Abstract
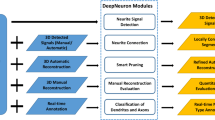
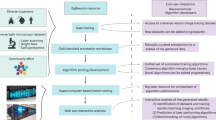
Tracing of neuron paths is important in neuroscience. Recent studies have shown that it is possible to segment and reconstruct three-dimensional morphology of axons and dendrites using fully automatic neuron tracing methods. A specific tracer may be better than others for a specific dataset, but another tracer could perform better for some other datasets. Ensemble of learners is an effective way to improve learning accuracy in machine learning. We developed automatic ensemble neuron tracers, which consistently perform well on 57 datasets of 5 species collected from 7 laboratories worldwide. Quantitative evaluation based on the data generated by human annotators shows that the proposed ensemble tracers are valuable for 3D neuron tracing and can be widely applied to different datasets.








Similar content being viewed by others
References
Abramoff, M. D., et al. (2004). Image processing with ImageJ. Biophotonics International, 11, 36–42.
Breiman, L. (1996). Bagging predictors. International Journal Machine Learning, 24, 134140.
Brown, K. M., et al. (2005). A cross-platform freeware tool for digital reconstruction of neuronal arborizations from image stacks. Neuroinformatics, 3, 343–359.
Chiang, et al. (2011). Three-dimensional reconstruction of brain-wide wiring networks in Drosophila at single-cell resolution. Current Biology, 21, 1–11.
Chklovskii, D. B. (2004). Synaptic connectivity and neuronal morphology: two sides of the same coin. Neuron, 43, 609–617.
Choromanska, A., et al. (2012). Automatic reconstruction of neural morphologies with multi-scale tracking. Neural Circuit, 6. doi:10.3389/fncir.2012.00025.
Collins, T. J. (2007). ImageJ for microscopy. Biotechniques, 43, 25–30.
Feng, L., et al. (2014). neuTube 1.0: A New Design for Efficient Neuron Reconstruction Software Based on The SWC Format eNeuro, 10.1523/ENEURO.0049-14.
Fiala, J. C. (2005). Reconstruct: A free editor for serial section microscopy. Journal of Microscopy, 218, 52–61.
Freund, Y., & Schapire, R. (1999). A Short Introduction to Boosting. Journal of Japanese Society for Artificial Intelligence, 14(5), 771– 780.
Kalisman, N., et al. (2003). Deriving physical connectivity from neuronal morphology. Biological Cybernetics, 88, 210–218.
Krichmar, J. L., et al. (2002). Effects of dendritic morphology on CA3 pyramidal cell electrophysiology: a simulation study. Brain Research, 941, 11–28.
Kim, E. J., et al. (2016). Neuroscience: Optimized tracing of neural circuits. Nature Methods, 13, 470.
Li, A., et al. (2010). Micro-optical sectioning tomography to obtain a high-resolution atlas of the mouse brain. Science, 330, 1404– 1408.
Lu, J. (2009). Semi-automated reconstruction of neural processes from large numbers of fluorescence images. PLoS One, 4, e5655.
Mainen, Z. F., & Sejnowski, T. J. (1996). Influence of dendritic structure on firing pattern in model neocortical neurons. Nature, 382, 363–366.
Meijering, E. (2010). Neuron Tracing in Perspective. Cytometry, 77, 693–704.
Ming, X., Li, A., Wu, J., Yan, C., Ding, W., Gong, H., Zeng, S., & Liu, Q. (2013). Rapid reconstruction of 3D neuronal morphology from light microscopy images with augmented rayburst sampling. PloS one, 8, e84557.
Peng, H., Ruan, R., Long, F., Simpson, J. H., & Myers, E.W. (2010). V3D enables real-time 3D visualization and quantitative analysis of large-scale biological image data sets. Nature Biotechnology, 28, 348–353.
Peng, H., Ruan, Z., Atasoy, D., & Sternson, S. (2010). Automatic reconstruction of 3D neuron structures using a graph-augmented deformable model. Bioinformatics, 26, i38–i46.
Peng, H., Long, F., & Myers, G. (2011). Automatics 3D neuron tracing using all-path pruning. Bioinformatics, 27, i239–i247.
Peng, H., et al. (2015). BigNeuron: large-scale 3D neuron reconstruction from optical microscopy images. Neuron, 87, 252–256.
Polikar, R. (2006). Ensemble based systems in decision making. IEEE Circuits and Systems Magazine, 6(3), 21–45.
Rokach, L. (2010). Ensemble-based classifiers. Artificial Intelligence Review, 33, 1V394.
Schaefer, A. T., et al. (2003). Coincidence detection in pyramidal neurons is tuned by their dendritic branching pattern. Journal of Neurophysiology, 89, 3143–3154.
Sholl, D. A. (1956). The measurable parameters of the cerebral cortex and their significance in its organization. Progress in Neurobiology, 2, 324–33.
Spruston, N. (2008). Pyramidal neurons: dendritic structure and synaptic integration. Nature Reviews Neuroscience, 4, 206–221.
Viola, P., & Jones, M. (2004). Robust real-time face detection. International Journal of Computer Vision, 57(2), 137V154.
Wang, Y., Narayanaswamy, A., Tsai, C. -L., & Roysam, B. (2011). A broadly applicable 3-D neuron tracing method based on open-curve snake. Neuroinformatics, 9, 193–217.
Wearne, S., Rodriguez, A., Ehlenberger, D., Rocher, A., Henderson, S., & Hof, P. (2005). New techniques for imaging, digitization and analysis of three-dimensional neural morphology on multiple scales. Neuroscience, 136, 661–680.
Xiao, H., & Peng, H. (2013). APP2: automatic tracing of 3D neuron morphology based on hierarchical pruning of a gray-weighted image distance-tree. Bioinformatics, 29, 1448–1454.
Yang, J., Gonzalez-Bellido, P.T., & Peng, H. (2013). A distance-field based automatic neuron tracing method. BMC Bioinformatics, 14. doi:10.1186/1471-2105-14-93.
Zhou, Z., Sorensen, S., Zeng, H., Hawrylycz, M., & Peng, H. (2014). Adaptive image enhancement for tracing 3D morphologies of neurons and brain vasculatures. Neuroinform, 13, 153–166.
Acknowledgments
Ching-Wei Wang, Hilmil Pradana and Yu-Ching Lee were supported by the Ministry of Science and Technology of Taiwan, under a grant (MOST-105-2221-E-011-121-MY2). Zhi Zhou and Hanchuan Peng were supported by Allen Institute for Brain Science, Seattle, WA, USA.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, CW., Lee, YC., Pradana, H. et al. Ensemble Neuron Tracer for 3D Neuron Reconstruction. Neuroinform 15, 185–198 (2017). https://doi.org/10.1007/s12021-017-9325-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12021-017-9325-1