Abstract
The effects of gene body DNA methylation on gene regulation still remains highly controversial. In this study, we generated whole genome bisulfite sequencing (WGBS) data with high sequencing depth in induced pluripotent stem cell (iPSC) and neuronal progentior cell (NPC), and investigated the relationship between DNA methylation changes in CpG islands (CGIs) and corresponding gene expression during NPC differentiation. Interestingly, differentially methylated CGIs were more abundant in intragenic regions compared to promoters and these methylated intragenic CGIs (iCGIs) were associated with neuronal development-related genes. When we compared gene expression level of methylated and unmethylated CGIs in intragenic regions, DNA methylation of iCGI was positively correlated with gene expression in contrast with promoter CGIs (pCGIs). To gain insight into regulatory mechanism mediated by iCGI DNA methylation, we executed motif searching in hypermethylated iCGIs and found NEUROD1 as a hypermethylated iCGI binding transcription factor. This study highlights give rise to possibility of activating role of hypermethylation in iCGIs and involvement of neuronal development related TFs.

The relationship between iCGI DNA methylation and expression of associated genes in neuronal developmental process. During iPSC to NPCdifferentiation, iCGI containing neural developmental genes show iCGI's DNA hypermethylation which is accompanied by gene activation and NEUROD1which is one of the core neuronal TFs interacts with hypermethylated iCGI regions.
Similar content being viewed by others
Introduction
It is known that several developmental lineages are under epigenetic regulation such as DNA methylation, histone modification and chromatin interaction [1], which all undergo dynamic modifications in response to various stimuli during differentiation [2]. DNA methylation is a key epigenetic modification that has been characterized extensively and is essential for normal developmental process [3]. While methyl groups can be added to adenine or cytosine of DNA bases, cytosine methylation is usually observed at CpG dinucleotides in mammals [4]. The majority of CpG dinucleotides across the entire genome are methylated, but those within CpG islands (CGIs) are predominantly unmethylated, with exceptions in inactive X chromosomes and imprinted genes [5, 6]. These CGIs are short, interspersed DNA regions differentiated from the genomic landscape by being GC-rich and CpG-rich, averaging 1000 base pairs in length [7]. Most promoters are associated with unmethylated CGIs and methylation status of CGIs exerts effects on gene transcription that determines cell-type and lineage-specific behavior during development [8,9,10], highlighting the importance of regulatory role of DNA methylation in such processes.
It has been suggested that CGIs are evolutionarily conserved to regulate expression of genes by controlling heterochromatin structures and transcription factor (TF) binding [11, 12]. Such important regulatory structures that have undergone selection are classified according to their genomic distribution, namely promoters (transcription start sites), intragenic (within gene bodies) and intergenic (in between gene bodies) [13]. Of interest was the difference in observed effects of DNA methylation on transcriptional activity between promoter CGIs (pCGIs) and intragenic CGIs (iCGIs) – while there is consensus over the role of pCGIs methylation in repressing gene expression, the impact of iCGI methylation is still controversial. Generally, low levels of DNA methylation in promoter CGIs are related with permissive chromatin structures that induce active transcription and proper binding of TFs [14, 15], with the opposite heterochromatin state that encourages transcription repression and interference with TF binding being associated with high levels of DNA methylation [16]. Loss of DNA methylation (hypomethylation) of pCGIs induced by genomic alterations and chemical treatment results in increasing expression of corresponding genes [17], while methylation levels in promoters of germline-specific genes during developmental periods are also inversely correlated with the expression of corresponding genes [18]. On the contrary, studies have reported differing consequences of iCGI methylation on gene expression. Several studies have showcased instances of inverse correlation between DNA methylation in gene body and gene expression, such as iCGIs that were placed under alternative promoters resulted in their transcription repression following gain of methylation (hypermethylation) of these CGIs [7]. In contrast to such repressive roles of hypermethylation in gene body, others have presented co-occurrence of DNA hypermethylation in gene body regions with H3K36me3, a widely-accepted transcription elongation mark [19]. Our previous study also reported on the positive correlation between DNA methylation in gene body and expression of corresponding genes in embryonic stem cell differentiation [20]. Taken together, the role of intragenic DNA methylation on gene expression remains debatable and calls for more investigation.
Following the ground-breaking report on generation of induced pluripotent stem cells (iPSCs) from dermal fibroblasts through over-expression of Yamanaka factors (Oct4, Sox2, Klf4, cMyc) by Kazutoshi Takahashi and Shinya Yamanaka, iPSCs have been heavily utilized in research involving differentiation into various lineages, and are preferred cell-models to study developmental process or diseases [21, 22]. Given their high-throughtput applications, much focus has been placed on characterizing cellular phenotypes and functions of iPSC and iPSC-derived cells [23, 24] to optimize repgroamming and differentiation efficiencies. Despite the emphasis, epigenetic profiling suggests an ‘epigenetic barrier’, resulting in varying and low reprogramming efficiencies of iPSCs, with issues that spill-over to affect properties of both iPSCs and iPSC-derived cells [25, 26]. Such observations underline the need for comprehensive analysis of epigenome for cellular reprogramming and differentiation to improve efficiency and potency of the iPSC and iPSC-derived cells, for optimum usage in clinical application.
Thus, we set out to refine the relationship between DNA methylation and gene expression during differentiation using iPSCs and iPSC-derived NPCs. Moreover, we generated whole genome bisulfite sequencing (WGBS) data with 10x sequencing depth to analyze DNA methylation at single base resolution to obtain more detailed information compared to the data from Illumina 450 K bead arrays used in most previous studies [27, 28]. In our study, we elucidated that DNA methylation of iCGI plays a crucial role in the activation of lineage determining genes during differentiation from iPSC to NPC. We observed that DNA methylation changes significantly occurred in iCGIs compared to pCGIs, particularly in neuronal developmental genes. We also found that neuronal transcription factors bound to hypermethylated iCGI regions, suggesting that interplay between fate determining factors and DNA methylation is one of the important regulatory mechanisms during differentiation process.
Materials and Methods
iPSC Maintenance and Differentiation into iPSC-Derived NPCs and ENs
Human iPS cell lines were kindly provided by the Yonsei University College of Medicine. The use of human iPSCs was approved by the Institutional Review Board (IRB) of Yonsei University (Permit Number: 7001988–201802-BR-119-01E). For preparation of feeder cells, STO fibroblasts [American Type Culture Collection (ATCC)] were cultured and inactivated with mitomycin C (10 μg/ml; Sigma-Aldrich, St. Louis, MO, USA). iPSCs were cultured on feeder cell layers and maintained using hiPSC maintenance medium consisting of DMEM/F12 (Invitrogen, Carlsbad, CA, USA), 20% knockout serum replacement (Invitrogen), 1% penicillin–streptomycin (Invitrogen), 1× nonessential amino acid (Invitrogen), 0.1 mM β-mercaptoethanol (Sigma-Aldrich), and 10 ng/ml basic fibroblast growth factor (bFGF) (Peprotech, Rocky Hill, NJ, USA) [29,30,31]. For neuronal differentiation, an embryoid body (EB) was generated by culturing human iPSCs for 5–6 days on non-adherent Petri dishes in hiPSCs maintained in medium lacking bFGF. To guide differentiation of EBs toward a neuroectodermal fate, 5 μM dorsomorphin (DM) (Sigma-Aldrich) and 5 μM SB431542 (Sigma-Aldrich) were added to the hiPSC medium during EB formation. EBs were then attached to culture dishes coated with Matrigel (BD Biosciences, San Jose, CA, USA) and cultured in neural induction medium consisting of DMEM/F12 medium (Invitrogen), 1× N2 supplement (Invitrogen), and 1× nonessential amino acids (Invitrogen) for 6 days [29,30,31]. When neural rosettes appeared in the center of the EBs, the NPCs were collected through a manipulated Pasteur pipet. Collected NPCs were re-plated on Matrigel-coated plates for neuronal maturation and cultured for 14 days in Neurobasal medium (Invitrogen) containing 1× B27 supplement without vitamin A (Invitrogen), 1× Glutamax (Invitrogen) supplemented with 10 ng/ml brain-derived neurotrophic factor (BDNF, Peprotech), 10 ng/ml glial cell-derived neurotrophic factor (GDNF, Peprotech), and 10 ng/ml neurotrophin-3 (NT3, Peprotech). ENs were collected by Accutase (Invitrogen) treatment [32].
WGBS Library Construction
Genomic DNA was extracted using the PureLink Genomic DNA Kit (Invitrogen). Total 300 ng of genomic DNA was used for bisulfite conversion as per the manual of EZ DNA Methylation-Gold Kit (Zymo Research, Irvine, CA, USA). Bisulfite-treated genomic DNA sample were used for library preparation by TruSeq DNA Methylation kit manual (Illumina, San Diego, California, United States). We sequenced the bisulfite converted genomic library (2 × 150 bp) on a HiSeqX system.
mRNA-Seq Library Construction
Total RNA from each cell type was extracted using the TRIzol Reagent (Invitrogen) by manufacturer’s protocol. RNA integrity (RIN) values of samples were assessed by Agilent Bioanalyzer and only samples with RIN value greater than 8 were used. Libraries were prepared from 200 ng of total RNA using the TruSeq Rapid SBS Kit or TruSeq SBS Kit v4 (Illumina) by manufacturer’s protocol. Libraries were sequenced on a HiSeq 2500 system for paired-end RNA sequencing data (2 × 101 bp).
Histone ChIP-Seq Library Preparation
We conducted ChIP-seq to profile the genome-wide landscape of histone H3 lysine 4 tri-methylation (H3K4me3) and lysine 27 tri-methylation (H3K27me3) in iPSC and NPC cells; each measurement was made in two biological replicates. Cells (1 × 106) were crosslinked in crosslinking buffer (100 mM NaCl, 0.1 mM EDTA, 5 mM HEPES [pH 8.0], 1% formaldehyde) for 10 min at 25 °C. The crosslinking reaction was quenched by addition of 125 mM glycine, followed by incubation at 25 °C for 5 min with rotation and two washes in ice-cold PBS. The crosslinked cells were re-suspended in SDS lysis buffer (1% SDS, 50 mM Tris-HCl [pH 8.0], 10 mM EDTA) with protease inhibitor (Roche, Basel, Switzerland). Mono- and di-nucleosome-sized chromatin was obtained through sonication (Covaris, S220). The sonicated chromatin was incubated with antibody (H3K4me3, Millipore, 07–473; H3K27me3, Active Motif, 61,017) and DynaBeads Protein A (Thermo Fisher Scientific, Waltham, MA, USA) for 4 h in 4 °C with rotation, and a fraction of the input chromatin was stored to be used as the input control. For H3K4me3 ChIP, the bead complex was subjected to four washes in RIPA wash buffer-β (300 mM NaCl, 1 mM EDTA [pH 8.0], 0.5 mM EGTA [pH 8.0], 1% Triton X-100, 0.2% SDS, 0.1% sodium deoxycholate, 10 mM Tris [pH 8.0]). For H3K27me3, the bead complex was washed five times with RIPA-α (140 mM NaCl, 1 mM EDTA [pH 8.0], 0.5 mM EGTA [pH 8.0], 1% Triton X-100, 0.1% SDS, 0.1% sodium deoxycholate, 10 mM Tris [pH 8.0]) and twice with RIPA-γ (250 mM LiCl, 10 mM Tris-HCl [pH 8.0], 1 mM EDTA, 0.5% NP-40, 0.5% NaDOC). The immunoprecipitated complex was treated with RNase A (QIAGEN, Germantown, MD, USA) and reverse-crosslinked overnight at 68 °C. The immunoprecipitated DNA was recovered using AMPure XP beads (Beckman Coulter, Carlsbad, CA, USA), and ChIP-seq libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit (New England Biolabs, Ipswich, Massachusetts, USA). ChIP-seq libraries were sequenced in 150 bp paired-end mode on the Illumina NovaSeq 6000 platform.
Whole Genome Bisulfite Sequencing Data Processing
We used trim-galore (v.0.4.0) to trim off the adapter sequences or low-quality reads using default options; trim_galore --paired --trim1 -o Filename_R1.fq Filename_R2.fq. Trimmed sequences were then aligned to the human reference genome (hg19) using Bismark (0.14.4) with bowtie2 option; bismark -p 4 --bowtie2 –bam reference −1 Filename_R1_val_1.fq −2 Filename_R2_val_2.fq. Duplicated reads were removed using Picard markduplicate with default options (1.141). Methylated value (Beta value, the ratio of intensities between methylated and un-methylated alleles) for all the covered CpG sites were calculated using bismark_methylation_extractor (0.14.4) with followed options; bismark_methylation_extractor -p --no_overlap --comprehensive --bedGraph --multicore 4. We searched for differentially methylated regions (DMRs) with metilene version 0.2.6, utilizing per-CpG methylation beta values as inputs. In order to obtain broad DMRs, we minimized the splitting of DMR by setting valley filter to 0.1 (−v 0.1). We further filtered the resulting DMRs with differential methylation threshold 30% and significance q-value threshold 0.05. DMR enrichment values were calculated by described methods [33].
mRNA-Sequencing Data Processing
Quality check for reads was executed using FastQC and read statistics were obtained with SAMtools. Raw reads were aligned to the human reference (hg19) using STAR (v2.6.0a) with the support of transcriptome reference (GRCh37, UCSC). From the mapped reads, we further calculated the TPM (Transcripts per Million) using RSEM (v1.2.31) at the gene level with followed options; rsem-calculate-expression --paired-end --star --star-path --paired-end -p 8 Sample_1.fastq.gz Sample_2.fastq.gz. To reduce any technical bias among the samples, we further quantile normalized using R library, preprocesscore [34].
ChIP-Seq Data Processing
For read alignment, we used BWA (0.7.7) to map reads to the human genome sequence (hg19) by followed codes; (1) bwa-0.7.7 aln -t 7 Reference sample_1.fastq > sample_bwa_hg19_1.sai, (2) bwa-0.7.7 aln -t 7 Reference sample_2.fastq > sample_bwa_hg19_2.sai, (3) bwa-0.7.7 sampe Reference sample_bwa_hg19_1.sai sample_bwa_hg19_2.sai sample_1.fastq sample_2.fastq > sample_bwa_hg19_pe.sam. We further removed duplicated reads using Picard MarkDuplicate (1.141) with default options. For peak calling, we used MACS2 (2.1.0) to call the significantly enriched ChIP region compare to the input IgG control by following options; macs2 callpeak –t sample_noDup.bam -c IgG_noDup.bam -f BAM -g hs -n sample_macs_hg19 -B --outdir output_macs_result.
Motif Scanning for Candidates of Hypermethylated iCGIs
For candidates of hypermethylated iCGIs in neuronal activating genes, we used FIMO algorithm in MEME suite [35]. From the chromosome coordinates of iCGI candidates, we extracted genome sequence by using the UCSC genome browser. Based on 11 candidates of iCGIs as input data, we executed FIMO with default options.
Quantitative PCR for NEUROD1 ChIP
We conducted NEUROD1 ChIP with same protocol with histone ChIP-seq with different washing condition. The chromatin-antibody-bead complex was subjected to serial washing with 5 times for RIPA wash buffer-α (140 mM NaCl, 1 mM EDTA [pH 8.0], 0.5 mM EGTA [pH 8.0], 1% Triton X-100, 0.1% SDS, 0.1% sodium deoxycholate, 10 mM Tris [pH 8.0]), and 2 times for RIPA wash buffer-γ (250 mM LiCl, 10 mM Tris-HCl [pH 8.0], 1 mM EDTA, 0.5% NP-40, 0.5% NaDOC). Quantitative PCR (qPCR) reactions were carried out in triplicate on specific genomic regions using KAPA SYBR FAST qPCR Master Mix (KK4600). The resulting signals were normalized for primer efficiency by carrying out qPCR for each primer pair using Input DNA. The following RT-qPCR condition was used: 95.0 °C for 3 min (1 cycle), 94.0 °C for 10 s, 60 °C for 20 s and 72 °C for 30 s (40 cycles). The following primer sequences were used: 5’-GGTGGAGGCTGTGACGC-3′ and 5′- GTCACCCAGTCCTCC CTCTC-3′ for LHX2 hypermethylated iCGI region; 5’-CCTGACGCTGGCTCCTG-3′ and 5’-ACCAGCATGTCTTCACCTCG-3′ for WNT3A hypermethylated iCGI.
Results
The more Frequent DNA Methylation Changes in Intragenic CpG Island during Neuronal Differentiation
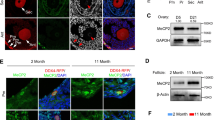
Prior to analyzing DNA methylation changes during differentiation, we validated our differentiation protocol from iPSC to early neuron (EN) by immunostaining of each cell type specific marker. Tra-1-60 and OCT4 for iPSC, Pax6 and Ki67 for NPC and Tuj1 and NeuN for EN were observed (Fig. 1A). We then performed whole genome bisulfite sequencing (WGBS) for iPSCs, NPC and EN produced using the above protocol. We were able to generate DNA methylation data of 10X sequencing depth with a minimum 600 million reads per sample. To observe global differences in DNA methylation, we analyzed differentially methylated regions (DMRs) from iPSC to NPC using Metilene [36] and examined DMR enrichment in promoter CGIs (pCGI) and intragenic CGIs (iCGI), respectively. Interestingly, the enrichment ratio of DNA methylation changes was higher in iCGI regions (Fig. 1B). When we checked DMR changes based on promoter, exon and intron regions, their enrichment ratio were at least 5 times lower compared to iCGI regions (Fig. 1C), so we decided to focus on CGIs in the promoter and intragenic regions. Next, we analyzed differential DNA methylation value based on pCGIs and iCGIs. We defined DNA methylation value of CGIs by averaging methylation ratio (mCG/CG) of all CpG sites in CGIs and then we executed hierarchical clustering using the difference of methylation which were calculated by subtracting iPSC from NPC and iPSC from EN (Fig. 1D). Based on methylation changing aspects, we made three groups; hypermethylation, hypomethylation and no change groups. Interestingly, the degree of hypermethylation gradually increases until NPC to EN, but that of hypomethylation is at the peak in NPC and sustained until EN (Fig. 1D). We next examined the distribution proportion of the three groups in pCGI and iCGI. There were 0.5% of hypomethylation and 10.9% of hypermethylation in pCGI, whereas 6% of hypomethylation and 28% of hypermethylation in iCGI which suggested that there are more dynamic methylation changes in iCGI (Fig. 1E). To examine which genes are related with methylation changes of CGI during differentiation into NPC, we performed gene ontology analysis. Interestingly, the only gene group related with hypermethylated iCGIs are categorized into neuron differentiation (Supplementary Fig. 1A, 1B, 1C, 1D). Thus, we concluded that lineage determining genes are associated with hypermethylation of iCGIs.
DNA methylation changes in intragenic CpG islands during differentiation of iPSCs into NPCs. (A) Characteristic of hiPSCs and neuronal lineage. Immunocytochemical staining of markers of pluripotent cells (Tra-1-60, OCT4), neural progenitor cells (Pax6, Ki67), and mature neurons (Tuj1, NeuN). Scale bar is μm. (B)Enrichment ratio of DMRs in pCGIs and iCGIs during differentiation of iPSCs into NPCs, determined using Metilene. The enrichment ratio was normalized against the length of the genomic elements. (C) Enrichment of differentially methylated regions (DMRs) during differentiation of iPSCs into NPCs in promoter, exon, and intron regions. Enrichment ratio is normalized against the length of the genomic elements. (D) Heatmaps of hierarchical clustering for differences in averaged DNA methylation level in pCGIs (left) and iCGIs (right). Differences in DNA methylation values were calculated by subtracting NPCs from iPSCs and EN from iPSCs. N, NPCs; E, ENs. (E) Ratio of hypermethylation and hypomethylation in pCGIs (left) and iCGIs (right)
The Positive Correlation between Gene Expression and DNA Hypermethylation in iCGIs
To define the epigenetic feature of CGI, we performed chromatin immunoprecipitation sequencing (ChIP-seq) for H3K4me3 in iPSC and NPC as H3K4me3 is regarded as an activation mark and the negative correlation between H3K4me3 and DNA methylation is well known [37,38,39,40]. After executing k-means clustering for total pCGIs and iCGIs, we observed two clusters which showed exclusive patterns between H3K4me3 and DNA methylation. According to the clustering pattern, we could define unmethylated (unmet; high H3K4me3 and low DNA methylation) CGIs and methylated (met; low H3K4me3 and high DNA methylation) CGIs in promoter (Fig. 2A) and intragenic regions (Fig. 2B). We calculated the portion of unmethylated and methylated CGIs and found that 91% of pCGIs in iPSC and 31% of iCGIs were unmethylated (Fig. 2A, B). During differentiation to NPC, these portions were maintained at 89.3% for pCGIs and 32.1% for iCGIs. We then tried to find the correlation between gene expression and DNA methylation status. When we examined the expression of concurrent genes in methylated and unmethylated CGI in promoter, DNA methylation was inversely correlated with gene expression in both iPSC and NPC (Fig. 2C). However, genes that contained methylated iCGIs showed higher expression level than genes that had unmethylated iCGIs in both iPSC and NPC (Fig. 2C). Next, we wanted to investigate whether gene expression alteration by DNA methylation changes in iCGI is still effective during differentiation to NPC. Among 2168 unmethylated iCGIs in iPSC, only 91 iCGIs were turned to be methylated in NPC but the others were still unmethylated (Supplementary Table 1). Likewise, among 4817 methylated iCGIs in iPSC, only 170 iCGIs were turned to be demethylated in NPC but the others were still methylated (Fig. 2D). Interestingly, when we examined the expression of concurrent genes, 91 iCGIs (unmethylated in iPSC but turned to be methylated in NPC) containing genes were significantly increased after differentiation, but the other iCGIs containing genes expressions were not significantly changed (Fig. 2E).
Differing epigenetic patterns of CGIs in promoter and gene body, and comparison of changes in DNA methylation and gene expression. (A) In pCGIs: Heatmaps based on k-means clustering for DNA methylation and H3K4me3 in pCGIs in the iPSC (left) and NPC (right) states. (B) In iCGIs: Heatmaps based on k-means clustering for DNA methylation and H3K4me3 in iCGIs in the iPSC (left) and NPC (right) states. (C) Boxplots representing gene expression levels for genes containing unmethylated or methylated pCGIs (left) and iCGIs (right) in iPSCs and NPCs. (D) Numbers of unmethylated iCGIs in iPSCs that remained unchanged in NPCs and those that changed their methylation status (left bar), with the corresponding numbers for methylated iCGIs in iPSC that changed or maintained their status in NPCs (right bar). Y-axis indicates the ratio of the number of iCGIs. (E) Boxplots of expression levels of genes containing iCGIs that changed from unmethylated to methylated (left) and iCGIs that maintained their unmethylated status (right). ****, p value <0.0001 (unpaired t-test, Mann–Whitney test, two-tailed). For all box-and-whisker plots, the upper and lower bounds of the boxes indicate the 75th and 25th percentiles, respectively, and the horizontal lines in the boxes present the median value. The upper and lower error bars indicate the 90th and 10th percentiles, respectively. ****, p value <0.0001 (unpaired t-test, Mann–Whitney test, two-tailed)
Concurrent Regulation of Bivalent Promoter and iCGI Methylation on Lineage Specific Genes during Differentiation to NPC
In stem cells, many developmental key genes have an unique epigenetic feature consisting the coexistence of active marker (H3K4me3) and repressive marker (H3K27me3) in promoters [41]. These bivalent modifications were generally observed in promoters of stem cells and considered to play a role to pause the expression of corresponding genes [42]. As we found relevance between expressions of neuronal differentiation genes and DNA methylation in their iCGIs during NPC differentiation, we sought to reveal the relationship between histone bivalency and iCGI methylation. We performed H3K27me3 ChIP-seq and analyzed it together with H3K4me3 ChIP-seq data on iCGI containing genes in iPSC. In the genes which have unmethylated iCGIs in iPSC, the intensities of H3K4me3 and H3K27me3 were high in not only promoters but also iCGI regions, however, in the genes which have methylated iCGI in iPSC, only H3K4me3 was detected in promoters (Fig. 3A). Next, we wanted to see whether iCGI DNA methylation affects the expression of bivalent promoter genes in iPSC. Before doing this, we defined bivalent promoters using H3K4me3 and H3K27me3 ChIP-seq data in our iPSC model (Fig. 3B). According to DNA methylation status of iCGIs, we divided bivalent promoter genes into 3 groups; bivalent promoters with unmethylated iCGIs, bivalent promoters with methylated iCGIs and bivalent promoter without iCGI (Fig. 3C). When we looked at expressions of bivalent promoter genes with methylated iCGIs, they showed higher expression than the other groups. The genes with bivalent promoters and unmethylated iCGIs showed the minimum level of expression (Fig. 3C). Thus, we hypothesized that iCGIs as well as bivalent promoters are involved in the regulation of developmental genes through DNA methylation in iPSC. We next set out to select genes which might be controlled by iCGI DNA methylation. Since we sought to look at the genes whose expressions are substantially increased during neuronal development, we selected genes that had bivalent promoters and unmethylated iCGIs in iPSC with low gene expression (log2(Transcript Per Million(TPM)) < 3). When we plotted expression level of selected genes in iPSC and NPC, we observed that some genes became activated in NPC (Fig. 3D). To see whether DNA hypermethylation and gene activation are associated, we profiled the genes by expression changes and DNA methylation differences (Fig. 3E). We observed both induction in neuronal differentiation genes such as Rax, Pax6, Lhx2 and Wnt3a [43,44,45,46] and hypermethylation of these genes during differentiation. The genes that are known to be important for other lineage development, for example, Onecut3a for beta cell, Tbx1 and Tbx5 for heart cell and Hoxb4 for hematopoietic cell, showed neither expression changes or methylation difference (Fig. 3E and Fig. 4A, B). Thus, we concluded that neuronal specific genes are regulated by iCGI methylation together with bivalent promoters during differentiation to NPC.
Bivalent promoter genes related to neural induction in NPC differentiation. (A) Average signal intensities of H3K4me3 and H3K27me3 in the gene structure of both unmethylated iCGI associated genes (blue) and methylated iCGI associated genes (red). Y-axis indicates expression level in log2(RPKM) of H3K4me3 (left) and H3K27me3 (right). (B) Using histone modification data for H3K4me3 and H3K27me3 as active and repressive marks, respectively, total promoters (TSS ± 1 kb) were grouped by hierarchical clustering. Promoters with low H3K4me3 and low H3K27me3 were eliminated. We calculated RPKM using an in-house script; the value in the heatmap is in log2(RPKM). (C) Expression levels of genes containing bivalent promoters with no iCGI (BivProm w/o iCGI), methylated iCGI (BivProm w. met iCGI), and unmethylated iCGI (BivProm w/ unmet iCGI) in iPSCs. The upper and lower whiskers of boxplots indicate the 90th and 10th percentiles, respectively. ****, p value <0.0001 (unpaired t-test, Mann–Whitney test, two-tailed). (D) Scatter plot of expression of genes with bivalent promoters containing unmethylated iCGIs in iPSCs (x-axis) and NPCs (y-axis). (E) Scatter plot of changes in DNA methylation (x-axis) and gene expression (y-axis) during differentiation of iPSCs into NPCs for genes with bivalent promoter containing unmethylated iCGIs
DNA methylation and expression patterns in genetic region of representative genes. (A) UCSC genome browser showing representative genes which were activated during NPC differentiation and hypermethylated in iCGIs. (B) UCSC genome browser showing representative gene which were not activated during NPC differentiation and unchanged in iCGIs
Lineage Determining Factor Binding on the Hypermethylated iCGI Regions of Lineage Specific Genes
Since it is established that cell type-specific transcription factor (TF) mediates lineage specific epigenetic changes [47], we investigated whether specific TF can bind onto hypermethylated iCGI regions. To find TF binding motifs, we executed the FIMO algorithm on iCGI regions [35]. Through our criteria which is that FIMO score is more than 10 and p value is under 5 × 10−5, we selected TF candidates. When we checked the expression pattern of each TF during NPC differentiation, 49 TFs were induced. Intriguingly, bHLH (beta helix-loop-helix) proneural factors such as ASCL1 and NEUROD1 (Fig. 5A) were part of the induced TF group. Although the crucial role of proneural factors on neuronal development is well known, most studies have been conducted in functional neurons converted directly from fibroblasts, with no similar studies in our current iPSC-NPC model to the best of our knowledge. Since Neurod1 was induced more than Ascl1 (Fig. 5A), we decided to execute ChIP-qPCR for NEUROD1 in iPSC and NPC to check whether NEUROD1 indeed binds onto hypermethylated iCGI regions. Among hypermethylated iCGI regions, we observed the binding of NEUROD1 to hypermethylated iCGIs in LHX2 gene body and WNT3A gene body in NPC, but not in iPSC (Fig. 5B). Taken together, we concluded that NEUROD1 as a proneural factor, binds onto the hypermethylated region in specific gene bodies and directs the differentiation into NPC. Methylation at this specific time point may be a pre-requirement for NEUROD1 binding to cognate sites.
Candidate TFs targeted to intragenic CGIs bind to hypermethylated regions. (A) Expression changes in activated candidate TFs during NPC differentiation. (B) ChIP-qPCR was performed on iPSCs and NPCs. NEUROD1 enrichment was quantified in LHX2 (hypermethylated iCGI) and WNT3A (hypermethylated iCGI). Data were normalized against the input sample. Error bars represent standard deviation of fold enrichment in three individual runs
Discussion
The repressive role of DNA methylation on gene expression has been suggested by many studies [17, 48]. While DNA methylation of promoter regions is tightly linked with transcription repression, effects of DNA methylation in gene body regions varies. In this study, we showed that DNA methylation differences were more frequently detected in iCGIs than pCGIs during iPSC to NPC differentiation. As expected, DNA hypermethylation in pCGIs was associated with gene repression, but DNA hypermethylation in iCGI regions of specific neuronal genes positively correlated with gene activation. Interestingly, we observed that NEUROD1, one of the core transcription factors involved in neuronal differentiation, showed physical binding with iCGI regions of LHX2 and WNT3A genes (Fig. 6).
Graphical research scheme. This graphical summary depicts the relationship between iCGI DNA methylation and gene expression in neuronal development associated genes. During iPSC to NPC differentiation, iCGI containing neuronal developmental genes show iCGI DNA hypermethylation which is accompanied by gene activation. NEUROD1 which is one of the core neuronal TFs interacts with hypermethylated iCGI regions
During developmental processes, cells are under strict control to differentiate into optimal lineages and DNA methylation confers epigenetic road signs to guide cellular differentiation and restrict reversion into an undifferentiated state [49]. It is known that a set of germline-specific genes are specifically regulated by promoter DNA methylation for maintaining developmental silencing and many CGIs in promoters maintain unmethylated status through continous exclusion of DNA methyltransferases (DNMTs) via binding with certain TFs [18, 50]. Thus, while the importance and mechanisms of pCGI DNA methlyation as an epigenetic modification in mammals have been established, the role of iCGI methylation in gene expression still remains an open question.Whether iCGI DNA methylation is the cause or consequence of transcriptional activity has also been examined – while it has been demonstrated that transcriptional processes through CGI regions are requried for DNMT3B-mediated DNA methylation [51], our previous study using embryonic stem cell (ESC) data in contrast showed that DNA methylation by DNMT was required for the transcriptional activation of iCGI-containing genes and these iCGI regions also included specific TF binding motifs [20]. Other studies have also implied that DNA methylation on iCGI is not just a consequence of active transcription [52]. In the context of continued uncertainty over the effect iCGI methylation on gene transcription, our study suggest that iCGI methylation is required for activation of neuronal differenation related genes.
The iPSC-NPC differentiation model has been investigated in a variety of ways [53], and TFs such as ASCL1, NEUROG2, TLX3, NEUROD1, NEUROD2 and ATOH1 are now known to be induced during the process [54, 55]. Among them, basic-helix–loop–helix (bHLH) containing TFs, so called proneural factors, carry greater weight in determining differentiation to NPC [56], and ASCL1 alongside NEUROD1 are key proneural factors which are sufficient to drive NPC development independently [57]. The binding sites for NEUROD1 in promoters and enhancers are well covered, but to our best knowledge, binding sites in intragenic regions have not been extensively studied [57]. Moreover, the relationship between proneural factor binding and DNA methylation during NPC differentiation has been rarely studied. Although a recent paper reported that NEUROD2 induced hypomethylation of cognate binding sites by interacting with ten-eleven translocation 2 (TET2), the detailed mechanism still remains elusive [58]. We present NEUROD1, a key proneural factor that binds to methylated iCGI regions to activate transcription of neuronal differentiation related genes such as LHX2 and WNT3A. While several studies indicate that DNA methylation generally blocks TFs from binding to their recognized sites and TF binding to methylated regions is permited through methyl-CpG-binding domains (MBDs) [14], we reasoned that our identified TF candidate behaves similarly to TFs such as Kaiso, CEBPa, ZFP57 and KAP1, which bind to methylated DNA even in the absence of MBDs [59, 60].
In this research, we performed WBGS and ChIP-qPCR analysis on one type of iPS cell line to present the pioneer factors which play a crucial role in neuronal differentiation. However, futher investigation of different types of iPSC with diverse origins and related differing cell lineages are required to further confirm the influence and mechanism of proneuroal factors such as NEUROD1.
Taken together, we conclude that iCGI DNA methylation is as critical as pCGI DNA methylation in controlling development-related genes and hypermethylation in iCGI regions is accompanied with lineage specfic TF binding with a positive correalation with corresponding gene expression. Although further studies such as ChIP-seq for proneural factors is recommended for further validation, our data serves as a spring platform for the discovery and validation of similar transcription factors that play a regulatory role in neuronal differentiation related genes in relation with iCGI methylation such as NEUROD1.
References
Ehrlich, M., & Lacey, M. (2013). DNA methylation and differentiation: Silencing, upregulation and modulation of gene expression. Epigenomics, 5(5), 553–568.
Dixon, J. R., Jung, I., Selvaraj, S., Shen, Y., Antosiewicz-Bourget, J. E., Lee, A. Y., Ye, Z., Kim, A., Rajagopal, N., Xie, W., Diao, Y., Liang, J., Zhao, H., Lobanenkov, V. V., Ecker, J. R., Thomson, J. A., & Ren, B. (2015). Chromatin architecture reorganization during stem cell differentiation. Nature, 518(7539), 331–336.
Nazor, K. L., Altun, G., Lynch, C., Tran, H., Harness, J. V., Slavin, I., Garitaonandia, I., Müller, F. J., Wang, Y. C., Boscolo, F. S., Fakunle, E., Dumevska, B., Lee, S., Park, H. S., Olee, T., D'Lima, D. D., Semechkin, R., Parast, M. M., Galat, V., Laslett, A. L., Schmidt, U., Keirstead, H. S., Loring, J. F., & Laurent, L. C. (2012). Recurrent variations in DNA methylation in human pluripotent stem cells and their differentiated derivatives. Cell Stem Cell, 10(5), 620–634.
Lister, R., Pelizzola, M., Dowen, R. H., Hawkins, R. D., Hon, G., Tonti-Filippini, J., Nery, J. R., Lee, L., Ye, Z., Ngo, Q. M., Edsall, L., Antosiewicz-Bourget, J., Stewart, R., Ruotti, V., Millar, A. H., Thomson, J. A., Ren, B., & Ecker, J. R. (2009). Human DNA methylomes at base resolution show widespread epigenomic differences. Nature, 462(7271), 315–322.
Barlow, D. P. (1995). Gametic imprinting in mammals. Science, 270(5242), 1610–1613.
Yen, P. H., Patel, P., Chinault, A. C., Mohandas, T., & Shapiro, L. J. (1984). Differential methylation of hypoxanthine phosphoribosyltransferase genes on active and inactive human X chromosomes. Proceedings of the National Academy of Sciences of the United States of America, 81(6), 1759–1763.
Deaton, A. M., & Bird, A. (2011). CpG islands and the regulation of transcription. Genes & Development, 25(10), 1010–1022.
Bird, A. (2002). DNA methylation patterns and epigenetic memory. Genes & Development, 16(1), 6–21.
Strichman-Almashanu, L. Z., Lee, R. S., Onyango, P. O., Perlman, E., Flam, F., Frieman, M. B., & Feinberg, A. P. (2002). A genome-wide screen for normally methylated human CpG islands that can identify novel imprinted genes. Genome Research, 12(4), 543–554.
Saxonov, S., Berg, P., & Brutlag, D. L. (2006). A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proceedings of the National Academy of Sciences of the United States of America, 103(5), 1412–1417.
Moore, L. D., Le, T., & Fan, G. (2013). DNA methylation and its basic function. Neuropsychopharmacology, 38(1), 23–38.
Saksouk, N., Simboeck, E., & Dejardin, J. (2015). Constitutive heterochromatin formation and transcription in mammals. Epigenetics & Chromatin, 8, 3.
Medvedeva, Y. A., Fridman, M. V., Oparina, N. J., Malko, D. B., Ermakova, E. O., Kulakovskiy, I. V., Heinzel, A., & Makeev, V. J. (2010). Intergenic, gene terminal, and intragenic CpG islands in the human genome. BMC Genomics, 11, 48–48.
Li, E., & Zhang, Y. (2014). DNA methylation in mammals. Cold Spring Harbor Perspectives in Biology, 6(5), a019133.
Medvedeva, Y. A., Khamis, A. M., Kulakovskiy, I. V., Ba-Alawi, W., Bhuyan, M. S. I., Kawaji, H., Lassmann, T., Harbers, M., Forrest, A. R. R., Bajic, V. B., & The FANTOM consortium. (2014). Effects of cytosine methylation on transcription factor binding sites. BMC Genomics, 15, 119.
Jones, P. A., & Baylin, S. B. (2007). The epigenomics of cancer. Cell, 128(4), 683–692.
Plachot, C., & Lelievre, S. A. (2004). DNA methylation control of tissue polarity and cellular differentiation in the mammary epithelium. Experimental Cell Research, 298(1), 122–132.
Hackett, J. A., Reddington, J. P., Nestor, C. E., Dunican, D. S., Branco, M. R., Reichmann, J., Reik, W., Surani, M. A., Adams, I. R., & Meehan, R. R. (2012). Promoter DNA methylation couples genome-defence mechanisms to epigenetic reprogramming in the mouse germline. Development, 139(19), 3623–3632.
Gates, L. A., Foulds, C. E., & O'Malley, B. W. (2017). Histone Marks in the 'Driver's Seat': Functional roles in steering the transcription cycle. Trends in Biochemical Sciences, 42(12), 977–989.
Lee, S. M., Lee, J., Noh, K. M., Choi, W. Y., Jeon, S., Oh, G. T., Kim-Ha, J., Jin, Y., Cho, S. W., & Kim, Y. J. (2017). Intragenic CpG islands play important roles in bivalent chromatin assembly of developmental genes. Proceedings of the National Academy of Sciences of the United States of America, 114(10), E1885–E1894.
Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126(4), 663–676.
Marchetto, M. C., Brennand, K. J., Boyer, L. F., & Gage, F. H. (2011). Induced pluripotent stem cells (iPSCs) and neurological disease modeling: Progress and promises. Human Molecular Genetics, 20(R2), R109–R115.
Liu, C., et al., (2018) Modeling human diseases with induced pluripotent stem cells: from 2D to 3D and beyond. Development, 145(5).
McCauley, H. A., & Wells, J. M. (2017). Pluripotent stem cell-derived organoids: Using principles of developmental biology to grow human tissues in a dish. Development, 144(6), 958–962.
Vaskova, E. A., Stekleneva, A. E., Medvedev, S. P., & Zakian, S. M. (2013). "epigenetic memory" phenomenon in induced pluripotent stem cells. Acta Naturae, 5(4), 15–21.
Ang, Y. S., Gaspar-Maia, A., Lemischka, I. R., & Bernstein, E. (2011). Stem cells and reprogramming: Breaking the epigenetic barrier? Trends in Pharmacological Sciences, 32(7), 394–401.
de Boni, L., et al., (2018) DNA methylation alterations in iPSC- and hESC-derived neurons: Potential implications for neurological disease modeling. Clinical Epigenetics. 10.
Roost, M. S., Slieker, R. C., Bialecka, M., van Iperen, L., Gomes Fernandes, M. M., He, N., Suchiman, H. E. D., Szuhai, K., Carlotti, F., de Koning, E. J. P., Mummery, C. L., Heijmans, B. T., & Chuva de Sousa Lopes, S. M. (2017). DNA methylation and transcriptional trajectories during human development and reprogramming of isogenic pluripotent stem cells. Nature Communications, 8, 908.
Kim, H. W., Yang, K., Jeong, W. J., Choi, S. J., Lee, J. S., Cho, A. N., Chang, G. E., Cheong, E., Cho, S. W., & Lim, Y. B. (2016). Photoactivation of noncovalently assembled peptide ligands on carbon nanotubes enables the dynamic regulation of stem cell differentiation. ACS Applied Materials & Interfaces, 8(40), 26470–26481.
Seo, H. I., Cho, A. N., Jang, J., Kim, D. W., Cho, S. W., & Chung, B. G. (2015). Thermo-responsive polymeric nanoparticles for enhancing neuronal differentiation of human induced pluripotent stem cells. Nanomedicine, 11(7), 1861–1869.
Shin, J., Choi, E. J., Cho, J. H., Cho, A. N., Jin, Y., Yang, K., Song, C., & Cho, S. W. (2017). Three-dimensional Electroconductive hyaluronic acid hydrogels incorporated with carbon nanotubes and Polypyrrole by catechol-mediated dispersion enhance neurogenesis of human neural stem cells. Biomacromolecules, 18(10), 3060–3072.
Li, L., Roh, J. H., Chang, E. H., Lee, Y., Lee, S., Kim, M., Koh, W., Chang, J. W., Kim, H. J., Nakanishi, M., Barker, R. A., Na, D. L., & Song, J. (2018). iPSC modeling of Presenilin1 mutation in Alzheimer's disease with cerebellar ataxia. Experimental Neurobiology, 27(5), 350–364.
Lee, S. M., Lee, Y. G., Bae, J. B., Choi, J. K., Tayama, C., Hata, K., Yun, Y., Seong, J. K., & Kim, Y. J. (2014). HBx induces hypomethylation of distal intragenic CpG islands required for active expression of developmental regulators. Proceedings of the National Academy of Sciences of the United States of America, 111(26), 9555–9560.
Bolstad, B. M., Irizarry, R. A., Astrand, M., & Speed, T. P. (2003). A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics, 19(2), 185–193.
Grant, C. E., Bailey, T. L., & Noble, W. S. (2011). FIMO: Scanning for occurrences of a given motif. Bioinformatics, 27(7), 1017–1018.
Juhling, F., et al. (2016). Metilene: Fast and sensitive calling of differentially methylated regions from bisulfite sequencing data. Genome Research, 26(2), 256–262.
Greenberg, M. V. C., & Bourc'his, D. (2019). The diverse roles of DNA methylation in mammalian development and disease. Nature Reviews. Molecular Cell Biology, 20(10), 590–607.
Chen, K.F., et al., (2015) Broad H3K4me3 is associated with increased transcription elongation and enhancer activity at tumor-suppressor genes. Nature Genetics. 47(10): p. 1149−+.
Heintzman, N. D., Stuart, R. K., Hon, G., Fu, Y., Ching, C. W., Hawkins, R. D., Barrera, L. O., van Calcar, S., Qu, C., Ching, K. A., Wang, W., Weng, Z., Green, R. D., Crawford, G. E., & Ren, B. (2007). Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genetics, 39(3), 311–318.
Barski, A., Cuddapah, S., Cui, K., Roh, T. Y., Schones, D. E., Wang, Z., Wei, G., Chepelev, I., & Zhao, K. (2007). High-resolution profiling of histone methylations in the human genome. Cell, 129(4), 823–837.
Voigt, P., Tee, W. W., & Reinberg, D. (2013). A double take on bivalent promoters. Genes & Development, 27(12), 1318–1338.
Azuara, V., Perry, P., Sauer, S., Spivakov, M., Jørgensen, H. F., John, R. M., Gouti, M., Casanova, M., Warnes, G., Merkenschlager, M., & Fisher, A. G. (2006). Chromatin signatures of pluripotent cell lines. Nature Cell Biology, 8(5), 532–U189.
Hou, P. S., Chuang, C. Y., Kao, C. F., Chou, S. J., Stone, L., Ho, H. N., Chien, C. L., & Kuo, H. C. (2013). LHX2 regulates the neural differentiation of human embryonic stem cells via transcriptional modulation of PAX6 and CER1. Nucleic Acids Research, 41(16), 7753–7770.
Bel-Vialar, S., Medevielle, F., & Pituello, F. (2007). The on/off of Pax6 controls the tempo of neuronal differentiation in the developing spinal cord. Developmental Biology, 305(2), 659–673.
Muranishi, Y., Terada, K., & Furukawa, T. (2012). An essential role for Rax in retina and neuroendocrine system development. Development, Growth & Differentiation, 54(3), 341–348.
Mattes, B., Weber, S., Peres, J., Chen, Q., Davidson, G., Houart, C., & Scholpp, S. (2012). Wnt3 and Wnt3a are required for induction of the mid-diencephalic organizer in the caudal forebrain. Neural Development, 7, 12.
Iwafuchi-Doi, M., & Zaret, K. S. (2014). Pioneer transcription factors in cell reprogramming. Genes & Development, 28(24), 2679–2692.
Rhee, I., Bachman, K. E., Park, B. H., Jair, K. W., Yen, R. W. C., Schuebel, K. E., Cui, H., Feinberg, A. P., Lengauer, C., Kinzler, K. W., Baylin, S. B., & Vogelstein, B. (2002). DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature, 416(6880), 552–556.
Messerschmidt, D. M., Knowles, B. B., & Solter, D. (2014). DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes & Development, 28(8), 812–828.
Smith, Z. D., & Meissner, A. (2013). DNA methylation: Roles in mammalian development. Nature Reviews. Genetics, 14(3), 204–220.
Jeziorska, D. M., Murray, R. J. S., de Gobbi, M., Gaentzsch, R., Garrick, D., Ayyub, H., Chen, T., Li, E., Telenius, J., Lynch, M., Graham, B., Smith, A. J. H., Lund, J. N., Hughes, J. R., Higgs, D. R., & Tufarelli, C. (2017). DNA methylation of intragenic CpG islands depends on their transcriptional activity during differentiation and disease. Proceedings of the National Academy of Sciences of the United States of America, 114(36), E7526–E7535.
Xu, W., Li, J., Rong, B., Zhao, B., Wang, M., Dai, R., Chen, Q., Liu, H., Gu, Z., Liu, S., Guo, R., Shen, H., Wu, F., & Lan, F. (2020). DNMT3A reads and connects histone H3K36me2 to DNA methylation. Protein & Cell, 11(2), 150–154.
Frati, G., Luciani, M., Meneghini, V., de Cicco, S., Ståhlman, M., Blomqvist, M., Grossi, S., Filocamo, M., Morena, F., Menegon, A., Martino, S., & Gritti, A. (2018). Human iPSC-based models highlight defective glial and neuronal differentiation from neural progenitor cells in metachromatic leukodystrophy. Cell Death & Disease, 9, 698.
D'Aiuto, L., et al. (2015). Large-scale generation of human iPSC-derived neural stem cells/early neural progenitor cells and their neuronal differentiation (vol 10, pg 365, 2014). Organogenesis, 11(2), 93–93.
Simao, D., et al. (2018). Recapitulation of human neural microenvironment signatures in iPSC-derived NPC 3D differentiation. Stem Cell Reports, 11(2), 552–564.
Skeath, J. B., & Carroll, S. B. (1994). The Achaete-Scute complex - generation of cellular-pattern and fate within the Drosophila nervous-system. FASEB Journal, 8(10), 714–721.
Wapinski, O. L., Vierbuchen, T., Qu, K., Lee, Q. Y., Chanda, S., Fuentes, D. R., Giresi, P. G., Ng, Y. H., Marro, S., Neff, N. F., Drechsel, D., Martynoga, B., Castro, D. S., Webb, A. E., Südhof, T. C., Brunet, A., Guillemot, F., Chang, H. Y., & Wernig, M. (2013). Hierarchical mechanisms for direct reprogramming of fibroblasts to neurons. Cell, 155(3), 621–635.
Hahn, M.A., et al., (2019) Reprogramming of DNA methylation at NEUROD2-bound sequences during cortical neuron differentiation. Science Advances. 5(10).
Kulis, M., Queirós, A. C., Beekman, R., & Martín-Subero, J. I. (2013). Intragenic DNA methylation in transcriptional regulation, normal differentiation and cancer. Biochimica et Biophysica Acta, 1829(11), 1161–1174.
Zhu, H., Wang, G. H., & Qian, J. (2016). Transcription factors as readers and effectors of DNA methylation. Nature Reviews Genetics, 17(9), 551–565.
Acknowledgments
This work was supported by Samsung Science and Technology Foundation Project SSTF-BA1601-13.
Accession Codes
All Whole Genome Bisulfite sequencing (WGBS), mRNA-seq and ChIP-seq data for iPSC and NPC have been submitted to the Gene Expression Omnibus (GEO) under accession GSE156723.
Author information
Authors and Affiliations
Contributions
W.-Y.C. and Y.-J.K. conceived and designed the study. W.-Y.C., A.J.L and J.L. produced and processed next generation sequencing data and W.-Y.C. analyzed the data. A.-N.C. performed cellular work for iPSC maintaining and differentiation. I.-K.J. and S.-W.C. contributed advice in experiment and data analysis. W.-Y.C, J.-H.H. and L.K.K wrote the manuscript. L.K.K and Y.-J.K. supervised the project.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Fig. 1.
Characterization of function of genes according to localization of CGIs. (A-D) Gene Ontology analysis of genes containing hypermethylated promoter CGIs (pCGIs), hypomethylated pCGIs, hypermethylated intragenic CGIs (iCGIs) and hypomethylated iCGIs. The p-values were calculated using the Benjamini–Hochberg method in DAVID. Supplementary Table 1. The list of unmethylated iCGIs which were activated by neural differentiation (PDF 85 kb) (PDF 85 kb)
Rights and permissions
About this article
Cite this article
Choi, WY., Hwang, JH., Cho, AN. et al. DNA Methylation of Intragenic CpG Islands are Required for Differentiation from iPSC to NPC. Stem Cell Rev and Rep 16, 1316–1327 (2020). https://doi.org/10.1007/s12015-020-10041-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-020-10041-6