and 1526 healthy controls from eight studies were included in this meta-analysis. As the main results, we observed a positive association between the 2DL3 (OR = 1.48, 95% CI = 1.17–1.88, P < 0.001) and susceptibility to COVID-19 and a negative association between the 2DP1 and the risk for COVID-19 (OR = 0.48, 95% CI = 0.23–0.99, P = 0.049). This meta-analysis demonstrated that KIR2DL3, as a member of iKIRs, might be associated with an increased risk of COVID-19 disease.




Similar content being viewed by others
References
Cucinotta D, Vanelli M. WHO declares COVID-19 a pandemic. Acta Biomed. 2020;91(1):157–60. https://doi.org/10.23750/abm.v91i1.9397.
Muhareb R, Giacaman R. Tracking COVID-19 responsibly. Lancet. 2020. https://doi.org/10.1016/s0140-6736(20)30693-0.
Pal M, Berhanu G, Desalegn C, Kandi V. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2): an update. Cureus. 2020;12(3):e7423. https://doi.org/10.7759/cureus.7423.
Payne S. Family Coronaviridae. Viruses. 2017;149–58. https://doi.org/10.1016/B978-0-12-803109-4.00017-9.
Kannan S, Shaik Syed Ali P, Sheeza A, Hemalatha K. COVID-19 (novel coronavirus 2019) - recent trends. Eur Rev Med Pharmacol Sci. 2020;24(4):2006–11. https://doi.org/10.26355/eurrev_202002_20378.
Lotfi R, Kalmarzi RN, Roghani SA. A review on the immune responses against novel emerging coronavirus (SARS-CoV-2). Immunol Res. 2021;69(3):213–24.
Mohammed RN, Tamjidifar R, Rahman HS, Adili A, Ghoreishizadeh S, Saeedi H, et al. A comprehensive review about immune responses and exhaustion during coronavirus disease (COVID-19). Cell Commun Signal. 2022;20(1):79. https://doi.org/10.1186/s12964-022-00856-w.
Hosseini A, Hashemi V, Shomali N, Asghari F, Gharibi T, Akbari M, et al. Innate and adaptive immune responses against coronavirus. Biomed Pharmacother. 2020;132:110859. https://doi.org/10.1016/j.biopha.2020.110859.
Agrati C, Carsetti R, Bordoni V, Sacchi A, Quintarelli C, Locatelli F, et al. The immune response as a double-edged sword: the lesson learnt during the COVID-19 pandemic. Immunology. 2022;167(3):287–302. https://doi.org/10.1111/imm.13564.
Saad N, Moussa S. Immune response to COVID-19 infection: a double-edged sword. Immunol Med. 2021;44(3):187–96. https://doi.org/10.1080/25785826.2020.1870305.
Taylor PC, Adams AC, Hufford MM, de la Torre I, Winthrop K, Gottlieb RL. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat Rev Immunol. 2021;21(6):382–93. https://doi.org/10.1038/s41577-021-00542-x.
Alhumaid S, Al Mutair A, Alali J, Al Dossary N, Albattat SH, Al HajjiMohammed SM, et al. Efficacy and safety of tixagevimab/cilgavimab to prevent COVID-19 (pre-exposure prophylaxis): a systematic review and meta-analysis. Diseases. 2022;10(4) https://doi.org/10.3390/diseases10040118.
Kyriazopoulou E, Panagopoulos P, Metallidis S, Dalekos GN, Poulakou G, Gatselis N, et al. An open label trial of anakinra to prevent respiratory failure in COVID-19. Elife. 2021:10. https://doi.org/10.7554/eLife.66125.
Vlaar APJ, de Bruin S, Busch M, Timmermans S, van Zeggeren IE, Koning R, et al. Anti-C5a antibody IFX-1 (vilobelimab) treatment versus best supportive care for patients with severe COVID-19 (PANAMO): an exploratory, open-label, phase 2 randomised controlled trial. Lancet Rheumatol. 2020;2(12):e764–e73. https://doi.org/10.1016/s2665-9913(20)30341-6.
Monk PD, Marsden RJ, Tear VJ, Brookes J, Batten TN, Mankowski M, et al. Safety and efficacy of inhaled nebulised interferon beta-1a (SNG001) for treatment of SARS-CoV-2 infection: a randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Respir Med. 2021;9(2):196–206. https://doi.org/10.1016/s2213-2600(20)30511-7.
Di Vito C, Calcaterra F, Coianiz N, Terzoli S, Voza A, Mikulak J, et al. Natural killer cells in SARS-CoV-2 infection: pathophysiology and therapeutic implications. Front Immunol. 2022;13:888248. https://doi.org/10.3389/fimmu.2022.888248.
Mikulak J, Bruni E, Oriolo F, Di Vito C, Mavilio D. Hepatic natural killer cells: organ-specific sentinels of liver immune homeostasis and physiopathology. Front Immunol. 2019;10:946. https://doi.org/10.3389/fimmu.2019.00946.
Vivier E, Tomasello E, Baratin M, Walzer T, Ugolini S. Functions of natural killer cells. Nat Immunol. 2008;9(5):503–10.
Dębska-Zielkowska J, Moszkowska G, Zieliński M, Zielińska H, Dukat-Mazurek A, Trzonkowski P, Stefańska K. KIR receptors as key regulators of nk cells activity in health and disease. Cells. 2021;10(7). https://doi.org/10.3390/cells10071777.
Sun PD. Structure and function of natural-killer-cell receptors. Immunol Res. 2003;27:539–48.
Rizzo S, Schiuma G, Beltrami S, Gentili V, Rizzo R, Bortolotti D. Role of KIR receptor in NK regulation during viral infections. Immuno. 2021;1(3):305–31.
Béziat V, Hilton HG, Norman PJ, Traherne JA. Deciphering the killer-cell immunoglobulin-like receptor system at super-resolution for natural killer and T-cell biology. Immunology. 2017;150(3):248–64.
Pende D, Falco M, Vitale M, Cantoni C, Vitale C, Munari E, et al. Killer Ig-like receptors (KIRs): their role in NK cell modulation and developments leading to their clinical exploitation. Front Immunol. 2019;10:1179. https://doi.org/10.3389/fimmu.2019.01179.
Lee HR, Baek KH. Role of natural killer cells for immunotherapy in chronic myeloid leukemia (Review). Oncol Rep. 2019;41(5):2625–35. https://doi.org/10.3892/or.2019.7059.
Cronk JM, Fafoutis E, Brown MG. Licensing natural killers for antiviral immunity. Pathogens. 2021;10(7). https://doi.org/10.3390/pathogens10070908.
Boudreau JE, Hsu KC. Natural killer cell education and the response to infection and cancer therapy: stay tuned. Trends Immunol. 2018;39(3):222–39. https://doi.org/10.1016/j.it.2017.12.001.
Rascle P, Woolley G, Jost S, Manickam C, Reeves RK. NK cell education: physiological and pathological influences. Front Immunol. 2023;14:1087155. https://doi.org/10.3389/fimmu.2023.1087155.
Paul S, Lal G. The molecular mechanism of natural killer cells function and its importance in cancer immunotherapy. Front Immunol. 2017;8:1124. https://doi.org/10.3389/fimmu.2017.01124.
Shan Z, Huang J, Liao Q, Huang K, Wang M, Xu R, et al. Association of killer cell immunoglobulin-like receptors with spontaneous clearance of hepatitis C virus in the Chinese population. Transfusion. 2018;58(4):1028–35. https://doi.org/10.1111/trf.14527.
Ligotti ME, Aiello A, Accardi G, Calabrò A, Ciaccio M, Colomba C, et al. Distribution of KIR genes and their HLA ligands in different viral infectious diseases: frequency study in Sicilian population. Int J Mol Sci. 2022;23(24):15466.
Jackson E, Zhang CX, Kiani Z, Lisovsky I, Tallon B, Del Corpo A, et al. HIV exposed seronegative (HESN) compared to HIV infected individuals have higher frequencies of telomeric killer immunoglobulin-like receptor (KIR) B motifs; contribution of KIR B motif encoded genes to NK cell responsiveness. PLoS One. 2017;12(9):e0185160. https://doi.org/10.1371/journal.pone.0185160.
Bernal E, Gimeno L, Alcaraz MJ, Quadeer AA, Moreno M, Martínez-Sánchez MV, et al. Activating killer-cell immunoglobulin-like receptors are associated with the severity of coronavirus disease 2019. J Infect Dis. 2021;224(2):229–40. https://doi.org/10.1093/infdis/jiab228.
Littera R, Chessa L, Deidda S, Angioni G, Campagna M, Lai S, et al. Natural killer-cell immunoglobulin-like receptors trigger differences in immune response to SARS-CoV-2 infection. PLoS One. 2021;16(8):e0255608. https://doi.org/10.1371/journal.pone.0255608.
Lesan V, Bewarder M, Metz C, Becker A, Mang S, Regitz E, et al. Killer immunoglobulin-like receptor 2DS5 is associated with recovery from coronavirus disease 2019. Intensive Care Med Exp. 2021;9(1):45. https://doi.org/10.1186/s40635-021-00409-4.
Hajeer A, Jawdat D, Massadeh S, Aljawini N, Abedalthagafi MS, Arabi YM, Alaamery M. Association of KIR gene polymorphisms with COVID-19 disease. Clin Immunol. 2022;234:108911. https://doi.org/10.1016/j.clim.2021.108911.
Maruthamuthu S, Rajalingam K, Kaur N, Morvan MG, Soto J, Lee N, et al. Individualized constellation of killer cell immunoglobulin-like receptors and cognate HLA class I ligands that controls natural killer cell antiviral immunity predisposes COVID-19. Front Genet. 2022;13:845474. https://doi.org/10.3389/fgene.2022.845474.
Hu S, Shao Z, Ni W, Sun P, Qiao J, Wan H, et al. The KIR2DL2/HLA-C1C1 Gene pairing is associated with an increased risk of SARS-CoV-2 infection. Front Immunol. 2022;13:919110. https://doi.org/10.3389/fimmu.2022.919110.
Alomar S, Alkhuriji A, Alkhulaifi FM, Mansour L, Al-Jurayyan A, Aldossari GS, et al. Relationship between KIR genotypes and HLA-ligands with SARS-CoV-2 infection in the Saudi population. J King Saud Univ Sci. 2023;35(1):102416. https://doi.org/10.1016/j.jksus.2022.102416.
Campbell KS, Purdy AK. Structure/function of human killer cell immunoglobulin-like receptors: lessons from polymorphisms, evolution, crystal structures and mutations. Immunology. 2011;132(3):315–25. https://doi.org/10.1111/j.1365-2567.2010.03398.x.
Long EO, Sik Kim H, Liu D, Peterson ME, Rajagopalan S. Controlling natural killer cell responses: integration of signals for activation and inhibition. Annu Rev Immunol. 2013;31:227–58.
Quatrini L, Della Chiesa M, Sivori S, Mingari MC, Pende D, Moretta L. Human NK cells, their receptors and function. Eur J Immunol. 2021;51(7):1566–79. https://doi.org/10.1002/eji.202049028.
Pegram HJ, Andrews DM, Smyth MJ, Darcy PK, Kershaw MH. Activating and inhibitory receptors of natural killer cells. Immunol Cell Biol. 2011;89(2):216–24. https://doi.org/10.1038/icb.2010.78.
Acknowledgements
We appreciate Dr. Peyman Bemani, Assistant Professor of Medical Immunology, for his valuable remarks in revising the manuscript.
Author information
Authors and Affiliations
Contributions
KK conceived and designed the study. NM and SM screened publications, extracted data, and wrote the first draft of the manuscript. SHT performed the statistical analysis and wrote part of the manuscript draft. DK, SMe, and KK reviewed and made edits to the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Teshnizi, S.H., Mirzazadeh, S., Mashhadi, N. et al. Association study between killer immunoglobulin-like receptor polymorphisms and susceptibility to COVID-19 disease: a systematic review and meta-analysis. Immunol Res 72, 175–184 (2024). https://doi.org/10.1007/s12026-023-09428-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12026-023-09428-7