Abstract
Objective
The aim of this study was to check the hypothesis concerning the crucial role of DNA methylation (one of the epigenetic mechanisms) within selected genes related to the destruction and regeneration of neural cells and its input in the pathogenesis of diabetic neuropathy, using a model of the DNA in peripheral blood cells.
Methods
A cross-sectional, case-control study was conducted, consisting of 24 adult Type 1 Diabetes Melitus (T1DM) patients with autonomic neuropathy (CAN), 25 T1DM patients without neuropathy and 25 matched, healthy adults acting as a control (Ctrl). The Ewing’s tests, using the ProSciCard apparatus (Mewicon CATEEM-Tec GmbH), was employed to assess the severity of the patients’ symptoms of autonomic neuropathy. For DNA methylation analysis, DNA material of each sample DNA after bisulfite conversion was used for the hybridization of BeadChips (Infinium Methylation EPIC Kit, Illumina), and imaged on the Illumina HiScan. The changes in the expression of selected genes were examined using real-time PCR. Probes were labeled using fluorescein amidite, FAM (Thermo Fisher Scientific). Amplification was performed using the continuous fluorescence detection 7900 HT Fast Real-Time PCR system (Thermo Fisher Scientific). The expression ratio of the target mRNA was normalized to the level of 18s RNA and compared with the control. Statistical analysis was performed using Statistica version 13.1. The statistically significant results were recognized, with a value of p < 0.05.
Results
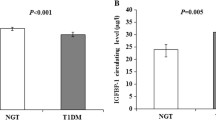
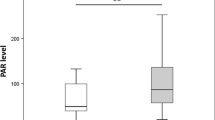
Clinical analysis of the investigated groups revealed a significantly higher percentage of personal insulin pump users in the group without neuropathy. The glucose metabolic control, based on the HbA1c level analysis, was also significantly better in T1DM patients without CAN. The Bumphunter method for DNA methylation analysis showed statistically significant regions related to the genes involved in nerve regeneration ninjurin 2 (NINJ2) and functionality (BR serine/threonine kinase 2 BRSK2, claudin 4 CLDN4). When compared with T1DM patients without neuropathy, T1DM patients with neuropathy showed significantly increased methylation in the first NINJ2 axon, and a lower level of DNA methylation in the region of the first intron of BRSK2, as well as the CLDN4 5′UTR regions. The qRT-PCR results confirmed the decreased expression of NINJ2 and CLDN4 genes in patients with T1DM with CAN.
Conclusions
The different DNA methylation profiles, correlating with the expression of genes related to nervous tissue development and regeneration in patients with T1DM with autonomic neuropathy provide evidence for the role of epigenetic mechanisms promoting the development of CAN, a chronic complication of T1DM.

Similar content being viewed by others
References
V. Spallone, D. Ziegler, R. Freeman, L. Bernardi, S. Frontoni, R. Pop-Busui, M. Stevens, P. Kempler, J. Hilsted, S. Tesfaye, P. Low, P. Valensi, Cardiovascular autonomic neuropathy in diabetes: clinical impact, assessment, diagnosis, and management. Diabetes Metab. Res Rev. 27(7), 639–653 (2011)
Aaron Vinik Carolina Casellini, Marie-Laure Nevoret. https://www.endotext.org/chapter/diabetic-complications/diabetic-neuropathies/ (2018)
D. Ziegler, Diabetic cardiovascular autonomic neuropathy: prognosis, diagnosis and treatment. Diabetes Metab. Rev. 10, 339–383 (1994)
The effect of intensive diabetes therapy on measures of autonomic nervous system function in the Diabetes Control and Complications Trial (DCCT) Diabetologia. 1998;41:416–423. https://doi.org/10.1007/s001250050924
R. Pop-Busui, P.A. Low, B.H. Waberski, C.L. Martin, J.W. Albers, E.L. Feldman, C. Sommer, P.A. Cleary, J.M. Lachin, W.H. Herman, Effects of prior intensive insulin therapy on cardiac autonomic nervous system function in type 1 diabetes mellitus: the Diabetes Control and Complications Trial/Epidemiology of Diabetes Interventions and Complications study (DCCT/EDIC). Circulation 119, 2886–2893 (2009)
M.A. Reddy, E. Zhang, R. Natarajan, Epigenetic mechanisms in diabetic complications and metabolic memory. Diabetologia 58, 443–455 (2015). https://doi.org/10.1007/s00125-014-3462-y
Peter A. Jones, Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484 (2012). no. 7
2018 Guidelines on the management of diabetic patients. A position of diabetes Poland. Clin. Diabetol. 7(1), 1–90 (2018). https://doi.org/10.5603/DK.2018.0001
J. Maksimovic, L. Gordon, A. Oshlack, SWAN: Subset-quantile within array normalization for illumina infinium HumanMethylation450 BeadChips. Genome Biol. 13(6), R44 (2012). https://doi.org/10.1186/gb-2012-13-6-r44
W.E. Johnson, C. Li, A. Rabinovic, Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007). https://doi.org/10.1093/biostatistics/kxj037
J.T. Leek, W.E. Johnson, H.S. Parker, et al. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012). https://doi.org/10.1093/bioinformatics/bts034
E.A. Houseman, W.P. Accomando, D.C. Koestler, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 13, 86 (2012). https://doi.org/10.1186/1471-2105-13-86
G.K. Smyth, Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl Genet. Mol. Biol. 3, 1–25 (2004). https://doi.org/10.2202/1544-6115.1027
A.E. Jaffe, P. Murakami, H. Lee, et al. Bump hunting to identify differentially methylated regions in epigenetic epidemiology studies. Int J. Epidemiol. 41(1), 200–209 (2012). https://doi.org/10.1093/ije/dyr238
R Core Team. R: a language and environment for statistical computing. (R Foundation for Statistical Computing, Vienna, Austria, 2013)
R.C. Gentleman, V.J. Carey, D.M. Bates et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5(10), R80 (2004)
J.M. Lizcano, O. Goransson, R. Toth, M. Deak, N.A. Morrice, J. Boudeau, S.A. Hawley, L. Udd, T.P. Mäkelä, D.G. Hardie, D.R. Alessi, LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. Embo J. 23, 833–843 (2004)
M. Kishi, Y.A. Pan, J.G. Crump, J.R. Sanes, Mammalian SAD kinase are required for neuronal polarization. Science 307, 929–932 (2005)
B.N. Lilley, Y.A. Pan, J.R. Sanes, SAD kinases sculpt axonal arbors of sensory neurons through long-and short-term responses to neurotrophin signals. Neuron 79(1), 39–53 (2013)
N.J. Bright, D. Carling, C. Thornton, Investigating the regulation of brain-specific kinases 1 and 2 by phosphorylation. J. Biol. Chem. 283(22), 14946–14954 (2008)
A. J. Davies, H. W. Kim, R. Gonzalez-Cano, J. Choi, S. K. Back, S. E. Roh, E. Johnson, M. Gabriac, M. S. Kim, J. Lee, J. E. Lee, Natural killer cells degenerate intact sensory afferents following nerve injury. Cell 176(4), 716–728 (2019)
B. Kiec-Wilk, B. Matejko, U. Razny, M. Stankiewicz, J. Skupien, T. Klupa, M.T. Malecki, Hypoglycemic episodes are associated with inflammatory status in patients with type 1 diabetes mellitus. Atherosclerosis 251, 334–338 (2016). https://doi.org/10.1016/j.atherosclerosis.2016.05.002
H. Honda, M.J. Pazin, H. Ji, R.P. Wernyj, P.J. Morin, Crucial roles of Sp1 and epigenetic modifications in the regulation of the CLDN4 promoter in ovarian cancer cells. J. Biol. Chem. 281(30), 21433–21444 (2006)
Y. Zhang, N. Fatima, M.L. Duafu, Coordinated changes in DNA methylation and histone modifications regulate silencing/derepression of luteinizing hormone receptor gene transcription. Mol. Cell Biol. 25(18), 7929–7939 (2005). https://doi.org/10.1128/MCB.25.18.7929-7939.2005
A. Blattler, P.J. Farnham, Cross-talk between site-specific transcription factors and DNA methylation states. J. Biol. Chem. 288(48), 34287–34294 (2013). https://doi.org/10.1074/jbc.R113.512517
L. Elkouby-Naor, T. Ben-Yosef, Functions of claudin tight junction proteins and their complex interactions in various physiological systems. Int Rev. Cell Mol. Biol. 279, 1–32 (2010)
M.J. Kwon, Emerging roles of claudins in human cancer. Int J. Mol. Sci. 14, 18148–18180 (2013)
D. Gunzel, A.S. Yu, Claudins and the modulation of tight junction permeability. Physiol. Rev. 93, 525–569 (2013)
Tatsuo Miyamoto at all, Tight junctions in Schwann cells of peripheral myelinated axons: a lesson from claudin-19–deficient mice. J. Cell Biol. 169, 527–538 (2005)
T. Araki, J. Milbrandt, Ninjurin2, a novel homophilic adhesion molecule, is expressed in mature sensory and enteric neurons and promotes neurite outgrowth. J. Neurosci. 20, 187–195 (2000)
Guozhuan Zhang, Yingjiao Sun, Lei Wang, Hui Tian, Lishuang Liang, Association of serum Ninjurin2 levels with neurologic damage and postherpetic neuralgia occurrence: an observational cohort study in chinese herpeszoster patients. Oncotarget 8(42), 71520–71527 (2017). https://doi.org/10.18632/oncotarget.17640
R. Jaenisch, A. Bird, Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 33(3), 245–254 (2003)
S. Ghosh, A.J. Yates, M.C. Frühwald, J.C. Miecznikowski, C. Plass, D. Smiraglia, Tissue specific DNA methylation of CpG islands in normal human adult somatic tissues distinguishes neural from non-neural tissues. Epigenetics 5(6), 527–538 (2010)
GUO, U. Junjie et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat. Neurosci. 14.10, 1345 (2011)
R. Lister, E.A. Mukamel, J.R. Nery, M. Urich, C.A. Puddifoot, N.D. Johnson, J. Lucero, Y. Huang, A.J. Dwork, M.D. Schultz, M. Yu, Global epigenomic reconfiguration during mammalian brain development. Science 341(6146), 1237905 (2013)
T.P. Wu, T. Wang, M.G. Seetin, Y. Lai, S. Zhu, K. Lin, Y. Liu, S.D. Byrum, S.G. Mackintosh, M. Zhong, A. Tackett, DNA methylation on N 6-adenine in mammalian embryonic stem cells. Nature 532(7599), 329 (2016)
Funding
Work co-financed from the scientific grant of the National Science Center no 2014/13/B/NZ4/00149.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Opinion no. KBET/192/B/2014.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gastoł, J., Kapusta, P., Polus, A. et al. Epigenetic mechanism in search for the pathomechanism of diabetic neuropathy development in diabetes mellitus type 1 (T1DM). Endocrine 68, 235–240 (2020). https://doi.org/10.1007/s12020-019-02172-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-019-02172-9