Abstract
Purposess
The purpose of this study was using next-generation sequencing technique to explore the potential association between germline variants of 14 targeted genes and papillary thyroid carcinoma (PTC) predisposition as well as disease progression.
Methods
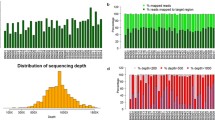
In all, 516 subjects were enrolled in this study including 416 PTC patients and 100 healthy controls. PTC patients were divided into distant metastasis group and non-distant metastasis group. Patients in distant metastasis group were further divided into radioiodine-refractory PTC (RR-PTC) and non-RR-PTC depending on their response to radioiodine therapy. Genomic DNA was extracted from peripheral blood sample and MiSeq Benchtop Sequencer was used for sequencing.
Results
We found rs11246050 in NLRP6 (dominant model, OR/95% CI: 2.028/1.091–3.769, p = 0.025), rs2286742 and rs3740530 in HABP2 (recessive model, OR/95% CI: 9.644/1.307–71.16, p = 0.026 and 3.989/1.413–11.26, p = 0.009), rs2736098 in TERT (recessive model, OR/95% CI: 2.322/1.028–5.242. p = 0.042) and rs62054619 in GAS8-AS1 (recessive model, OR/95% CI: 2.219/1.067–4.617, p = 0.033) were associated with the risk of PTC. rs1137282 in KRAS (dominant model, OR/95% CI: 0.5430/0.3192–0.9236, p = 0.024), rs1347591 and rs4461062 in NUP93 (dominant model, OR/95% CI: 0.6121/0.4128–0.9076, p = 0.015 and 0.6156/0.4157–0.9117, p = 0.015) were associated with low risk of distant metastatic disease in PTC patients. rs33954691 in TERT was associated with the risk of RR-PTC under dominant model (OR/95% CI: 3.161/1.596–6.262).
Conclusions
Germline variants of related genes could be associated with the susceptibility of PTC as well as disease progression (distant metastasis and radioiodine-refractory status). However, these results must be further verified and the potential biological functions of these germline variants in the pathogenesis of PTC remain to be determined in future studies.

Similar content being viewed by others
References
Z.L. Qiu, H.J. Song, Y.H. Xu, Q.Y. Luo, Efficacy and survival analysis of 131I therapy for bone metastases from differentiated thyroid cancer. J. Clin. Endocrinol. Metab. 96(10), 3078–3086 (2011). https://doi.org/10.1210/jc.2011-0093
C.T. Shen, Z.L. Qiu, Q.Y. Luo, Sorafenib in the treatment of radioiodine-refractory differentiated thyroid cancer: a meta-analysis. Endocr. Relat. Cancer 21(2), 253–261 (2014). https://doi.org/10.1530/ERC-13-0438
H.J. Song, Z.L. Qiu, C.T. Shen, W.J. Wei, Q.Y. Luo, Pulmonary metastases in differentiated thyroid cancer: efficacy of radioiodine therapy and prognostic factors. Eur. J. Endocrinol. 173(3), 399–408 (2015). https://doi.org/10.1530/EJE-15-0296
J. Gudmundsson, P. Sulem, D.F. Gudbjartsson, J.G. Jonasson, A. Sigurdsson, J.T. Bergthorsson, H. He, T. Blondal, F. Geller, M. Jakobsdottir, D.N. Magnusdottir, S. Matthiasdottir, S.N. Stacey, O.B. Skarphedinsson, H. Helgadottir, W. Li, R. Nagy, E. Aguillo, E. Faure, E. Prats, B. Saez, M. Martinez, G.I. Eyjolfsson, U.S. Bjornsdottir, H. Holm, K. Kristjansson, M.L. Frigge, H. Kristvinsson, J.R. Gulcher, T. Jonsson, T. Rafnar, H. Hjartarsson, J.I. Mayordomo, A. de la Chapelle, J. Hrafnkelsson, U. Thorsteinsdottir, A. Kong, K. Stefansson, Common variants on 9q22.33 and 14q13.3 predispose to thyroid cancer in European populations. Nat. Genet. 41(4), 460–464 (2009). https://doi.org/10.1038/ng.339
M. Takahashi, V.A. Saenko, T.I. Rogounovitch, T. Kawaguchi, V.M. Drozd, H. Takigawa-Imamura, N.M. Akulevich, C. Ratanajaraya, N. Mitsutake, N. Takamura, L.I. Danilova, M.L. Lushchik, Y.E. Demidchik, S. Heath, R. Yamada, M. Lathrop, F. Matsuda, S. Yamashita, The FOXE1 locus is a major genetic determinant for radiation-related thyroid carcinoma in Chernobyl. Hum. Mol. Genet. 19(12), 2516–2523 (2010). https://doi.org/10.1093/hmg/ddq123
J. Gudmundsson, P. Sulem, D.F. Gudbjartsson, J.G. Jonasson, G. Masson, H. He, A. Jonasdottir, A. Sigurdsson, S.N. Stacey, H. Johannsdottir, H.T. Helgadottir, W. Li, R. Nagy, M.D. Ringel, R.T. Kloos, M.C. de Visser, T.S. Plantinga, M. den Heijer, E. Aguillo, A. Panadero, E. Prats, A. Garcia-Castano, A. De Juan, F. Rivera, G.B. Walters, H. Bjarnason, L. Tryggvadottir, G.I. Eyjolfsson, U.S. Bjornsdottir, H. Holm, I. Olafsson, K. Kristjansson, H. Kristvinsson, O.T. Magnusson, G. Thorleifsson, J.R. Gulcher, A. Kong, L.A. Kiemeney, T. Jonsson, H. Hjartarson, J.I. Mayordomo, R.T. Netea-Maier, A. de la Chapelle, J. Hrafnkelsson, U. Thorsteinsdottir, T. Rafnar, K. Stefansson, Discovery of common variants associated with low TSH levels and thyroid cancer risk. Nat. Genet. 44(3), 319–322 (2012). https://doi.org/10.1038/ng.1046
H.Y. Son, Y. Hwangbo, S.K. Yoo, S.W. Im, S.D. Yang, S.J. Kwak, M.S. Park, S.H. Kwak, S.W. Cho, J.S. Ryu, J. Kim, Y.S. Jung, T.H. Kim, S.J. Kim, K.E. Lee, D.J. Park, N.H. Cho, J. Sung, J.S. Seo, E.K. Lee, Y.J. Park, J.I. Kim, Genome-wide association and expression quantitative trait loci studies identify multiple susceptibility loci for thyroid cancer. Nat. Commun. 8, 15966 (2017). https://doi.org/10.1038/ncomms15966
Y.L. Wang, S.H. Feng, S.C. Guo, W.J. Wei, D.S. Li, Y. Wang, X. Wang, Z.Y. Wang, Y.Y. Ma, L. Jin, Q.H. Ji, J.C. Wang, Confirmation of papillary thyroid cancer susceptibility loci identified by genome-wide association studies of chromosomes 14q13, 9q22, 2q35 and 8p12 in a Chinese population. J. Med. Genet. 50(10), 689–695 (2013). https://doi.org/10.1136/jmedgenet-2013-101687
M. Matsuse, M. Takahashi, N. Mitsutake, E. Nishihara, M. Hirokawa, T. Kawaguchi, T. Rogounovitch, V. Saenko, A. Bychkov, K. Suzuki, K. Matsuo, K. Tajima, A. Miyauchi, R. Yamada, F. Matsuda, S. Yamashita, The FOXE1 and NKX2-1 loci are associated with susceptibility to papillary thyroid carcinoma in the Japanese population. J. Med. Genet. 48(9), 645–648 (2011). https://doi.org/10.1136/jmedgenet-2011-100063
S. Liyanarachchi, A. Wojcicka, W. Li, M. Czetwertynska, E. Stachlewska, R. Nagy, K. Hoag, B. Wen, R. Ploski, M.D. Ringel, I. Kozlowicz-Gudzinska, W. Gierlikowski, K. Jazdzewski, H. He, A. de la Chapelle, Cumulative risk impact of five genetic variants associated with papillary thyroid carcinoma. Thyroid 23(12), 1532–1540 (2013). https://doi.org/10.1089/thy.2013.0102
A.M. Jones, K.M. Howarth, L. Martin, M. Gorman, R. Mihai, L. Moss, A. Auton, C. Lemon, H. Mehanna, H. Mohan, S.E. Clarke, J. Wadsley, E. Macias, A. Coatesworth, M. Beasley, T. Roques, C. Martin, P. Ryan, G. Gerrard, D. Power, C. Bremmer, T. Consortium, I. Tomlinson, L.G. Carvajal-Carmona, Thyroid cancer susceptibility polymorphisms: confirmation of loci on chromosomes 9q22 and 14q13, validation of a recessive 8q24 locus and failure to replicate a locus on 5q24. J. Med. Genet. 49(3), 158–163 (2012). https://doi.org/10.1136/jmedgenet-2011-100586
H. He, W. Li, S. Liyanarachchi, Y. Wang, L. Yu, L.K.Genutis, S. Maharry, J.E. Phay, R. Shen, P. Brock, A. de la Chapelle, The role of NRG1 in the predisposition to papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. (2017). https://doi.org/10.1210/jc.2017-01798
S.K. Gara, L. Jia, M.J. Merino, S.K. Agarwal, L. Zhang, M. Cam, D. Patel, E. Kebebew, Germline HABP2 mutation causing familial nonmedullary thyroid cancer. N. Engl. J. Med. 373(5), 448–455 (2015). https://doi.org/10.1056/NEJMoa1502449
T. Zhang, M. Xing, HABP2 G534E mutation in familial nonmedullary thyroid cancer. J. Natl. Cancer Inst. 108(6), djv415 (2016). https://doi.org/10.1093/jnci/djv415
A. Kowalik, D. Gasior-Perczak, M. Gromek, M. Siolek, A. Walczyk, I. Palyga, M. Chlopek, J. Kopczynski, R. Mezyk, A. Kowalska, S. Gozdz, The p.G534E variant of HABP2 is not associated with sporadic papillary thyroid carcinoma in a Polish population. Oncotarget 8(35), 58304–58308 (2017). https://doi.org/10.18632/oncotarget.16870
L.E.B. de Mello, A.N. Araujo, C.X. Alves, F.J.P. de Paiva, J. Brandao-Neto, J.M. Cerutti, The G534E variant in HABP2 is not associated with increased risk of familial nonmedullary thyroid cancer in Brazilian Kindreds. Clin. Endocrinol. 87(1), 113–114 (2017). https://doi.org/10.1111/cen.13352
C. Colombo, M. Muzza, M.C. Proverbio, G. Ercoli, M. Perrino, V. Cirello, L. Vicentini, S. Ferrero, L. Fugazzola, Segregation and expression analyses of hyaluronan-binding protein 2 (HABP2): insights from a large series of familial non-medullary thyroid cancers and literature review. Clin. Endocrinol. 86(6), 837–844 (2017). https://doi.org/10.1111/cen.13316
S. Cantara, C. Marzocchi, M.G. Castagna, F. Pacini, HABP2 G534E variation in familial non-medullary thyroid cancer: an Italian series. J. Endocrinol. Invest. 40(5), 557–560 (2017). https://doi.org/10.1007/s40618-016-0583-9
A.L. Weeks, S.G. Wilson, L. Ward, J. Goldblatt, J. Hui, J.P. Walsh, HABP2 germline variants are uncommon in familial nonmedullary thyroid cancer. BMC Med. Genet. 17(1), 60 (2016). https://doi.org/10.1186/s12881-016-0323-1
M. Ruiz-Ferrer, R.M. Fernandez, E. Navarro, G. Antinolo, S. Borrego, G534E variant in HABP2 and nonmedullary hyroid Cancer. Thyroid 26(7), 987–988 (2016). https://doi.org/10.1089/thy.2016.0193
A.S. Alzahrani, A.K. Murugan, E. Qasem, H. Al-Hindi, HABP2 gene mutations do not cause familial or sporadic non-medullary thyroid cancer in a highly inbred middle eastern population. Thyroid 26(5), 667–671 (2016). https://doi.org/10.1089/thy.2015.0537
R. Sahasrabudhe, J. Stultz, J. Williamson, P. Lott, A. Estrada, M. Bohorquez, C. Palles, G. Polanco-Echeverry, E. Jaeger, L. Martin, M. Magdalena Echeverry, I. Tomlinson, L.G. Carvajal-Carmona, Tcukin: The HABP2 G534E variant is an unlikely cause of familial non-medullary thyroid cancer. J. Clin. Endocrinol. Metab. 10(3), 1098–1103 (2016). https://doi.org/10.1210/jc.2015-3928
Q. Zhang, F. Song, H. Zheng, X. Zhu, F. Song, X. Yao, L. Zhang, K. Chen, Association between single-nucleotide polymorphisms of BRAF and papillary thyroid carcinoma in a Chinese population. Thyroid 23(1), 38–44 (2013). https://doi.org/10.1089/thy.2012.0228
X. Shen, G. Zhu, R. Liu, D. Viola, R. Elisei, E. Puxeddu, L. Fugazzola, C. Colombo, B. Jarzab, A. Czarniecka, A.K. Lam, C. Mian, F. Vianello, L. Yip, G. Riesco-Eizaguirre, P. Santisteban, C.J. O’Neill, M.S. Sywak, R. Clifton-Bligh, B. Bendlova, V. Sykorova, M. Xing, Patient age-associated mortality risk is differentiated by BRAF V600E status in papillary thyroid cancer. J. Clin. Oncol. JCO2017745497 (2017). https://doi.org/10.1200/JCO.2017.74.5497
M. Xing, The T1799A BRAF mutation is not a germline mutation in familial nonmedullary thyroid cancer. Clin. Endocrinol. 63(3), 263–266 (2005). https://doi.org/10.1111/j.1365-2265.2005.02332.x
H. Li, R. Durbin, Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5), 589–595 (2010). https://doi.org/10.1093/bioinformatics/btp698
A. McKenna, M. Hanna, E. Banks, A. Sivachenko, K. Cibulskis, A. Kernytsky, K. Garimella, D. Altshuler, S. Gabriel, M. Daly, M.A. DePristo, The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20(9), 1297–1303 (2010). https://doi.org/10.1101/gr.107524.110
D.C. Koboldt, Q. Zhang, D.E. Larson, D. Shen, M.D. McLellan, L. Lin, C.A. Miller, E.R. Mardis, L. Ding, R.K. Wilson, VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22(3), 568–576 (2012). https://doi.org/10.1101/gr.129684.111
K. Wang, M. Li, H. Hakonarson, ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38(16), e164 (2010). https://doi.org/10.1093/nar/gkq603
I.A. Adzhubei, S. Schmidt, L. Peshkin, V.E. Ramensky, A. Gerasimova, P. Bork, A.S. Kondrashov, S.R. Sunyaev, A method and server for predicting damaging missense mutations. Nat. Methods 7(4), 248–249 (2010). https://doi.org/10.1038/nmeth0410-248
P.C. Ng, S. Henikoff, SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 31(13), 3812–3814 (2003)
J.M. Schwarz, C. Rodelsperger, M. Schuelke, D. Seelow, MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 7(8), 575–576 (2010). https://doi.org/10.1038/nmeth0810-575
S. Purcell, B. Neale, K. Todd-Brown, L. Thomas, M.A. Ferreira, D. Bender, J. Maller, P. Sklar, P.I. de Bakker, M.J. Daly, P.C. Sham, PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81(3), 559–575 (2007). https://doi.org/10.1086/519795
E.Y. Zhou, Z. Lin, Y. Yang, HABP2 mutation and nonmedullary thyroid cancer. N. Engl. J. Med. 373(21), 2084–2085 (2015). https://doi.org/10.1056/NEJMc1511631#SA2
M. Levy, H. Shapiro, C.A. Thaiss, E. Elinav, NLRP6: a multifaceted innate immune sensor. Trends Immunol. 38(4), 248–260 (2017). https://doi.org/10.1016/j.it.2017.01.001
S. Normand, A. Delanoye-Crespin, A. Bressenot, L. Huot, T. Grandjean, L. Peyrin-Biroulet, Y. Lemoine, D. Hot, M. Chamaillard, Nod-like receptor pyrin domain-containing protein 6 (NLRP6) controls epithelial self-renewal and colorectal carcinogenesis upon injury. Proc. Natl Acad. Sci. USA 108(23), 9601–9606 (2011). https://doi.org/10.1073/pnas.1100981108
H. Wang, G. Xu, Z. Huang, W. Li, H. Cai, Y. Zhang, D. Xiong, G. Liu, S. Wang, Z. Xue, Q. Luo, LRP6 targeting suppresses gastric tumorigenesis via P14(ARF)-Mdm2-P53-dependent cellular senescence. Oncotarget 8(67), 111597–111607 (2017). https://doi.org/10.18632/oncotarget.22876
M. Ge, M. Shi, C. An, W. Yang, X. Nie, J. Zhang, Z. Lv, J. Li, L. Zhou, Z. Du, M. Yang, Functional evaluation of TERT-CLPTM1L genetic variants associated with susceptibility of papillary thyroid carcinoma. Sci. Rep. 6, 26037 (2016). https://doi.org/10.1038/srep26037
T. Pang, M. Zhou, R. Liu, J. Luo, R. Xia, TERT rs2736098 (Ex2-659G>A) polymorphism and cancer susceptibility: evidence from a comprehensive meta-analysis. Oncotarget 8(56), 96433–96441 (2017). https://doi.org/10.18632/oncotarget.21703
G. Atzmon, M. Cho, R.M. Cawthon, T. Budagov, M. Katz, X. Yang, G. Siegel, A. Bergman, D.M. Huffman, C.B. Schechter, W.E. Wright, J.W. Shay, N. Barzilai, D.R. Govindaraju, Y. Suh, Evolution in health and medicine Sackler colloquium: Genetic variation in human telomerase is associated with telomere length in Ashkenazi centenarians. Proc. Natl Acad. Sci. USA 107(Suppl. 1), 1710–1717 (2010). https://doi.org/10.1073/pnas.0906191106
W. Pan, L. Zhou, M. Ge, B. Zhang, X. Yang, X. Xiong, G. Fu, J. Zhang, X. Nie, H. Li, X. Tang, J. Wei, M. Shao, J. Zheng, Q. Yuan, W. Tan, C. Wu, M. Yang, D. Lin, Whole exome sequencing identifies lncRNA GAS8-AS1 and LPAR4 as novel papillary thyroid carcinoma driver alternations. Hum. Mol. Genet. 25(9), 1875–1884 (2016). https://doi.org/10.1093/hmg/ddw056
Y. Qin, W. Sun, H. Zhang, P. Zhang, Z. Wang, W. Dong, L. He, T. Zhang, L. Shao, W. Zhang, C. Wu, LncRNA GAS8-AS1 inhibits cell proliferation through ATG5-mediated autophagy in papillary thyroid cancer. Endocrine 59(3), 555–564 (2018). https://doi.org/10.1007/s12020-017-1520-1
L. Ning, W. Rao, Y. Yu, X. Liu, Y. Pan, Y. Ma, R. Liu, S. Zhang, H. Sun, Q. Yu, Association between the KRAS gene polymorphisms and papillary thyroid carcinoma in a Chinese Han population. J. Cancer 7(15), 2420–2426 (2016). https://doi.org/10.7150/jca.16507
J.H. Lee, X.M. Zhao, I. Yoon, J.Y. Lee, N.H. Kwon, Y.Y. Wang, K.M. Lee, M.J. Lee, J. Kim, H.G. Moon, Y. In, J.K. Hao, K.M. Park, D.Y. Noh, W. Han, S. Kim, Integrative analysis of mutational and transcriptional profiles reveals driver mutations of metastatic breast cancers. Cell Discov. 2, 16025 (2016). https://doi.org/10.1038/celldisc.2016.25
Acknowledgements
The authors thank Genesky Bio-Tech Co., Ltd. for their help with gene sequencing and analysis.
Funding
This work was supported by Shanghai key discipline of medical imaging (no. 2017ZZ02005) and the National Natural Science Foundation of China (no. 81771865).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was approved by the ethics committee of Shanghai Jiao Tong University Affiliated Sixth People’s Hospital.
Informed consent
Written informed consent was achieved from each subject (for subject whose age was younger than 18 years, written informed consent was obtained from his/her statutory guardian additionally).
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shen, CT., Zhang, GQ., Qiu, ZL. et al. Targeted next-generation sequencing in papillary thyroid carcinoma patients looking for germline variants predisposing to the disease. Endocrine 64, 622–631 (2019). https://doi.org/10.1007/s12020-019-01878-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-019-01878-0