Abstract
This manuscript aims to: 1) provide specific guidelines on PMM techniques in the setting of minimally invasive autopsy (MIA), both for pathologists collecting samples and for microbiologists advising pathologists and interpreting the results and 2) introduce standardization in PMM sampling at MIA. Post-mortem microbiology (PMM) is crucial to identify the causative organism in deaths due to infection. MIA including the use of post-mortem (PM) computed tomography (CT) and PM magnetic resonance imaging (MRI), is increasingly carried out as a complement or replacement for the traditional PM. In this setting, mirroring the traditional autopsy, PMM aims to: detect infectious organisms causing sudden unexpected deaths; confirm clinically suspected but unproven infection; evaluate the efficacy of antimicrobial therapy; identify emergent pathogens; and recognize medical diagnostic errors. Meaningful interpretation of PMM results requires careful evaluation in the context of the clinical history, macroscopic and microscopic findings. These guidelines were developed by a multidisciplinary team with experts in various fields of microbiology and pathology on behalf of the ESGFOR (ESCMID – European Society of Clinical Microbiology and Infectious Diseases - Study Group of Forensic and Post-mortem Microbiology, in collaboration with the ESP -European Society of Pathology-) based on a literature search and the author’s expertise. Microbiological sampling methods for MIA are presented for various scenarios: adults, children, developed and developing countries. Concordance between MIA and conventional invasive autopsy is substantial for children and adults and moderate for neonates and maternal deaths. Networking and closer collaboration among microbiologists and pathologists is vital to maximize the yield of PMM in MIA.
Similar content being viewed by others
Introduction
Post-mortem microbiology (PMM) is crucial to identify the causative organism in deaths due to infection. Although the traditional autopsy remains the gold standard procedure to obtain samples for PMM, the lack of trained human resources, and other constraints such as cultural and religious requirements, have led to the introduction of minimally invasive autopsy (MIA) techniques as an alternative [1]. In some cases, MIA includes the procurement of tissue samples, also known as Minimally Invasive Tissue Sampling (MITS), using fine needles to collect small amounts of tissue from key organs and body fluids. Currently, MIA is increasingly carried out as a complement or replacement for the traditional complete autopsy, mostly in low- and middle-income countries (LMIC).
The dead human body is a complex microbial ecological system that varies significantly between individuals as a result of diet, lifestyle and geographical factors [2, 3]. It is therefore important to highlight that a positive result from PMM may reflect a genuine infection but may also reflect sample contamination, commensal organisms and/or post-mortem bacterial translocation (PMBT). The incidence of post-mortem contamination and PMBT has been estimated to approximate 20% in traditional autopsies where PMM has been conducted [4, 5]. However, when standardized procedures are adhered to, post-mortem contamination can be kept below 10% [6].
PMBT is a natural phenomenon in which endogenous commensal gut bacteria, most commonly Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, Enterococcus spp., Clostridia, and streptococci multiply and migrate into the blood and tissues. PMBT is a driver of putrefactive decomposition, which begins approximately four minutes after death [2, 7,8,9]. Although PMBT begins within eight hours of death, it is not solely dependent on the post-mortem interval (PMI). Agonal spread of microorganisms throughout the body during the process of dying and resuscitation has been suggested but is not universally accepted [6, 10, 11]. Aerobic bacteria proliferate early during the putrefactive process, to be overtaken later by anaerobes [2].
Sample contamination during collection of peripheral blood is most commonly due to the introduction of coagulase-negative staphylococci from the skin [10]. In general, a mixed bacterial growth likely represents sample contamination whilst single isolates are generally interpreted to represent true positives [6, 11, 12]. Despite close adherence to sampling protocols, a pure growth of a single microorganism from multiple sites may still represent contamination rather than infection [4, 10]. Yet isolation of a pathogenic organism from multiple sites at autopsy most likely represents a true ante-mortem bacteremia [4].
A prolonged PMI and putrefaction are usually perceived to impair the yield of PMM [13]. Tuomisto et al. [14], using real-time quantitative PCR to investigate bacterial migration from the gut into the blood, liver, portal vein, mesenteric lymph node, and pericardial fluid of cadavers stored at 4°C, were able to demonstrate that the relative amounts of intestinal bacterial DNA (bifidobacteria, Bacteroides, Enterobacter and Clostridia) increased with time. However, a prolonged PMI does not uniformly increase the risk of PMBT or the yield in obtaining positive post-mortem cultures [6, 15]. Differences in detection techniques (routine bacteriology cultures versus molecular techniques) could be part of the explanation in contradictory study results.
Palmiere et al. [4] showed that bacterial cultures and real-time polymerase chain reaction (qPCR) analyses were reliable, not yielding false-positive results in the detection of Neisseria meningitidis, Listeria monocytogenes, Streptococcus pneumoniae or Haemophilus influenzae in CSF obtained from the lateral ventricles and cisterna magna in severely decomposing adults known to have not died infectious deaths. Positive results were obtained in both bacterial culture and qPCR analysis of CSF from decomposing controls with known meningitis. Also, reliable results were obtained in other N. meningitis qPCR validation studies performed in post-mortem samples [16].
Overall, PMBT is a frustrating “side-effect” of death. From a practical point of view, refrigeration of the body minimizes this phenomenon [6]. There is increasing interest in the possibility that exploration of successional changes in the human microbiome after death using metagenomics may have a role to play in determining the PMI [3, 8], particularly in decomposing bodies [2, 17, 18].
As PMM is not yet an extended practice in MIA, the contaminating effects of PMBT have not been evaluated yet. In the following paragraphs, we develop in more detail microbiological sampling in MIA. With careful interpretation, PMM is a useful tool to identify correctly an infectious cause of death (COD). Our study aims to provide useful guidelines on PMM sampling techniques in MIA, both to the pathologist requesting the analysis and to the microbiologist advising the pathologist and interpreting the results.
General post-mortem microbiology sampling technique
PMM samples should be collected as soon as possible and ideally within 24 hours of death [19], but it is often impossible to perform MIA in such a short time frame, especially in poor settings. The collection of samples for PMM can still be fruitful [6], however the PMI should always be considered along with the imaging and/or macroscopic and microscopic features of the organs when interpreting the results (See Table 1).
The nature of the samples to be collected varies depending on the clinical history. Specific examples are considered below. In infant and childhood death without symptoms, a standard set of samples is recommended in the traditional PM: nasopharyngeal swab, CSF, blood, lung, spleen, heart, and bowel content. If sepsis, bacteremia or other conditions are suspected, it is recommended to add samples from other tissues (Table 1). [19,20,21]. According to the method of MIA used, the aim would be to replicate these samples as much as possible. Some centers complement the traditional full autopsy or the MIA MSCT (Multi-Slice-Computed Tomography), MRI (Magnetic Resonance Imaging), using MITS (tissue sampling through fine needle) or laparoscopic/thoracoscopic approaches which allow direct visualization and sampling of organs [22, 23].
A variety of other conditions may cause systemic inflammatory response syndrome and mimic sepsis in the absence of infection [24]. According to the context of the MIA, the provision of non-microbiological samples, such as those referred for toxicological screening, metabolic analysis and histopathological examination may be required.
Where samples are obtained by percutaneous needle puncture/aspiration in MITS, the body must first be appropriately positioned, and the skin should be cleaned with sterile water and disinfected either with alcohol-based solution containing chlorhexidine or iodine. Isopropyl alcohol should only be used after samples for toxicological analysis have been obtained (or the toxicologist must be notified of its use) [19]. Prior post-mortem angiography via femoral cannulation does not impede the collection of peripheral blood for PMM provided that the skin is disinfected prior to cannulation for angiography [5].
Ideally, a separate set of sterile instruments should be used to collect each sample, in order to avoid cross-contamination [13].
Tissue samples for microbiology should be stored in sterile containers or bottles without additives, refrigerated until transport and sent to the microbiology laboratory within 24-48 hours after the autopsy [13, 18]. It is recommended to cryopreserve additional samples at -80ºC for future molecular analyses in the event they are required.
Consideration must be given to the health and safety of mortuary workers during the collection of PMM samples, as the decedent’s microbiome may pose a risk. Specific measures to protect mortuary staff from potential infectious hazards have been described elsewhere [13, 25]. Direct contact and puncture wounds are the most important routes for transmission of blood-borne infections, and aerosol transmission is mainly a risk for airborne pathogens such as Mycobacterium tuberculosis complex or the MERS, SARS or COVID-19 coronaviruses [26]. Universal safety precautions to be applied are personal protective equipment including eye protection, a surgical mask, surgical gown, waterproof apron, gauntlets, surgical gloves and, optionally, a cut-resistant glove on the non-dominant hand. However, when airborne pathogens are anticipated, additional respiratory protection, such as a free flight phase two (FFP2) or even FFP3 masks for highly contagious pathogens such as COVID-19 is required [27]. Where fungal growth is evident macroscopically either during the crime scene review, exhumation or at autopsy, respiratory protection should also be worn due to the risk of inhaling spores from Aspergillus fumigatus complex and Candida albicans [18].
Different settings for microbiology in MIA
The adult MIA
Changing societal, cultural, religious and political attitudes around the world have seen the conventional invasive autopsy become less acceptable and less frequent [1, 28, 29]. MIA has gained acceptability and is a valid alternative to the traditional autopsy in the investigation of natural adult deaths, especially in low-income countries [28, 30,31,32,33].
As in the conventional PM, MIA protocol includes a review of the available history of events leading to death and a thorough external examination. Unlike the conventional complete autopsy, in MIA the body cavities are not opened. Instead, key organs may be sampled through percutaneous puncture using specific needles (see below under MIA in developing countries). Swabs represent another alternative to be used in noninvasive post mortems. Where facilities and resources permit, non-invasive or MIAs using either multi-slice computed tomography (MSCT) or MRI scans with or without angiography may be used to replace or limit the conventional autopsy [31, 34, 35]. In adults, MSCT has been shown to be superior to MRI in detecting the COD [34]. The combination of post-mortem whole body MSCT scan, CT angiography and tissue biopsy has been shown to be the most sensitive technique in detecting the COD when MIAs are undertaken and surpass non-invasive techniques alone [35].
Microbiology sampling during the adult MIA
While MIA can reduce the need to undertake invasive autopsies in patients with known or suspected Hazard Group 3 infections such as HIV or COVID-19 [31, 35,36,37], very few recent articles detail the collection of samples for microbiological examination in adults during MIA. Previous studies have focused on the collection of biopsies for histopathological examination [38, 39]. In some studies, patients with known high-risk infections have been actively excluded and/or tissue sampling limited to histopathological examination [30, 40]. Previous research has focused on the role of MIA to assess the prevalence of tuberculosis and other infections in patients with known HIV infection [41]. Collection of blood, CSF and samples from the lungs, liver and at MIA conducted 3-6 days after death have demonstrated that tuberculosis and other bacterial infections are the commonest COD in patients with advanced HIV infection. Such findings concur with those from conventional invasive autopsies performed in other studies [42].
Swab samples, either nasopharyngeal or oropharyngeal, for SARS-CoV-2 can be taken without damage to the body, before performing the MIA protocol [43]. Swabbing the upper respiratory tract, particularly the nasopharyngeal site, is relevant for post-mortem diagnosis of viral respiratory pathogens [44] while the throat is relevant for measles [45].
Prior to procuring needle tissue samples, the skin should be cleaned with sterile water and disinfected either with alcohol-based solution containing chlorhexidine or iodine. Under the suspicion of legionellosis, it is particularly important to avoid contact with tap water to prevent contamination [41]. Blood samples are collected into an EDTA-containing tube, and an aerobic blood culture bottle. CSF is collected into a sterile tube for analysis and stored at -80°C in Eppendorf Safe-Lock Tubes. Tissue samples obtained by needle biopsy are placed both in thioglycolate broth and in lysis buffer for microbiological analyses. Similarly, samples from these sites and from the heart, spleen, kidney, uterus and skin can be collected for histopathological examination. MIA is ideally performed within 24 hours of death to minimize the effects of PMBT. MIAs performed after 24h after death can still render reliable diagnostic results, although in this situation the contribution of some microorganisms such as Enterobacteriaceae and Pseudomonas spp. might be overestimated [46] and may reflect bacterial translocation rather than true infections. This translocation may become a larger problem given the delay in performing MIA while waiting for the result of a SARS-CoV-2 PCR taken after admission of a body in the forensic pathology laboratory.
Castillo et al. [32] compared MIA to conventional invasive autopsy. Within 24 hours of death, the authors disinfected and sampled the skin, blood and CSF. This was followed by targeted collection of tissue using needle biopsies for microbiological and histopathological analysis from the liver, lungs, bone marrow and central nervous system [47]. The conventional autopsy was performed immediately after MIA. MIA sensitivity was highest for disseminated infections (98%, 95% Confidence Interval (CI) 87-100%) and lowest for pulmonary infections (79%, 95% CI: 58-93%) due to difficulties in targeting lesions within the lung on blind needle biopsy. MIA specificity was highest for gastrointestinal infections (100%, 95% CI 97-100%) and lowest for disseminated infections (99%, 95% CI 92-100%). The single most frequent infectious COD identified in this study was M. tuberculosis complex. [32, 48].
In an innovative study, Van der Linden et al. [49] used needle biopsies under imaging guidance during MIA to collect samples from the heart, liver and kidney. The authors found that the integrity of the patient’s RNA in these samples was larger than in samples subsequently obtained by conventional autopsy, though glyceraldehyde-3-phosphate dehydrogenase (GAPDH) expression was lower in samples with a PMI greater than 15 hours. Needle biopsies had also been used during MIA with CT-guidance to perform PCR analysis for the presence of bacterioplankton DNA in lungs, spleen, kidney and brain [50] as a diagnostic marker in drowning six days after the discovery of the body. Each samples wascollected using a new sterile cutting needle after cleaning the skin with alcohol and was stored at 4°C in sterile containers without preservative until processing.
Protocol for the collection of microbiology samples at MIA in adults
At present, the literature reflects that there is insufficient worldwide experience to allow a definitive recommendation on how and when samples should be collected for microbiological investigation during MIA. Relatively few studies have addressed this issue, and almost all have been undertaken in areas of high HIV and tuberculosis prevalence [47, 51]. Microbiological investigation during adult MIA is likely to have a lower diagnostic yield with longer PMIs and in areas of low HIV prevalence. It is clear that samples can be successfully obtained for microbiology at MIA with or without imaging-guidance. Provided these have been obtained after first appropriately disinfecting the skin, these samples can be submitted to a wide variety of microbiological investigations, including serology, culture and PCR-based analyses (Table 2). What remains unclear is how many biopsies should be obtained from each site, whether all samples should be collected in every MIA or whether sample collection should be directed by the clinical history and what impact the PMI has on the sensitivity and specificity of such investigations. These are areas for future research.
For several BSL3 pathogens, such as HIV, M. tuberculosis complex and SARS-CoV-2, it is difficult to determine the duration of infectiousness of the body. Tuberculosis could be infectious for more than 12 months [52], HIV and SARS-CoV-2 may be detected by PCR for several days post-mortem [53, 54]. To the best of our knowledge the duration of their infectiousness, i.e. isolation in viral culture in dead human bodies, is not known. Therefore, it is worthwhile taking swabs or punctures for molecular analyses for these BSL3 pathogens, following stringent safety measures [26] even with post-mortem intervals of more than 24 h.
The MIA in stillbirth, neonates and children
Microbiology sampling in pediatric MIA can be obtained using the protocol described above for adult patients (Table 3) [47, 51, 55, 56]. Successful implementation of pediatric MIA into routine clinical practice requires further development and has many issues to consider in order to ensure that the best possible service is provided [22, 23]. However, provided that MIA is undertaken jointly by pediatric pathologists and radiologists, it could be an acceptable alternative to conventional autopsy in selected cases [50]. Currently, the pediatric MIA uses MRI rather than CT [57, 58]. The examination of the placenta becomes an invaluable tool in the post-mortem examination of stillbirth, and in neonatal deaths the study of the placenta becomes key, providing significant information in more than 50% of the MIA cases [59].
In low/medium income countries, where resources are not available, some studies suggest using minimally invasive tissue sampling (MITS) with available clinical data, for attributing underlying and immediate causes of neonatal deaths [60].
In pediatric post-mortems swab samples for SARS-CoV-2 a rectal swab should be added in addition to the nasopharyngeal swab used in adults [61].
MIA in developing countries
Most of the world's population dies without qualified medical personnel either certifying their death or identifying a specific cause. The conventional invasive autopsy is indisputably the gold standard for COD determination. However, it is rarely performed in low-income countries due to the lack of resources, trained available pathologists, poor acceptability and the fact that in these countries most of the deaths occur outside the hospital. Minimally invasive techniques have been proposed as an alternative to the traditional complete autopsy to better refine the mortality statistics in these settings and therefore contribute to effective public health interventions.
In 2013, the Barcelona Institute for Global Health (ISGlobal) started the CaDMIA research project (validation of the MIA tool for COD investigation in developing countries), funded by the Bill & Melinda Gates Foundation, aiming to validate the use of a less invasive methodology for COD ascertainment. The MIA protocol is based on the collection of post-mortem biopsies from key organs (central nervous system, lungs, heart, liver, kidney, bone marrow, spleen and skin) and fluids (CSF and blood). MIA and conventional invasive autopsy samples are studied using traditional and advanced histopathological and microbiological techniques. Pathology and microbiology results are analyzed by a multidisciplinary team in order to reach a consensus regarding the most plausible COD, both for the MIA and the conventional autopsy. The procedure involves the collection of 20mL of blood and CSF and puncture of liver, lungs, heart, spleen, kidneys, bone marrow and brain in all cases plus the uterus in women of childbearing age, using biopsy needles [22, 55, 56]. The type and main characteristics of the different needles used in the MIA procedure for each particular biopsy, and the sites of puncture and the number of samples to be obtained are summarized in Table 4 [47]. The organs are presented in the order in which the samples are collected.
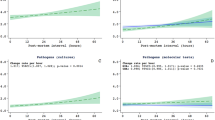
Extensive microbiological investigations are included to identify infectious COD [51]. The sampling and testing scheme for microbiology includes a universal screening for several key pathogens (i.e. HIV, HBV, malaria) and bacterial/fungal culture of biological fluids and tissue samples. In addition to universal screening, frozen aliquots of plasma, CSF or other biological fluids and tissue samples in nucleic acid-preserving lysis buffer are collected for molecular assays (i.e. broad-spectrum PCR assays and specific qPCR tests) guided by histopathological findings. Preliminary results suggest that CSF and lung tissue are among the samples offering a better performance for microbiological analysis to detect an infectious COD. The MIA tool has recently been validated for neonates, children, maternal deaths and other adult deaths from Mozambique [32, 55, 56, 62]. In these validation studies, the concordance between MIA and conventional complete autopsy is moderate for neonates and maternal deaths (kappa = 0.404 and 0.485 respectively) and substantial for children and adults (kappa = 0.704 and 0.732 respectively). The overall concordance between the MIA diagnosis and conventional invasive autopsy was 75.9% (85/112). The concordance was higher for infectious diseases and malignant tumors (63/80 (78.8%) and 13/16 (81.3%), respectively) than for other diseases (9/16; 56.2%). The specific microorganisms causing death were identified in the MIA in 62/74 (83.8%) of the infectious disease deaths with a recognized cause.
Discussion
PMM has shown to be useful to detect an infectious COD, though it must be interpreted with caution, and in the context of the clinical history (including ante-mortem microbiology) as well as imaging, macroscopic and microscopic findings [9, 10]. Where there is concordance between PMM, the history and the autopsy findings, it is reasonable to attribute the identified microorganism as a causative factor. Where concordance is lacking, it is likely that the microbiological findings are spurious and represent contamination or PMBT [4].
Laboratory interpretative criteria for post-mortem bacterial culturing have been previously defined by many authors [11, 12, 63,64,65,66,67,68]. The possibility of sample degradation affecting the detection of nucleic acids and therefore compromising the molecular detection of pathogens should also be considered when dealing with post-mortem samples [65]. However, these recommendations are generally addressed to the interpretation of the results obtained in samples taken during a traditional autopsy.
The PMM protocols using MIA offer an alternative to better ascertain infection under specific settings. As mentioned above, some validation studies have already been performed, showing their value [32, 55, 56, 62]. However, additional investigations in a variety of scenarios and the evaluation of how the PMI affects MIA microbiological results compared to those obtained by a traditional autopsy could be of help for a better estimation of its possible utility under different circumstances. Also, in the MIA setting, more interpretation criteria based on prospective studies including not only post-mortem cultures, but also molecular analyses are required for a better comprehension of PMM.
Key Points
-
1
This study provides specific guidelines on post mortem microbiology (PMM) techniques in the setting of minimally invasive autopsy (MIA)
-
2
In low and middle -income countries MIA combines clinical history, external examination, minimally invasive tissue sampling (MITS) and microbiology.
-
3
In developed countries MIA adds to multi slice computed tomography (MSCT) and/or magnetic resonance imaging (MRI).
-
4
Swabbing is an additional approach to be considered as part of MIA, as it is useful for SARS-CoV-2 and other respiratory viruses.
-
5
Where there is concordance between PMM, the history, histology and/or imaging findings, it is reasonable to attribute the identified microorganism as a causative factor. Where concordance is lacking, it is likely that the microbiological findings are spurious and represent contamination or post-mortem bacterial translocation.
References
Bassat Q, Ordi J, Vila J, Ismail MR, Carrilho C, et al. Development of a post-mortem procedure to reduce the uncertainty regarding causes of death in developing countries. Lancet. 2013;1:e125-6.
Finley SJ, Benbow ME, Javan GT. Microbial communities associated with human decomposition and their potential use as postmortem clocks. Int J Legal Med. 2015;129:623–32.
Hauther KA, Cobaugh KL, Meadows Jantz L, Sparer TE, DeBruyn JM. Estimating time since death from postmortem human gut microbial communities. J Forensic Sci. 2015;60:1234–40.
Palmiere C, Vanhaebost J, Ventura F, Bonsignore A, Bonetti LR. Cerebrospinal fluid PCR analysis and biochemistry in bodies with severe decomposition. J Forensic Leg Med. 2015;30:21–4.
Palmiere C, Egger C, Grabherr S, Jaton-Ogay K, Greub G. Postmortem angiography using femoral cannulation and postmortem microbiology. Int J Legal Med. 2015;129:861–7.
Morris JA, Harrison LM, Partridge SM. Practical and theoretical aspects of postmortem bacteriology. Curr Diagn Pathol. 2007;13:65–74.
Buchelo SR, Lynne AM. The microbiome of human decomposition. Microbe Magazine. 2016;11:165–71.
Burcham ZM, Hood JA, Pechal JL, Krausz KL, Schmidt CJ, et al. Fluorescently labeled bacteria provide insight on post-mortem microbial transmigration. Forensic Sci Int. 2016;264:63–9.
Christoffersen S. The importance of microbiological testing for establishing cause of death in 42 forensic autopsies. Forensic Sci Int. 2015;250:27–32.
Palmiere C, Egger C, Prod’Hom G, Greub G. Bacterial translocation and sample contamination in postmortem microbiological analyses. J Forensic Sci. 2016;61:367-74.
Weber MA, Sebire NJ. Postmortem investigation of sudden unexpected death in infancy: current issues and autopsy protocol. Diagn Histopathol. 2009;15:510–23.
Lobmaier IV, Vege A, Gaustad P, Rognum TO. Bacteriological investigation-significance of time lapse after death. Eur J Clin Microbiol Infect Dis. 2009;28:1191–8.
Saegeman V, Cohen MC, Alberola J, Ziyade N, Farina C, et al. How is post-mortem microbiology appraised by pathologists? Results from a practice survey conducted by ESGFOR. Eur J Clin Microbiol Infect Dis. 2017;36:1381–5.
Tuomisto S, Karhunen PJ, Vuento R, Aittoniemi J, Pessi T. Evaluation of postmortem bacterial migration using culturing and real-time quantitative PCR. J Forensic Sci. 2013;58:910–6.
Weber MA, Hartley JC, Brooke I, Lock PE, Klein NJ, et al. Post-mortem interval and bacteriological culture yield in sudden unexpected death in infancy (SUDI). Forensic Sci Int. 2010;198:121–5.
Fernández-Rodríguez A, Alcalá B, Alvarez-Lafuente R. Real-time PCR detection of Neisseria meningitidis in formalin-fixed tissues from sudden deaths. Diagn Microbiol Infect Dis. 2008;60:339–46.
Damann FE, Williams DE, Layton AC. Potential use of bacterial community succession in decaying human bone for estimating postmortem interval. J Forensic Sci. 2015;60:844–50.
Schwarz P, Dannaoui E, Gehl A, Felske-Zech H, Birngruber CG, et al. Molecular identification of fungi found on decomposed human bodies in forensic autopsy cases. Int J Legal Med. 2015;129:785–91.
Fernández-Rodríguez A, Cohen MC, Lucena J, Van de Voorde W, Angelini A, et al. How to optimise the yield of forensic and clinical post-mortem microbiology with an adequate sampling: a proposal for standardisation. Eur J Clin Microbiol Infect Dis. 2015;34:1045–57.
Fernández-Rodríguez A, Burton J, Andreoletti L, Alberola J, Fornes P, et al. Post-mortem microbiology in Sudden Death: Sampling protocols proposed in different clinical settings. Clin Microb Infect. 2019;25:570–9.
Burton JL, Saegeman V, Arribi A, Rello J, Andreoletti L, et al. Post-mortem microbiology sampling following death in hospital: An ESGFOR Task Force Consensus Statement. J Clin Pathol. 2019. https://doi.org/10.1136/jclinpath.
Addison S, Arthurs OJ, Thayyil S. Post-mortem MRI as an alternative to non-forensic autopsy in foetuses and children: from research into clinical practice. Br J Radiol. 2014;87:20130621.
Cohen MC, Paley MN, Griffiths PD, Whitby EH. Less invasive autopsy: benefits and limitations of the use of magnetic resonance imaging in the perinatal postmortem. Pediatr Dev Pathol. 2008;11:1–9.
Tsokos M. Postmortem diagnosis of sepsis. Forensic Sci Int. 2007;165:155–64.
Burton JL. Health and safety at autopsy. J Clin Pathol. 2003;56:254–60.
Hanley B, Lucas SB, Youd E, et al. Autopsy in suspected COVID-19 cases. J Clin Pathol. 2020;73:239–42.
The Royal College of Pathologists and the Association for Anatomical Pathology Technology in conjunction with Public Health England. Transmission-based precautions. Guidance for care of deceased. during COVID-19 pandemic. Issued 19 March 2020. https://www.rcpath.org/uploads/assets/0b7d77fa-b385-4c60-b47dde930477494b/G200-TBPs-Guidance-for-care-of-deceased-during-COVID-19-pandemic.pdf.
Burton JL, Underwood JCE. Clinical, educational and epidemiological value of autopsy. Lancet. 2007;369:1471–80.
Fan JK, Tong DK, Poon JT, Lo OS, Beh PS, et al. Multimodality minimally invasive autopsy – a feasible and accurate approach to post-mortem examination. Forensic Sci Int. 2010;195:93–8.
Weustinik AC, Hunink MGM, van Dijke CF, Renken CF, Krestin GP, et al. Minimally invasive autopsy: an alternative to conventional autopsy? Radiology. 2009;250:897–904.
Roberts ISD, Traill ZC. Minimally invasive autopsy employing post-mortem CT and targeted coronary angiography: evaluation of its application to a routine coronial service. Histopathology. 2014;64:211–7.
Castillo P, Martínez MJ, Ussene E, Jordao D, Lovane L, et al. Validity of a minimally invasive autopsy for cause of death determination in adults in Mozambique: an observational study. PLoS Med. 2016;13:e1002171.
Maixenchs M, Anselma R, Zielinski-Gutiérrez E, Odiambo FO, Akello C, et al. Willingness to know the cause of death and hypothetical acceptability of the minimally invasive autopsy in six diverse African and Asian settings: a mixed methods socio-behavioural study. PLoS Med. 2016;13:e1002172.
Roberts ISD, Benamore RE, Benbow EW, Lee SH, Harris JN, et al. Post-mortem imaging as an alternative to autopsy in the diagnosis of adult deaths: a validation study. Lancet. 2012;379:136–42.
Blokker BM, Wagensveld IM, Weustink AC, Oosterhuis JW, Hunink MG. Non-invasive or minimally invasive autopsy compared to conventional autopsy of suspected natural deaths in adults: a systematic review. Eur Radiol. 2016;26:1159–79.
Fryer EP, Traill ZC, Benamore RE, Roberts IS. High risk medicolegal autopsies: is a full postmortem examination necessary? J Clin Pathol. 2012;66:1–7.
Gonzalez-Arnay E, Martin-Olivera R, Quintero-Quintero YC, Hernandez-Guerra AI. Proposal for a harmonized protocol for COVID-19 screening and necropsy in forensic sciences facilities. J Forensic Leg Med. 2020;76:102067.
Bolliger SA, Filograna L, Spendlove D, Thali MJ, Sirnhofer S, et al. Postmortem imaging-guided biopsy as an adjuvant to minimally invasive autopsy with CT and postmortem angiography: a feasibility study. Am J Roentgenol. 2010;195:1051–6.
Ebert LC, Ptacek W, Breitbeck R, Fürst M, Kronreif G, et al. Virtobot 2.0: the future of automated surface documentation and CT-guided needle placement in forensic medicine. Forensic Sci Med Pathol. 2014;10:179-86.
Ross SG, Thali MJ, Bollinger S, Germerott T, Ruder TD, et al. Sudden death after chest pain: feasibility of virtual autopsy with post-mortem CT angiography and biopsy. Radiology. 2012;264:250–9.
Karat AS, Omar T, von Gottberg A, Tlali M, Chihota VN, et al. Autopsy prevalence of tuberculosis and other potentially treatable infections among adults with advanced HIV enrolled in out-patient care in South Africa. PLOS One. 2016;11:e0166158.
Cox JA, Lukande RL, Lucas S, Nelson AM, Van Marck E, et al. Autopsy causes of death in HIV-positive individuals in sub-Saharan Africa and correlation with clinical diagnoses. AIDS Rev. 2010;12:183–94.
Santurro A, Scopetti M, D’Errico S, Fineschi V. A technical report from the Italian SARS-CoV-2 outbreak. Postmortem sampling and autopsy investigation in cases of suspected or probable COVID-19. Forensic Sci Med Pathol. 2020. https://doi.org/10.1007/s12024-020-00258-9.
Moore C, Jones R. The use of coroner’s autopsy reports to validate the use of targeted swabbing rather than tissue collection for rapid confirmation of virological causes of sudden death in the community. J Clin Virol. 2015;63:59–62.
Wise J. Largest group of children affected by measles outbreak in Wales is 10–18 year olds. BMJ. 2013;346:f2545.
Hurtado JC, Quintó L, Castillo P, Carrilho C, Fernandes F, et al. Postmortem interval and diagnostic performance of the autopsy methods. Sci Rep. 2018;8:16112.
Castillo P, Ussene E, Ismail MR, Jordao D, lovane L, et al. Pathological methods applied to the investigation of causes of death in developing countries: minimally invasive autopsy approach. PLoS ONE. 2015;10:e0132057.
Palhares AEM, Ferreira L, Freire M, Castillo P, Martínez MJ, et al. Performance of the minimally invasive autopsy tool for cause of death determination in adult deaths from the Brazilian Amazon: an observational study. Virchows Arch. 2019;475:649–58.
Van der Linden A, Blokker BM, Kap M, Weustink AC, Riegman PHJ, et al. Post-mortem tissue biopsies obtained at minimally invasive autopsy: an RNA-quality analysis. PLOS One. 2014;9:e115675.
Rutty GN, Johnson C, Amoroso J, Robinson C, Bradley CJ, et al. Post-mortem computed tomography coaxial cutting needle biopsy to facilitate the detection of bacterioplankton using PCR probes as a diagnostic indicator for drowning. Int J Legal Med. 2017;131:211–6.
Martínez MJ, Massora S, Mandomando I, Ussene E, Jordao D, et al. Infectious cause of death determination using minimally invasive autopsies in developing countries. Diagn Microbiol Infect Dis. 2016;84:80–6.
Smith CR. Survival of tubercle bacilli: The viability of dried tubercle bacilli in unfiltered roomlight, in the dark and in the refrigerator. Am Rev Tuberculosis. 1942;45:334–45.
Hoffman PN, Healing TD. Zz Guide to infection control in the healthcare setting, The infection hazards of human cadavers. International Society of Infectious Diseases. https://www.ialm.info/UserFiles/files/INFO%20COVID19/ISID_GUIDE_THE_INFECTION_HAZARDS_OF_HUMAN_CADAVERS.pdf.
Luo L, Liu D, Zhang H, et al. Air and surface contamination in non-health care settings among 641 environmental specimens of 39 COVID-19 cases. PLoS Negl Trop Dis. 2020;9(14):e0008570.
Bassat Q, Castillo P, MartÍnez MJ, Jordao D, Lovane L, Hurtado JC, et al. Validity of a minimally invasive autopsy tool for cause of death determination in pediatric deaths in Mozambique: An observational study. PLoS Med. 2017;14:e1002317.
Menendez C, Castillo P, Martínez MJ, Jordao D, Lovane L, Ismail MR, et al. Validity of a minimally invasive autopsy for cause of death determination in stillborn babies and neonates in Mozambique: An observational study. PLoS Med. 2017;14:e1002318.
Thayyil S, Sebire NJ, Chitty LS, Wade A, Chong W, et al. Post-mortem MRI versus conventional autopsy in fetuses and children: a prospective validation study. Lancet. 2013;382:223–33.
Sethuraman C, Cohen M, Whitby E, Marven S, Lindley R. Sheffield minimally invasive autopsy research technique (smart)- a technical report on feasibility of keyhole autopsy and magnetic resonance imaging. Platform presentation at the 50th Meeting of the Pediatric Pathology Society. Boston, US. 2015. https://doi.org/10.2350/15-02-1606-MISC.1.
Rummery R, DiGiovanni J, Raghavan A, Whitby E, Cohen M. Benefits and limitations of the minimally invasive perinatal autopsy: A review of the first 5 years of an innovative service development. Society of Pediatric Pathology Spring Meeting. Vancouver, BC, Canada; 2018.
Madhi SA, Pathirana J, Baillie V, Izu A, Bassat Q, et al. Unraveling specific causes of neonatal mortality using minimally invasive tissue sampling: An observational study. Clin Infec Dis. 2019;69:S351-60.
Xu Y, Li X, Zhu B, et al. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat Med. 2020;26:502–5.
Castillo P, Hurtado JC, Martínez MJ, Jordao D, Lovane L, et al. Validity of a minimally invasive autopsy for cause of death determination in maternal deaths in Mozambique: an observational study. PLOS Medicine. 2017;14:e1002431.
Prtak L, Al-Adnani M, Fenton P, Kudesia G, Cohen MC. Contribution of bacteriology and virology in sudden unexpected death in infancy. Arch Dis Child. 2010;95:371–6.
Roberts FJ. A review of postmortem bacteriological cultures. Can Med Assoc J. 1969;100:70–4.
Alberola J, Cohen MC, Fernandez Rodriguez A. Análisis microbiológico post mortem. Recomendaciones de la Sociedad Española de Enfermedades Infecciosas y Microbiología Clínica. In: Cercenado E, Canton R, editors. Proc Microbio Clin. 2012. https://seimc.org/contenidos/documentoscientificos/procedimientosmicrobiologia/seimc-procedimientomicrobiologia43.pdf.
Sunagawa K, Sugitani M. Post-mortem detection of bacteremia using pairs of blood culture samples. Leg Med. 2017;24:92–7.
Weber M, Klein N, Hartley J, Lock P, Malone M, et al. Infection and sudden unexpected death in infancy: a systematic retrospective case review. Lancet. 2008;371:1848–53.
Fernández-Rodríguez A, Alberola J, Cohen MC. Análisis microbiológico post-mortem. Enferm Infecc Microbiol Clin. 2013;31:685–91.
Zhao W, Zhong Z, Xie X, Yu Q, Liu J, et al. Relation between chest CT findings and clinical conditions of coronavirus disease (COVID-19) pneumonia: a multicenter study. Am J Roentgenol. 2020;214:1072–7.
Yao XH, Li TY, He ZC, Ping YF, Liu HW, et al. A pathological report of three COVID-19 cases by minimally invasive autopsies. Zhonghua Bing Xue Za Zhi. 2020;49:411–7.
Driggin E, Madhavan MV, Bikdeli B, Chuich T, Laracy J, et al. Cardiovascular considerations for patients, health care workers, and health systems during the coronavirus disease 2019 (COVID-19) pandemic. J Am College Cardiol. 2020. https://doi.org/10.1016/j.jacc.2020.03.031.
Dubey SP, Larawin V, Molumi CP. Intracranial spread of chronic middle ear suppuration. Am J Otolaryngol. 2010;31:73–7.
Geyik MF, Kokoglu OF, Hosoglu S, Ayaz C. Acute bacterial meningitis as a complication of otitis media and related mortality factors. Yonsei Med J. 2002;43:573–8.
Saegeman V, Bammens B, De Munter P, Neyt J, Cossey V, et al. How to isolate a patient with Cladophialophora bantiana infection? An opinion J Hosp Infect. 2015;91:89–90.
Johnson JA, Boyce TG, Cetta F, Steckelberg JM, Johnson JN. Infective endocarditis in the pediatric patient: A 60-year single-institution review. Mayo Clin Proc. 2012;87:629–35.
Achermann Y, Rossle M, Hoffmann M, Deggim V, Kister S, et al. Prosthetic valve endocarditis and bloodstream infection due to Mycobacterium chimaera. J Clin Microbiol. 2013;51:1769–73.
Williamson D, Howden B, Stinear T. Mycobacterium chimaera spread from heating and cooling units in heart surgery. N Engl J Med. 2017;376:600–2.
Satoh F, Tachibana H, Hasegawa I, Osawa M. Sudden death caused by chronic Chagas disease in a non-endemic country: Autopsy report. Pathol Int. 2010;60:235–40.
Liu A, Hu Y, Coates A. Sudden cardiac death and tuberculosis - How much do we know? Tuberculosis. 2012;92:307–31.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Saegeman, V., Cohen, M.C., Burton, J.L. et al. Microbiology in minimally invasive autopsy: best techniques to detect infection. ESGFOR (ESCMID study group of forensic and post-mortem microbiology) guidelines. Forensic Sci Med Pathol 17, 87–100 (2021). https://doi.org/10.1007/s12024-020-00337-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12024-020-00337-x