Abstract
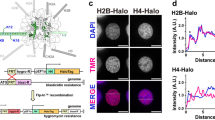
Mitoxantrone (MTX), a choice of drug in cancer chemotherapeutic regime, is a potent and less toxic among anthracycline class of drugs. Here, we study the molecular interaction of MTX, with histone and its acetylation dynamics. Its binding with histone core protein was predicted with CD and UV–visible spectroscopic techniques. The MTX–protein complex resulted in the impediment of the histone acetyltransferase (HAT) activity in a dose dependent manner on MTX binding. Interestingly, the concentration dependent reduction in acetylated state of specific lysines K9/K14 was also observed on MTX treatment in vivo. The molecular distance r, between donor (histone H3) and acceptor (MTX) was estimated using Förster’s theory of non-radiation energy transfer and the detailed binding phenomenon was expounded. MTX binding site near N-terminal lysines is characterized with an association constant of the order of 104. The positive thermodynamic values of both ∆H° and ∆S° were suggestive that the hydrophobic interactions dominate in MTX–protein binding. The binding site allocation predicted by computational modeling placed the drug molecule near N-terminal lysine K9 and K14 of histone H3, and corroborate with the thermodynamic interaction model. The study establishes that MTX–histone interaction affects protein acetylation state and also provided a mechanistic model for its binding. Hence, MTX interaction may affect chromatin structure and implicates its role in transcriptional regulation at epigenetic level.






Similar content being viewed by others
References
Myers, C. E., Mimnaugh, E. G., Yeh, G. C., & Stone, B. K. (1988). In J. W. Lown (Ed.), Anthracycline and anthracenedione-based anticancer agents (p. 527). Amsterdam: Elsevier Science Publishers B.V.
Van Holde, K. E. (1988). Chromatin (p. 497). New York: Springer-Verlag.
Strahl, B. D., & Allis, C. D. (2000). The language of covalent histone modifications. Nature, 403, 41–45.
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F., & Richmond, T. J. (1997). Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature, 389, 251–260.
Luger, K., & Richmond, T. J. (1998). The histone tails of the nucleosome. Current Opinion in Genetics and Development, 8, 140–146.
Gray, S. G., & The, B. T. (2001). Histone acetylation/deacetylation and cancer: An “open” & “shut” case? Current Molecular Medicine, 1, 401–429.
Gregory, P. D., Wagner, K., & Hörz, W. (2001). Histone acetylation and chromatin remodeling. Experimental Cell Research, 265, 195–202.
Förster, T., & Sinanoglu, O. (eds.). (1996) Modern quantum chemistry (Vol. 3, p. 93). New York: Academic Press.
Schreiber, E., Matthias, P., Muller, M. M., & Schaffner, W. (1989). Rapid detection of octamer binding proteins with ‘mini-extracts’, prepared from a small number of cells. Nucleic Acids Research, 17, 6419–6421.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Analytical Biochemistry, 72, 248–254.
Fischle, W. (2005). In nucleo enzymatic assays for the identification and characterization of histone modifying activities. Methods, 36, 362–367.
Jones, G., Willett, P., & Glen, V. J. (1995). Molecular recognition of receptor sites using a genetic algorithm with a description of desolvation. Molecular Biology, 245, 43–53.
Wang, R., Lai, L., & Wang, S. J. (2002). Further development and validation of empirical scoring functions for structure-based binding affinity prediction. Journal of Computer-Aided Molecular Design, 16, 11–26.
Kuntz, I. D., Blaney, J. M., Oatley, S. J., Langridge, R., & Ferrin, T. E. (1982). Geometric approach to macromolecule–ligand interactions. Journal of Molecular Biology, 161, 269–288.
Munishkina, L. A., Fink, A. L., & Uversky, V. N. (2004). Conformational prerequisites for formation of amyloid fibrils from histones. Journal of Molecular Biology, 342, 1305–1324.
Miles, J. L., Morey, E., Crain, F., Gross, S., San Julian, J. S., & Canady, W. J. (1962). Inhibition of alpha-chymotrypsin by diethyl ether and certain alcohols: A new type of competitive inhibition. The Journal of Biological Chemistry, 237, 1319–1322.
Hu, Y. J., Liu, Y., Wang, J. B., Xiao, X. H., & Qu, S. S. (2004). Study of the interaction between monoammonium glycyrrhizinate and bovine serum albumin. Journal of Pharmaceutical and Biomedical Analysis, 36, 915–919.
Cui, F. L., Fan, J., Li, J. P., & Hu, Z. D. (2004). Interactions between 1-benzoyl-4-p-chlorophenyl thiosemicarbazide and serum albumin: Investigation by fluorescence spectroscopy. Bioorganic and Medicinal Chemistry, 12, 151–157.
Lakowicz, J. R., & Weber, G. (1973). Quenching of fluorescence by oxygen. Probe for structural fluctuations in macromolecules. Biochemistry, 12, 4161–4170.
Chen, Y., Elangovan, M., & Periasamy, A. (2005). FRET data analysis: The algorithm. In A. Periasamy & R. N. Day (Eds.), Molecular imaging: FRET microscopy and spectroscopy (p. 126). Oxford: Oxford University Press.
Luebben, W. R., Sharma. N., & Nyborg, J. K. (2010). Nucleosome eviction and activated transcription require p300 acetylation of histone H3 lysine 14. Proceedings of the National Academy of Sciences of the United States of America, 107, 19254–19259.
Nishida, H., Suzuki, T., Kondo, S., Miura, H., Fujimura, Y., & Hayashizaki, Y. (2006). Histone H3 acetylated at lysine 9 in promoter is associated with low nucleosome density in the vicinity of transcription start site in human cell. Chromosome Research, 14, 203–211.
Fuks, F. (2005). DNA methylation and histone modifications: Teaming up to silence genes. Current Opinion in Genetics and Development, 15, 490–495.
Gao, H., Lei, L., Liu, J., Qin, K., Chen, X., & Hu, Z. (2004). The study on the interaction between human serum albumin and a new reagent with antitumour activity by spectrophotometric methods. Journal of Photochemistry and Photobiology A, 167, 213–221.
Khan, S. N., Islam, B., Yennamalli, R., Sultan, A., Subbarao, N., & Khan, A. U. (2008). Interaction of mitoxantrone with human serum albumin: Spectroscopic and molecular modeling studies. European Journal of Pharmaceutical Sciences, 35, 371–382.
Khan, S. N., Danishuddin, M., & Khan, A. U. (2010). Inhibition of transcription factor assembly and structural stability on mitoxantrone binding with DNA. Bioscience Reports, 30, 331–340.
Ross, P. D., & Subramanian, S. (1981). Thermodynamics of protein association reactions. Forces contributing to stability. Biochemistry, 20, 3096–3102.
Khan, S. N., Islam, B., Yennamalli, R., Sultan, A., Subbarao, N., & Khan, A. U. (2008). Characterization of doxorubicin binding site and drug induced alteration in the functionally important structural state of oxyhemoglobin. Journal of Pharmaceutical and Biomedical Analysis, 48, 1096–1104.
Mahlknecht, U., & Hoelzer, D. (2000). Histone acetylation modifiers in the pathogenesis of malignant disease. Molecular Medicine, 6, 623–644.
Khan, S. N., & Khan, A. U. (2010). Role of histone acetylation in cell physiology and diseases: An update. Clinica Chimica Acta, 411, 1401–1411.
Bi, S., Ding, L., Tian, Y., Song, D., Zhou, X., Liu, X., et al. (2004). Investigation of the interaction between flavonoids and human serum albumin. Journal of Molecular Structure, 703, 37–45.
Kang, J., Liu, Y., Xie, M. X., Li, S., Jiang, M., & Wang, Y. D. (2004). Interactions of human serum albumin with chlorogenic acid and ferulic acid. Biochimica et Biophysica Acta, 1674, 205–214.
Mahesha, H. G., Singh, S. A., Srinivasan, N., & Rao, A. G. (2006). A spectroscopic study of the interaction of isoflavones with human serum albumin. The FEBS Journal, 273, 451–467.
Acknowledgments
We are thankful to the central instrumentation facility (CIF) of IBU. This work was supported by the CSIR sanction no. 37(1209)04 EMR II. Author is also grateful to Prof Alok Bhattacharya, JNU, New Delhi for his support and critical discussion.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.

Rights and permissions
About this article
Cite this article
Khan, S.N., Yennamalli, R., Subbarao, N. et al. Mitoxantrone Induced Impediment of Histone Acetylation and Structural Flexibility of the Protein. Cell Biochem Biophys 60, 209–218 (2011). https://doi.org/10.1007/s12013-010-9141-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-010-9141-9