Abstract
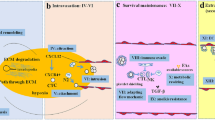
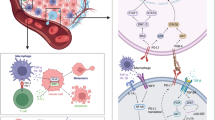
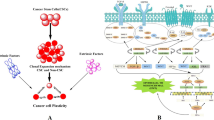
A tumor represents a highly intricate tissue entity, characterized by an exceptionally complex microenvironment that starkly contrasts with the typical physiological surroundings of healthy tissues. Within this tumor microenvironment (TME), every component and factor assume paramount importance in the progression of malignancy and exerts a pivotal influence on a patient’s clinical outcome. One of the remarkable aspects of the TME is its remarkable heterogeneity, not only across different types of cancers but even within the same histological category of tumors. In-depth research has illuminated the intricate interplay between specific immune cells and molecules and the dynamic characteristics of the TME. Recent investigations have yielded compelling evidence that several mutations harbored by tumor cells possess the capacity to instigate substantial alterations in the TME. These mutations, often acting as drivers of tumorigenesis, can orchestrate a cascade of events that remodel the TME, thereby influencing crucial aspects of cancer behavior, including its invasiveness, immune evasion, and response to therapies. It is within this nuanced context that the present study endeavors to provide a concise yet comprehensive summary of how specific mutations, within the genetic landscape of cancer cells, can instigate profound changes in TME features. By elucidating the intricate relationship between genetic mutations and the TME, this research aims to contribute to a deeper understanding of cancer biology. Ultimately, the knowledge gained from this study holds the potential to inform the development of more targeted and effective treatments, thereby offering new hope to patients grappling with the complexities of cancer.

Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- TME:
-
Tumor microenvironment
- ECM:
-
Extracellular matrix
- TAMs:
-
Tumor-associated macrophages
- TILs:
-
Tumor-infiltrating lymphocytes
- MMP:
-
Matrix metalloproteinase
- CTLA-4:
-
Cytotoxic T lymphocyte-associated protein 4
- CIMP:
-
CpG island methylator phenotype
- 2-HG:
-
2-hydroxygluratate
- HGSC:
-
High-grade serous tubo-ovarian cancer
- FTE:
-
Fallopian tube epithelium
- g-MDSC:
-
Granulocytic myeloid-derived suppressor cells
- m-MDSC:
-
Monocytic myeloid-derived suppressor cells
- NSCLC:
-
Non-small-cell lung carcinoma
- CNAs:
-
Copy number abnormalities
- HIF1:
-
Hypoxia-responsive factor
- DAG:
-
Diacylglycerol
- LGGs:
-
Lower-grade glioma
- SDC2:
-
Syndecan-2
- GAG:
-
Glycosaminoglycan
- HS:
-
Heparin sulfate
- FGL1:
-
Fibrinogen-like protein 1
- SCCE:
-
Small Cell carcinoma of the esophagus
- VEGF:
-
Vascular endothelial growth factor
- RCC:
-
Renal cell carcinoma
- Tregs:
-
Regulatory T cells
- MDSCs:
-
Myeloid-derived suppressor cells
- MCT1:
-
Monocarboxylate transporter 1
References
Tabrez, S., Khan, A. U., Hoque, M., Suhail, M., Khan, M. I., & Zughaibi, T. A. (2022). Investigating the anticancer efficacy of biogenic synthesized MgONPs: An in vitro analysis. Frontiers in Chemistry, 10, 970193.
Alafaleq, N. O., Zughaibi, T. A., Jabir, N. R., Khan, A. U., Khan, M. S., & Tabrez, S. (2023). Biogenic Synthesis of Cu-Mn Bimetallic Nanoparticles Using Pumpkin Seeds Extract and Their Characterization and Anticancer Efficacy. Nanomaterials (Basel, Switzerland), 13, 7.
Ahmad, I., Hoque, M., Alam, S. S. M., Zughaibi, T. A., & Tabrez, S. (2023). Curcumin and Plumbagin Synergistically Target the PI3K/Akt/mTOR Pathway: A Prospective Role in Cancer Treatment. International Journal of Molecular Sciences, 24(7), 6651.
Hanahan, D., & Coussens, L. M. (2012). Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell, 21(3), 309–322.
Ebos, J. M., & Kerbel, R. S. (2011). Antiangiogenic therapy: impact on invasion, disease progression, and metastasis. Nature reviews Clinical oncology, 8(4), 210–221.
Segovia-Mendoza, M., & Morales-Montor, J. (2019). Immune Tumor Microenvironment in Breast Cancer and the Participation of Estrogen and Its Receptors in Cancer Physiopathology. Frontiers in immunology, 10, 348.
Anderson, N. M., & Simon, M. C. (2020). The tumor microenvironment. Current Biology, 30(16), R921–r5.
Peña-Romero, A. C., & Orenes-Piñero, E. (2022). Dual Effect of Immune Cells within Tumour Microenvironment: Pro- and Anti-Tumour Effects and Their Triggers. Cancers, 14, 7.
Ma, Y., Shurin, G. V., Peiyuan, Z., & Shurin, M. R. (2013). Dendritic cells in the cancer microenvironment. Journal of Cancer, 4(1), 36–44.
Balkwill, F. R., Capasso, M., & Hagemann, T. (2012). The tumor microenvironment at a glance. Journal of cell science, 125(Pt 23), 5591–5596.
Barzegari, A., Saeedi, N., Zarredar, H., Barar, J., & Omidi, Y. (2014). The search for a promising cell factory system for production of edible vaccine. Human Vaccines & Immunotherapeutics, 10(8), 2497–2502.
de Looff, M., de Jong, S., & Kruyt, F. A. E. (2019). Multiple Interactions Between Cancer Cells and the Tumor Microenvironment Modulate TRAIL Signaling: Implications for TRAIL Receptor Targeted Therapy. Frontiers in immunology, 10, 1530.
Ma, C., Luo, H., Cao, J., Zheng, X., Zhang, J., & Zhang, Y., et al. (2020). Identification of a Novel Tumor Microenvironment-Associated Eight-Gene Signature for Prognosis Prediction in Lung Adenocarcinoma. Frontiers in molecular biosciences, 7, 571641.
Xiao, B., Peng, J., Wang, Y., Deng, Y., Ou, Q., & Wu, X., et al. (2020). Prognostic value of tumor infiltrating lymphocytes combined with PD-L1 expression for patients with solitary colorectal cancer liver metastasis. Annals of translational medicine, 8(19), 1221.
Maimela, N. R., Liu, S., & Zhang, Y. (2019). Fates of CD8+ T cells in Tumor Microenvironment. Computational and structural biotechnology journal, 17, 1–13.
Ohue, Y., & Nishikawa, H. (2019). Regulatory T (Treg) cells in cancer: Can Treg cells be a new therapeutic target? Cancer science, 110(7), 2080–2089.
Fridman, W. H., Meylan, M., Petitprez, F., Sun, C. M., Italiano, A., & Sautès-Fridman, C. (2022). B cells and tertiary lymphoid structures as determinants of tumour immune contexture and clinical outcome. Nature reviews Clinical oncology, 19(7), 441–457.
Boutilier, A. J., & Elsawa, S. F. (2021). Macrophage Polarization States in the Tumor Microenvironment. International Journal of Molecular Science, 22, 13.
Poh, A. R., & Ernst, M. (2018). Targeting Macrophages in Cancer: From Bench to Bedside. Frontiers in Oncology, 8, 49.
Masucci, M. T., Minopoli, M., & Carriero, M. V. (2019). Tumor Associated Neutrophils. Their Role in Tumorigenesis, Metastasis, Prognosis and Therapy. Frontiers in Oncology, 9, 1146.
Tabrez, S., Khan, A. U., Mirza, A. A., Suhail, M., Jabir, N. R., & Zughaibi, T. A., et al. (2022). Biosynthesis of copper oxide nanoparticles and its therapeutic efficacy against colon cancer. Nanotechnology Reviews, 11(1), 1322–1331.
Prete, A. D., Salvi, V., Soriani, A., Laffranchi, M., Sozio, F. & Bosisio, D. et al. (2023). Dendritic cell subsets in cancer immunity and tumor antigen sensing. Cellular & molecular immunology, 20(5), 432–447.
Binnewies, M., Roberts, E. W., Kersten, K., Chan, V., Fearon, D.F. & Merad, M. (2018). Understanding the tumor immune microenvironment (TIME) for effective therapy. Nature Medicine, 24(5), 541–550.
Yu, Y. R., & Ho, P. C. (2019). Sculpting tumor microenvironment with immune system: from immunometabolism to immunoediting. Clinical and experimental immunology, 197(2), 153–160.
Quail, D. F., & Joyce, J. A. (2013). Microenvironmental regulation of tumor progression and metastasis. Nature medicine, 19(11), 1423–1437.
Kitamura, T., Qian, B. Z., & Pollard, J. W. (2015). Immune cell promotion of metastasis. Nature reviews Immunology, 15(2), 73–86.
Chaudhary, B., & Elkord, E. (2016). Regulatory T Cells in the Tumor Microenvironment and Cancer Progression: Role and Therapeutic Targeting. Vaccines., 4, 3.
Laviron, M., & Boissonnas, A. (2019). Ontogeny of Tumor-Associated Macrophages. Frontiers in immunology, 10, 1799.
Allavena, P., Sica, A., Solinas, G., Porta, C., & Mantovani, A. (2008). The inflammatory micro-environment in tumor progression: the role of tumor-associated macrophages. Critical reviews in oncology/hematology, 66(1), 1–9.
Gowd, V., Ahmad, A., Tarique, M., Suhail, M., Zughaibi, T. A., & Tabrez, S., et al. (2022). Advancement of cancer immunotherapy using nanoparticles-based nanomedicine. Seminars in cancer biology, 86(Pt 2), 624–644.
Capece, D., Fischietti, M., Verzella, D., Gaggiano, A., Cicciarelli, G., & Tessitore, A., et al. (2013). The inflammatory microenvironment in hepatocellular carcinoma: a pivotal role for tumor-associated macrophages. BioMed research international, 2013, 187204.
Wang, J., Li, D., Cang, H., & Guo, B. (2019). Crosstalk between cancer and immune cells: Role of tumor-associated macrophages in the tumor microenvironment. Cancer medicine, 8(10), 4709–4721.
Xu, L., Zou, C., Zhang, S., Chu, T. S. M., Zhang, Y., & Chen, W., et al. (2022). Reshaping the systemic tumor immune environment (STIE) and tumor immune microenvironment (TIME) to enhance immunotherapy efficacy in solid tumors. Journal of hematology & oncology, 15(1), 1–30.
Kim, H. J., Ji, Y. R., & Lee, Y. M. (2022). Crosstalk between angiogenesis and immune regulation in the tumor microenvironment. Archives of Pharmacal Research, 45(6), 401–416.
Groner, B., & von Manstein, V. (2017). Jak Stat signaling and cancer: Opportunities, benefits and side effects of targeted inhibition. Molecular and cellular endocrinology, 451, 1–14.
Mimura, K., Teh, J. L., Okayama, H., Shiraishi, K., Kua, L. F., & Koh, V., et al. (2018). PD-L1 expression is mainly regulated by interferon gamma associated with JAK-STAT pathway in gastric cancer. Cancer science, 109(1), 43–53.
Abaza, A., Idris, F. S., Shaikh, H. A., Vahora, I., Moparthi, K. P., & Al Rushaidi, M. T., et al. (2023). Programmed Cell Death Protein 1 (PD-1) and Programmed Cell Death Ligand 1 (PD-L1) Immunotherapy: A Promising Breakthrough in Cancer Therapeutics. Cureus., 15, 9.
Yenyuwadee, S., Aliazis, K., Wang, Q., Christofides, A., Shah, R. & Patsoukis, N, et al. (2022). Immune cellular components and signaling pathways in the tumor microenvironment. Seminars in cancer biology, 86(2), 187–201.
Han, Y.-L., Luo, D., Habaxi, K., Tayierjiang, J., Zhao, W., & Wang, W., et al. (2022). COL5A2 Inhibits the TGF-β and Wnt/β-Catenin signaling pathways to inhibit the invasion and metastasis of osteosarcoma. Frontiers in Oncology, 12, 813809.
Kim, H. R., Park, H. J., Son, J., Lee, J. G., Chung, K. Y., & Cho, N. H., et al. (2019). Tumor microenvironment dictates regulatory T cell phenotype: Upregulated immune checkpoints reinforce suppressive function. Journal for immunotherapy of cancer, 7(1), 339.
Leivonen, S. K., Pollari, M., Brück, O., Pellinen, T., Autio, M., & Karjalainen-Lindsberg, M. L., et al. (2019). T-cell inflamed tumor microenvironment predicts favorable prognosis in primary testicular lymphoma. Haematologica, 104(2), 338–346.
Shanehbandi, D., Zarredar, H., Asadi, M., Zafari, V., Esmaeili, S., & Seyedrezazadeh, E., et al. (2021). Anticancer impacts of Terminalia catappa extract on SW480 colorectal neoplasm cell line. Journal of Gastrointestinal Cancer, 52(1), 99–105.
Noushmehr, H., Weisenberger, D. J., Diefes, K., Phillips, H. S., Pujara, K., & Berman, B. P., et al. (2010). Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer cell, 17(5), 510–522.
Dang, L., White, D. W., Gross, S., Bennett, B. D., Bittinger, M. A., & Driggers, E. M., et al. (2009). Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature, 462(7274), 739–744.
Turcan, S., Rohle, D., Goenka, A., Walsh, L. A., Fang, F., & Yilmaz, E., et al. (2012). IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature, 483(7390), 479–483.
Kong, L. Y., Wu, A. S., Doucette, T., Wei, J., Priebe, W., & Fuller, G. N., et al. (2010). Intratumoral mediated immunosuppression is prognostic in genetically engineered murine models of glioma and correlates to immunotherapeutic responses. Clinical cancer research: an official journal of the American Association for Cancer Research, 16(23), 5722–5733.
Ceccarelli, M., Barthel, F. P., Malta, T. M., Sabedot, T. S., Salama, S. R., & Murray, B. A., et al. (2016). Molecular Profiling Reveals Biologically Discrete Subsets and Pathways of Progression in Diffuse Glioma. Cell, 164(3), 550–563.
Amankulor, N. M., Kim, Y., Arora, S., Kargl, J., Szulzewsky, F., & Hanke, M., et al. (2017). Mutant IDH1 regulates the tumor-associated immune system in gliomas. Genes & development, 31(8), 774–786.
Nazam, N., Jabir, N. R., Ahmad, I., Alharthy, S. A., Khan, M. S., & Ayub, R., et al. (2023). Phenolic Acids-Mediated Regulation of Molecular Targets in Ovarian Cancer: Current Understanding and Future Perspectives. Pharmaceuticals (Basel, Switzerland), 16, 2.
Alafaleq, N. O., Alomari, A., Khan, M. S., Shaik, G. M., Hussain, A., & Ahmed, F., et al. (2022). Anticancer potential of gold nanoparticles (AuNPs) using a battery of in vitro tests. Nanotechnology Reviews, 11(1), 3292–3304.
Peng, W., Chen, J. Q., Liu, C., Malu, S., Creasy, C., & Tetzlaff, M. T., et al. (2016). Loss of PTEN Promotes Resistance to T Cell-Mediated Immunotherapy. Cancer discovery, 6(2), 202–216.
Miao, D., Margolis, C. A., Gao, W., Voss, M. H., Li, W., & Martini, D. J, et al. (2018). Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science, 359(6377), 801–806.
Zhang, S., Iyer, S., Ran, H., Dolgalev, I., Gu, S., & Wei, W, et al. (2021). Genetically Defined, Syngeneic Organoid Platform for Developing Combination Therapies for Ovarian Cancer. Cancer discovery, 11(2), 362–383.
Wang, G., Lu, X., Dey, P., Deng, P., Wu, C. C., & Jiang, S., et al. (2016). Targeting YAP-Dependent MDSC Infiltration Impairs Tumor Progression. Cancer discovery, 6(1), 80–95.
Kapur, P., Peña-Llopis, S., Christie, A., Zhrebker, L., Pavía-Jiménez, A., & Rathmell, W. K., et al. (2013). Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. The Lancet Oncology, 14(2), 159–167.
Simiczyjew, A., Dratkiewicz, E., Mazurkiewicz, J., Ziętek, M., Matkowski, R. & Nowak, D. (2020). The Influence of Tumor Microenvironment on Immune Escape of Melanoma. International Journal of Molecular Sciences, 21(21), 8359.
Fauriat, C., Long, E. O., Ljunggren, H. G., & Bryceson, Y. T. (2010). Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood., 115(11), 2167–2176.
Zarredar, H., Farajnia, S., Ansarin, K., Baradaran, B., Aria, M., & Asadi, M. (2019). Synergistic effect of novel EGFR inhibitor AZD8931 and p38α siRNA in lung adenocarcinoma cancer cells. Anti-Cancer Agents in Medicinal Chemistry (Formerly Current Medicinal Chemistry-Anti-Cancer Agents), 19(5), 638–644.
Thorsson, V., Gibbs, D. L., Brown, S. D., Wolf, D., Bortone, D. S., & Ou Yang, T. H., et al. (2018). The Immune Landscape of Cancer. Immunity., 48(4), 812–30.
Lambrecht, B. N., & Hammad, H. (2015). The immunology of asthma. Nature Immunology, 16(1), 45–56.
Izumi, H., Yamasaki, A., Takeda, K., Kodani, M., Touge, H., & Tanaka, N., et al. (2018). Acute-phase reaction induced by zoledronate and its effect on prognosis of patients with advanced non-small cell lung cancer. Lung Cancer, 122, 200–205.
Li, J., & Stanger, B. Z. (2019). The tumor as organizer model. Science, 363(6431), 1038–1039.
Wellenstein, M. D., & de Visser, K. E. (2018). Cancer-Cell-Intrinsic Mechanisms Shaping the Tumor Immune Landscape. Immunity, 48(3), 399–416.
Drakes, M. L., & Stiff, P. J. (2018). Regulation of Ovarian Cancer Prognosis by Immune Cells in the Tumor Microenvironment. Cancers, 10, 9.
Ruffell, B., & Coussens, L. M. (2015). Macrophages and therapeutic resistance in cancer. Cancer cell, 27(4), 462–472.
Suzuki, H., Aoki, K., Chiba, K., Sato, Y., Shiozawa, Y., & Shiraishi, Y., et al. (2015). Mutational landscape and clonal architecture in grade II and III gliomas. Nature genetics, 47(5), 458–468.
Eckel-Passow, J. E., Lachance, D. H., Molinaro, A. M., Walsh, K. M., Decker, P. A., & Sicotte, H., et al. (2015). Glioma Groups Based on 1p/19q, IDH, and TERT Promoter Mutations in Tumors. The. New England journal of medicine, 372(26), 2499–2508.
Rodriguez, G. M., Galpin, K. J. C., McCloskey, C. W. & Vanderhyden, B. C. (2018). The Tumor Microenvironment of Epithelial Ovarian Cancer and Its Influence on Response to Immunotherapy. Cancers, 10(8), 242.
Färkkilä, A., Gulhan, D. C., Casado, J., Jacobson, C. A., Nguyen, H. & Kochupurakkal, B. et al. (2020). Immunogenomic profiling determines responses to combined PARP and PD-1 inhibition in ovarian cancer. Nature Communications, 11(1), 1459
Curiel, T. J., Coukos, G., Zou, L., Alvarez, X., Cheng, P., & Mottram, P., et al. (2004). Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nature Medicine, 10(9), 942–949.
Zhang, L., Conejo-Garcia, J. R., Katsaros, D., Gimotty, P. A., Massobrio, M., & Regnani, G., et al. (2003). Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. The New England journal of medicine, 348(3), 203–213.
Bronger, H., Singer, J., Windmüller, C., Reuning, U., Zech, D., & Delbridge, C., et al. (2016). CXCL9 and CXCL10 predict survival and are regulated by cyclooxygenase inhibition in advanced serous ovarian cancer. British journal of cancer, 115(5), 553–563.
Nagarsheth, N., Wicha, M. S., & Zou, W. (2017). Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nature Reviews Immunology, 17(9), 559–572.
Dangaj, D., Bruand, M., Grimm, A. J., Ronet, C., Barras, D., & Duttagupta, P. A., et al. (2019). Cooperation between Constitutive and Inducible Chemokines Enables T Cell Engraftment and Immune Attack in Solid Tumors. Cancer cell, 35(6), 885–900.
Strickland, K. C., Howitt, B. E., Shukla, S. A., Rodig, S., Ritterhouse, L. L., & Liu, J. F., et al. (2016). Association and prognostic significance of BRCA1/2-mutation status with neoantigen load, number of tumor-infiltrating lymphocytes and expression of PD-1/PD-L1 in high grade serous ovarian cancer. Oncotarget, 7(12), 13587–13598.
Clarke, B., Tinker, A. V., Lee, C. H., Subramanian, S., van de Rijn, M., & Turbin, D., et al. (2009). Intraepithelial T cells and prognosis in ovarian carcinoma: novel associations with stage, tumor type, and BRCA1 loss. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc, 22(3), 393–402.
Liu, T., Xia, Q., Zhang, H., Wang, Z., Yang, W., & Gu, X., et al. (2020). CCL5-dependent mast cell infiltration into the tumor microenvironment in clear cell renal cell carcinoma patients. Aging, 12(21), 21809–21836.
Capaci, V., Mantovani, F., & Sal, G. D. (2020). A mutant p53/Hif1α/miR-30d axis reprograms the secretory pathway promoting the release of a prometastatic secretome. Cell stress, 4(11), 261–264.
Yin, W., Jiang, X., Tan, J., Xin, Z., Zhou, Q., & Zhan, C., et al. (2020). Development and Validation of a Tumor Mutation Burden-Related Immune Prognostic Model for Lower-Grade Glioma. Frontiers in oncology, 10, 1409.
Rovira-Clavé, X., Angulo-Ibáñez, M., Noguer, O., Espel, E., & Reina, M. (2012). Syndecan-2 can promote clearance of T-cell receptor/CD3 from the cell surface. Immunology, 137(3), 214–225.
Bastos, P., Gomes, T., & Ribeiro, L. (2017). Catechol-O-Methyltransferase (COMT): An Update on Its Role in Cancer, Neurological and Cardiovascular Diseases. Reviews of Physiology, Biochemistry and Pharmacology, 173, 1–39.
Ahmed, M. E., & Falasiri, S. (2020). The Immune Microenvironment in Penile Cancer and Rationale for. Immunotherapy, 9, 10.
Nechama, M., Kwon, J., Wei, S., Kyi, A. T., Welner, R. S., & Ben-Dov, I. Z., et al. (2018). The IL-33-PIN1-IRAK-M axis is critical for type 2 immunity in IL-33-induced allergic airway inflammation. Nature communications, 9(1), 1603.
Ohta, A., Kini, R., Ohta, A., Subramanian, M., Madasu, M., & Sitkovsky, M. (2012). The development and immunosuppressive functions of CD4(+) CD25(+) FoxP3(+) regulatory T cells are under influence of the adenosine-A2A adenosine receptor pathway. Frontiers in immunology, 3, 190.
Yang, W., Chen, N., Li, L., Chen, X., Liu, X., & Zhang, Y., et al. (2020). Favorable Immune Microenvironment in Patients with EGFR and MAPK Co-Mutations. Lung Cancer (Auckland, NZ), 11, 59–71.
Ryzhov, S., Novitskiy, S. V., Goldstein, A. E., Biktasova, A., Blackburn, M. R., & Biaggioni, I., et al. (2011). Adenosinergic regulation of the expansion and immunosuppressive activity of CD11b+Gr1+ cells. Journal of immunology (Baltimore, Md: 1950), 187(11), 6120–6129.
Ouyang, L., Zhang, K., Chen, J., Wang, J. & Huang, H. (2018). Roles of platelet-derived growth factor in vascular calcification. Journal of Cellular Physiology, 233(4), 2804–2814.
Wang, X., Teng, F., Kong, L., & Yu, J. (2016). PD-L1 expression in human cancers and its association with clinical outcomes. Onco Targets & Therapy, 9, 5023–5039.
Evrard, D., Hourseau, M., Couvelard, A., Paradis, V., Gauthier, H. & Raymond, E, et al. (2020). PD-L1 expression in the microenvironment and the response to checkpoint inhibitors in head and neck squamous cell carcinoma. Oncoimmunology, 9(1), 1844403
Zhao, Q., Chen, Y. X., Wu, Q. N., Zhang, C., Liu, M., & Wang, Y. N., et al. (2020). Systematic analysis of the transcriptome in small-cell carcinoma of the oesophagus reveals its immune microenvironment. Clinical & translational immunology, 9(10), e1173.
Shi, A. P., Tang, X. Y., Xiong, Y. L., Zheng, K. F., Liu, Y. J., & Shi, X. G., et al. (2021). Immune Checkpoint LAG3 and Its Ligand FGL1 in Cancer. Frontiers in immunology, 12, 785091.
Wang, J., Sanmamed, M. F., Datar, I., Su, T. T., Ji, L., & Sun, J., et al. (2019). Fibrinogen-like Protein 1 Is a Major Immune Inhibitory Ligand of LAG-3. Cell, 176(1-2), 334–347.
Guo, M., Yuan, F., Qi, F., Sun, J., Rao, Q., & Zhao, Z., et al. (2020). Expression and clinical significance of LAG-3, FGL1, PD-L1 and CD8(+)T cells in hepatocellular carcinoma using multiplex quantitative analysis. Journal of translational medicine, 18(1), 306.
Song, L., Ye, W., Cui, Y., Lu, J., Zhang, Y., & Ding, N., et al. (2017). Ecto-5’-nucleotidase (CD73) is a biomarker for clear cell renal carcinoma stem-like cells. Oncotarget., 8(19), 31977–31992.
Chambers, A. M., Wang, J., Lupo, K. B., Yu, H., Atallah Lanman, N. M., & Matosevic, S. (2018). Adenosinergic Signaling Alters Natural Killer Cell Functional Responses. Frontiers in immunology, 9, 2533.
Arab, S., & Hadjati, J. (2019). Adenosine Blockage in Tumor Microenvironment and Improvement of Cancer Immunotherapy. Immune network, 19(4), e23.
Takayama, H., Trenn, G., & Sitkovsky, M. V. (1988). Locus of inhibitory action of cAMP-dependent protein kinase in the antigen receptor-triggered cytotoxic T lymphocyte activation pathway. The. Journal of Biological Chemistry, 263(5), 2330–2336.
Tripathi, A., Lin, E., Xie, W., Flaifel, A., Steinharter, J. A., & Gatof, E. N. S., et al. (2020). Prognostic significance and immune correlates of CD73 expression in renal cell carcinoma. Journal for immunotherapy of cancer, 8(2), e001467.
Rafii, S., Ghouzlani, A., Naji, O., Ait Ssi, S., Kandoussi, S., & Lakhdar, A., et al. (2023). A(2A)R as a Prognostic Marker and a Potential Immunotherapy Target in Human Glioma. International Journal of Molecular Science, 24, 7.
Ernens, I., Bousquenaud, M., Lenoir, B., Devaux, Y., & Wagner, D. R. (2015). Adenosine stimulates angiogenesis by up-regulating production of thrombospondin-1 by macrophages. Journal of leukocyte biology, 97(1), 9–18.
Jin, M., Cao, W., Chen, B., Xiong, M., & Cao, G. (2022). Tumor-Derived Lactate Creates a Favorable Niche for Tumor via Supplying Energy Source for Tumor and Modulating the Tumor Microenvironment. Frontiers in cell and developmental biology, 10, 808859.
Romero-Garcia, S., Moreno-Altamirano, M. M., Prado-Garcia, H., & Sánchez-García, F. J. (2016). Lactate Contribution to the Tumor Microenvironment: Mechanisms, Effects on Immune Cells and Therapeutic Relevance. Frontiers in immunology, 7, 52.
Bola, B. M., Chadwick, A. L., Michopoulos, F., Blount, K. G., Telfer, B. A., & Williams, K. J., et al. (2014). Inhibition of monocarboxylate transporter-1 (MCT1) by AZD3965 enhances radiosensitivity by reducing lactate transport. Molecular cancer therapeutics, 13(12), 2805–2816.
Li, B., Yang, Q., Li, Z., Xu, Z., Sun, S., & Wu, Q., et al. (2020). Expression of Monocarboxylate Transporter 1 in Immunosuppressive Macrophages Is Associated With the Poor Prognosis in Breast Cancer. Frontiers in Oncology, 10, 574787.
Tiainen, S., Tumelius, R., Rilla, K., Hämäläinen, K., Tammi, M., & Tammi, R., et al. (2015). High numbers of macrophages, especially M2-like (CD163-positive), correlate with hyaluronan accumulation and poor outcome in breast cancer. Histopathology. 66(6), 873–883.
Kaleem, M., Dalhat, M. H., Azmi, L., Asar, T. O., Ahmad, W., & Alghanmi, M., et al. (2022). An Insight into Molecular Targets of Breast Cancer Brain Metastasis. International Journal of Molecular Sciences, 23(19), 11687.
Funding
This study was funded by Tuberculosis and Lung Disease Research Center, Tabriz University of Medical Science, Tabriz, Iran.
Author information
Authors and Affiliations
Contributions
Study Design: A.C., D.S.; Data Collection: M.A., H.Z.; Statistical Analysis: M.A., V.Z.; Data Interpretation: H.Z., Z.S.; Manuscript Preparation: Z.S., H.S.; Literature Search: A.C., D.S.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Asadi, M., Zarredar, H., Zafari, V. et al. Immune Features of Tumor Microenvironment: A Genetic Spotlight. Cell Biochem Biophys 82, 107–118 (2024). https://doi.org/10.1007/s12013-023-01192-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-023-01192-7