Abstract
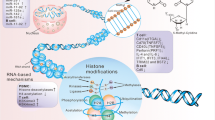
System lupus erythematosus (SLE) is a chronic autoimmune disease characterized by the production of autoantibodies that cause widespread tissue damage. The underlying etiology remains largely unknown. Aberrant epigenetics plays essential roles in the pathogenesis of SLE. This review explores the links between DNA methylation, histone modifications, and miRNAs in SLE and highlights how these factors may interact in SLE pathogenesis. We also discuss how furthering our knowledge of epigenetics in lupus provides hope for finding new diagnostic and prognostic biomarkers and novel therapeutic targets and strategies.


Similar content being viewed by others
References
Wakeland EK, Liu K, Graham RR, Behrens TW (2001) Delineating the genetic basis of systemic lupus erythematosus. Immunity 15:397–408
Tsao BP (2003) The genetics of human systemic lupus erythematosus. Trends Immunol 24:595–602
Cooper GS, Gilbert KM, Greidinger EL et al (2008) Recent advances and opportunities in research on lupus: environmental influences and mechanisms of disease. Environ Health Perspect 116:695–702
Pan Y, Sawalha AH (2009) Epigenetic regulation and the pathogenesis of systemic lupus erythematosus. Transl Res 153:4–10
Richardson B, Scheinbart L, Strahler J et al (1990) Evidence for impaired T cell DNA methylation in systemic lupus erythematosus and rheumatoid arthritis. Arthritis Rheum 33:1665–1673
Corvetta A, Della Bitta R, Luchetti MM, Pomponio G (1991) 5-Methylcytosine content of DNA in blood, synovial mononuclear cells and synovial tissue from patients affected by autoimmune rheumatic diseases. J Chromatogr 566:481–491
Yung RL, Quddus J, Chrisp CE, Johnson KJ, Richardson BC (1995) Mechanism of drug-induced lupus. I. Cloned Th2 cells modified with DNA methylation inhibitors in vitro cause autoimmunity in vivo. J Immunol 154:3025–3035
Quddus J, Johnson KJ, Gavalchin J et al (1993) Treating activated CD4+ T cells with either of two distinct DNA methyltransferase inhibitors, 5-azacytidine or procainamide, is sufficient to cause a lupus-like disease in syngeneic mice. J Clin Invest 92:38–53
Cornacchia E, Golbus J, Maybaum J et al (1988) Hydralazine and procainamide inhibit T cell DNA methylation and induce autoreactivity. J Immunol 140:2197–2200
Richardson B (2007) Primer: epigenetics of autoimmunity. Nat Clin Pract Rheumatol 3:521–527
Richardson BC, Strahler JR, Pivirotto TS et al (1992) Phenotypic and functional similarities between 5-azacytidine-treated T cells and a T cell subset in patients with active systemic lupus erythematosus. Arthritis Rheum 35:647–662
Lu Q, Kaplan M, Ray D et al (2002) Demethylation of ITGAL (CD11a) regulatory sequences in systemic lupus erythematosus. Arthritis Rheum 46:1282–1291
Kaplan MJ, Lu Q, Wu A, Attwood J, Richardson B (2004) Demethylation of promoter regulatory elements contributes to perforin overexpression in CD4+ lupus T cells. J Immunol 172:3652–3661
Lu Q, Wu A, Ray D et al (2003) DNA methylation and chromatin structure regulate T cell perforin gene expression. J Immunol 170:5124–5132
Lu Q, Wu A, Richardson BC (2005) Demethylation of the same promoter sequence increases CD70 expression in lupus T cells and T cells treated with lupus-inducing drugs. J Immunol 174:6212–6219
Oelke K, Lu Q, Richardson D et al (2004) Overexpression of CD70 and overstimulation of IgG synthesis by lupus T cells and T cells treated with DNA methylation inhibitors. Arthritis Rheum 50:1850–1860
Lu Q, Wu A, Tesmer L et al (2007) Demethylation of CD40LG on the inactive X in T cells from women with lupus. J Immunol 179:6352–6358
Mi XB, Zeng FQ (2008) Hypomethylation of interleukin-4 and -6 promoters in T cells from systemic lupus erythematosus patients. Acta Pharmacol Sin 29:105–112
Ballestar E, Esteller M, Richardson BC (2006) The epigenetic face of systemic lupus erythematosus. J Immunol 176:7143–7147
Wen ZK, Xu W, Xu L et al (2007) DNA hypomethylation is crucial for apoptotic DNA to induce systemic lupus erythematosus-like autoimmune disease in SLE-non-susceptible mice. Rheumatology (Oxford) 46:1796–1803
Qiao B, Wu J, Chu YW et al (2005) Induction of systemic lupus erythematosus-like syndrome in syngeneic mice by immunization with activated lymphocyte-derived DNA. Rheumatology (Oxford) 44:1108–1114
Sawalha AH (2008) Epigenetics and T-cell immunity. Autoimmunity 41:245–252
Luo Y, Li Y, Su Y et al (2008) Abnormal DNA methylation in T cells from patients with subacute cutaneous lupus erythematosus. Br J Dermatol 159:827–833
Deng C, Kaplan MJ, Yang J et al (2001) Decreased Ras-mitogen-activated protein kinase signaling may cause DNA hypomethylation in T lymphocytes from lupus patients. Arthritis Rheum 44:397–407
Scheinbart LS, Johnson MA, Gross LA, Edelstein SR, Richardson BC (1991) Procainamide inhibits DNA methyltransferase in a human T cell line. J Rheumatol 18:530–534
Balada E, Ordi-Ros J, Serrano-Acedo S et al (2008) Transcript levels of DNA methyltransferases DNMT1, DNMT3A and DNMT3B in CD4+ T cells from patients with systemic lupus erythematosus. Immunology 124:339–347
Deng C, Lu Q, Zhang Z et al (2003) Hydralazine may induce autoimmunity by inhibiting extracellular signal-regulated kinase pathway signaling. Arthritis Rheum 48:746–756
Sawalha AH, Jeffries M, Webb R et al (2008) Defective T-cell ERK signaling induces interferon-regulated gene expression and overexpression of methylation-sensitive genes similar to lupus patients. Genes Immun 9:368–378
Gorelik G, Fang JY, Wu A, Sawalha AH, Richardson B (2007) Impaired T cell protein kinase C delta activation decreases ERK pathway signaling in idiopathic and hydralazine-induced lupus. J Immunol 179:5553–5563
Miyamoto A, Nakayama K, Imaki H et al (2002) Increased proliferation of B cells and auto-immunity in mice lacking protein kinase Cdelta. Nature 416:865–869
Garcia BA, Busby SA, Shabanowitz J, Hunt DF, Mishra N (2005) Resetting the epigenetic histone code in the MRL-lpr/lpr mouse model of lupus by histone deacetylase inhibition. J Proteome Res 4:2032–2042
Hu N, Qiu X, Luo Y et al (2008) Abnormal histone modification patterns in lupus CD4+ T cells. J Rheumatol 35:804–810
Mishra N, Reilly CM, Brown DR, Ruiz P, Gilkeson GS (2003) Histone deacetylase inhibitors modulate renal disease in the MRL-lpr/lpr mouse. J Clin Invest 111:539–552
Reilly CM, Mishra N, Miller JM et al (2004) Modulation of renal disease in MRL/lpr mice by suberoylanilide hydroxamic acid. J Immunol 173:4171–4178
Mishra N, Brown DR, Olorenshaw IM, Kammer GM (2001) Trichostatin A reverses skewed expression of CD154, interleukin-10, and interferon-gamma gene and protein expression in lupus T cells. Proc Natl Acad Sci U S A 98:2628–2633
Nan X, Ng HH, Johnson CA et al (1998) Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393:386–389
Fuks F, Hurd PJ, Wolf D et al (2003) The methyl-CpG-binding protein MeCP2 links DNA methylation to histone methylation. J Biol Chem 278:4035–4040
Fuks F, Burgers WA, Brehm A, Hughes-Davies L, Kouzarides T (2000) DNA methyltransferase Dnmt1 associates with histone deacetylase activity. Nat Genet 24:88–91
Bartel DP, Chen CZ (2004) Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet 5:396–400
Li QJ, Chau J, Ebert PJ et al (2007) miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129:147–161
Rodriguez A, Vigorito E, Clare S et al (2007) Requirement of bic/microRNA-155 for normal immune function. Science 316:608–611
Vigorito E, Perks KL, Abreu-Goodger C et al (2007) microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 27:847–859
Calame K (2007) MicroRNA-155 function in B Cells. Immunity 27:825–827
Lu LF, Thai TH, Calado DP et al (2009) Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity 30:80–91
Taganov KD, Boldin MP, Chang KJ, Baltimore D (2006) NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci U S A 103:12481–12486
Zhou B, Wang S, Mayr C, Bartel DP, Lodish HF (2007) miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc Natl Acad Sci U S A 104:7080–7085
Johnnidis JB, Harris MH, Wheeler RT et al (2008) Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 451:1125–1129
Mendell JT (2008) miRiad roles for the miR-17–92 cluster in development and disease. Cell 133:217–222
Xiao C, Srinivasan L, Calado DP et al (2008) Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat Immunol 9:405–414
Stanczyk J, Pedrioli DM, Brentano F et al (2008) Altered expression of MicroRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum 58:1001–1009
Rigby RJ, Vinuesa CG (2008) SiLEncing SLE: the power and promise of small noncoding RNAs. Curr Opin Rheumatol 20:526–531
Dai Y, Huang YS, Tang M et al (2007) Microarray analysis of microRNA expression in peripheral blood cells of systemic lupus erythematosus patients. Lupus 16:939–946
Dai Y, Sui W, Lan H et al (2008) Comprehensive analysis of microRNA expression patterns in renal biopsies of lupus nephritis patients. Rheumatol Int 29(7):749–54
Yu D, Tan AH, Hu X et al (2007) Roquin represses autoimmunity by limiting inducible T-cell co-stimulator messenger RNA. Nature 450:299–303
Fabbri M, Garzon R, Cimmino A et al (2007) MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci U S A 104:15805–15810
Tuddenham L, Wheeler G, Ntounia-Fousara S et al (2006) The cartilage specific microRNA-140 targets histone deacetylase 4 in mouse cells. FEBS Lett 580:4214–4217
Scott GK, Mattie MD, Berger CE, Benz SC, Benz CC (2006) Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res 66:1277–1281
Saito Y, Liang G, Egger G et al (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9:435–443
Wang Y, Liang Y, Lu Q (2008) MicroRNA epigenetic alterations: predicting biomarkers and therapeutic targets in human diseases. Clin Genet 74:307–315
Arnheim N, Calabrese P (2009) Understanding what determines the frequency and pattern of human germline mutations. Nat Rev Genet 10:478–488
Barros SP, Offenbacher S (2009) Epigenetics: connecting environment and genotype to phenotype and disease. J Dent Res 88:400–408
Figueiredo LM, Cross GA, Janzen CJ (2009) Epigenetic regulation in African trypanosomes: a new kid on the block. Nat Rev Microbiol 7:504–513
Hewagama A, Richardson B (2009) The genetics and epigenetics of autoimmune diseases. J Autoimmun 33:3–11
Invernizzi P (2009) Future directions in genetic for autoimmune diseases. J Autoimmun 33:1–2
Invernizzi P, Pasini S, Selmi C, Gershwin ME, Podda M (2009) Female predominance and X chromosome defects in autoimmune diseases. J Autoimmun 33:12–16
Larizza D, Calcaterra V, Martinetti M (2009) Autoimmune stigmata in Turner syndrome: when lacks an X chromosome. J Autoimmun 33:25–30
Persani L, Rossetti R, Cacciatore C, Bonomi M (2009) Primary ovarian insufficiency: X chromosome defects and autoimmunity. J Autoimmun 33:35–41
Sawalha AH, Harley JB, Scofield RH (2009) Autoimmunity and Klinefelter’s syndrome: when men have two X chromosomes. J Autoimmun 33:31–34
Wells AD (2009) New insights into the molecular basis of T cell anergy: anergy factors, avoidance sensors, and epigenetic imprinting. J Immunol 182:7331–7341
Zernicka-Goetz M, Morris SA, Bruce AW (2009) Making a firm decision: multifaceted regulation of cell fate in the early mouse embryo. Nat Rev Genet 10:467–477
Acknowledgments
This study received financial support from the National Natural Science Foundation of China (nos. 30730083 and 30671883), the National Basic Research Program of China (973 Plan) (2009CB825605), Hunan Natural Science Foundation (no. 06C0049), and The Science Foundation of Hunan Province (no. 06SK3033).
Author information
Authors and Affiliations
Corresponding author
Additional information
Sha Zhao and Hai Long contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhao, S., Long, H. & Lu, Q. Epigenetic Perspectives in Systemic Lupus Erythematosus: Pathogenesis, Biomarkers, and Therapeutic Potentials. Clinic Rev Allerg Immunol 39, 3–9 (2010). https://doi.org/10.1007/s12016-009-8165-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-009-8165-7