Abstract
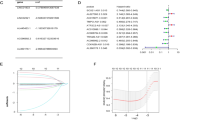
Cuproptosis, a newly discovered form of programmed cell death, relies on mitochondrial respiration, the chain of which has been found to be altered in ovarian cancer (OC). The current work probed into the effects of Cuproptosis on the prognosis, immune microenvironment and therapeutic response of OC based on Cuproptosis-related lncRNAs. Data on OC gene expression and clinical characteristics were collected from TCGA, ICGC and GEO databases, and mRNA and lncRNA were distinguished. Cuproptosis-related lncRNAs were screened for consensus clustering analysis. Differentially expressed lncRNAs (DElncRNAs) were identified between clusters, and least absolute shrinkage and selection operator (LASSO) and Cox regression analysis were performed to establish a prognostic signature. Its potential value in OC was evaluated by Gene Set Enrichment Analysis (GSEA), tumor cell mutation and immune microenvironment analysis, and response to immunotherapy and antineoplastic drugs. According to the classification scheme of Cuproptosis-related lncRNAs, OC was divided into four molecular subtypes, which were different in survival time, immune characteristics and somatic mutation. The prognostic signature between subtypes included 10 lncRNAs, which were significantly correlated with the prognosis, immune microenvironment related indexes, the expression of immune checkpoint molecules and the sensitivity of antineoplastic drug Paclitaxel and Gefitinib of OC. We examined the expression of ten LncRNAs in OC cell lines and found that LINC00189, ZFHX4-AS1, RPS6KA2-IT1 and C9orf106 were expressed elevated in OC cell lines, and LINC00861, LINC00582, DEPDC1-AS1, LINC01556, LEMD1-AS1, TYMSOS expression was decreased in OC cell lines. The results of CCK8 showed that the cell viability of OC cells decreased after inhibition of C9orf106, whereas the cell viability of OC cells increased after inhibition of LEMD1-AS1. This work revealed new Cuproptosis-related lncRNA molecular subtypes exhibiting tumor microenvironment (TME) heterogeneity for OC and proposed a prognostic signature that may have benefits in understanding the prognosis, pathological features and immune microenvironment of OC patients.









Similar content being viewed by others
Data Availability
The datasets generated and/or analyzed during the current study are available in the [GSE102073] repository, [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102073].
Abbreviations
- OC:
-
Ovarian cancer
- DElncRNAs:
-
Differentially expressed Long non-coding RNAs
- LASSO:
-
Least absolute shrinkage and selection operator
- GSEA:
-
Gene Set Enrichment Analysis
- RNA-seq:
-
RNA sequencing
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
Gene Expression Omnibus
- FPKM:
-
Fragments per kilobase of exon model per million mapped fragments
- SNV:
-
Single nucleotide variation
- CNV:
-
Copy number variation
- ICGC:
-
International Cancer Genome Consortium
- TPM:
-
Transcripts per kilobase
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the curve
- TME:
-
Tumor microenvironment
- Fges:
-
Functional gene expression signatures
- MATH:
-
Mutant-allele tumor heterogeneity
- TMB:
-
Tumor mutational burden
- HRD:
-
Homologous recombination DNA repair deficiency
- GSEA:
-
Gene Set Enrichment Analysis
- TIDE:
-
Tumor Immune Dysfunction and Exclusion
- ICI:
-
Immune checkpoint inhibitor
- GDSC:
-
Genome of Drug Sensitivity in Cancer
- IC50:
-
Half maximal inhibitory concentration
- MDSCs:
-
Myeloid-derived suppressor cells
- CAFs:
-
Cancer-associated fibroblasts
- TAMs:
-
Tumor-associated macrophages
References
Kurnit KC, Fleming GF, Lengyel E (2021) Updates and new options in advanced epithelial ovarian cancer treatment. Obstet Gynecol 137(1):108–121
Ovejero-Sanchez M, Gonzalez-Sarmiento R, Herrero AB (2023) DNA damage response alterations in ovarian cancer: from molecular mechanisms to therapeutic opportunities. Cancers (Basel) 15(2):448
Battistini C, Cavallaro U (2023) Patient-derived in vitro models of ovarian cancer: powerful tools to explore the biology of the disease and develop personalized treatments. Cancers (Basel) 15(2):368
van Zyl B, Tang D, Bowden NA (2018) Biomarkers of platinum resistance in ovarian cancer: what can we use to improve treatment. Endocr Relat Cancer 25(5):R303–R318
Marrelli D, Ansaloni L, Federici O, Asero S, Carbone L, Marano L et al (2022) Cytoreductive surgery (CRS) and HIPEC for advanced ovarian cancer with peritoneal metastases: italian PSM oncoteam evidence and study purposes. Cancers (Basel) 14(23):6010
Shukla P, Singh KK (2021) The mitochondrial landscape of ovarian cancer: emerging insights. Carcinogenesis 42(5):663–671
De Rasmo D, Cormio A, Cormio G, Signorile A (2023) Ovarian cancer: a landscape of mitochondria with emphasis on mitochondrial dynamics. Int J Mol Sci; 24(2).
Tsvetkov P, Coy S, Petrova B, Dreishpoon M, Verma A, Abdusamad M et al (2022) Copper induces cell death by targeting lipoylated TCA cycle proteins. Science 375(6586):1254–1261
Huang X, Zhou S, Toth J, Hajdu A (2022) Cuproptosis-related gene index: a predictor for pancreatic cancer prognosis, immunotherapy efficacy, and chemosensitivity. Front Immunol 13:978865
Chen B, Zhou X, Yang L, Zhou H, Meng M, Zhang L et al (2022) A Cuproptosis Activation Scoring model predicts neoplasm-immunity interactions and personalized treatments in glioma. Comput Biol Med 148:105924
Tong X, Tang R, Xiao M, Xu J, Wang W, Zhang B et al (2022) Targeting cell death pathways for cancer therapy: recent developments in necroptosis, pyroptosis, ferroptosis, and cuproptosis research. J Hematol Oncol 15(1):174
Yoshihara K, Shahmoradgoli M, Martinez E, Vegesna R, Kim H, Torres-Garcia W et al (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612
Bagaev A, Kotlov N, Nomie K, Svekolkin V, Gafurov A, Isaeva O et al (2021) Conserved pan-cancer microenvironment subtypes predict response to immunotherapy. Cancer Cell 39(6):845-65e7
Xu L, Deng C, Pang B, Zhang X, Liu W, Liao G et al (2018) TIP: a web server for resolving tumor immunophenotype profiling. Cancer Res 78(23):6575–6580
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457
Mroz EA, Rocco JW (2013) MATH, a novel measure of intratumor genetic heterogeneity, is high in poor-outcome classes of head and neck squamous cell carcinoma. Oral Oncol 49(3):211–215
Stover EH, Fuh K, Konstantinopoulos PA, Matulonis UA, Liu JF (2020) Clinical assays for assessment of homologous recombination DNA repair deficiency. Gynecol Oncol 159(3):887–898
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X et al (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558
Auslander N, Zhang G, Lee JS, Frederick DT, Miao B, Moll T et al (2018) Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat Med 24(10):1545–1549
Varanasi SK, Kaech SM, Bui JD (2022) SnapShot: cancer immunoediting. Cell 185(21):4038-e1
van den Bulk J, Verdegaal EM, de Miranda NF (2018) Cancer immunotherapy: broadening the scope of targetable tumours. Open Biol 8(6):180037
Tang D, Chen X, Kroemer G (2022) Cuproptosis: a copper-triggered modality of mitochondrial cell death. Cell Res 32(5):417–8
Liu H, Tang T (2022) Pan-cancer genetic analysis of cuproptosis and copper metabolism-related gene set. Front Oncol 12:952290
De Leo A, Santini D, Ceccarelli C, Santandrea G, Palicelli A, Acquaviva G et al (2021) What is new on ovarian carcinoma: integrated morphologic and molecular analysis following the new 2020 World Health Organization classification of female genital tumors. Diagnostics (Basel) 11(4):697
Palmirotta R, Silvestris E, D’Oronzo S, Cardascia A, Silvestris F (2017) Ovarian cancer: novel molecular aspects for clinical assessment. Crit Rev Oncol Hematol 117:12–29
Yu F, Quan F, Xu J, Zhang Y, Xie Y, Zhang J et al (2019) Breast cancer prognosis signature: linking risk stratification to disease subtypes. Brief Bioinform 20(6):2130–40
Radu MR, Pradatu A, Duica F, Micu R, Cretoiu SM, Suciu N et al (2021) Ovarian cancer: biomarkers and targeted therapy. Biomedicines 9(6):693
Pan Y, Yu Y, Wang X, Zhang T (2020) Tumor-associated macrophages in tumor immunity. Front Immunol 11:583084
Reina-Campos M, Scharping NE, Goldrath AW (2021) CD8(+) T cell metabolism in infection and cancer. Nat Rev Immunol 21(11):718–38
Hu X, Bian C, Zhao X, Yi T (2022) Efficacy evaluation of multi-immunotherapy in ovarian cancer: from bench to bed. Front Immunol 13:1034903
Marth C, Wieser V, Tsibulak I, Zeimet AG (2019) Immunotherapy in ovarian cancer: fake news or the real deal? Int J Gynecol Cancer 29(1):201–11
Zhang Y, Zhang X, Zhu H, Liu Y, Cao J, Li D et al (2020) Identification of potential prognostic long non-coding RNA biomarkers for predicting recurrence in patients with cervical cancer. Cancer Manag Res 12:719–30
Gao Q, Shi Y, Sun Y, Zhou S, Liu Z, Sun X et al (2023) Identification and verification of aging-related lncRNAs for prognosis prediction and immune microenvironment in patients with head and neck squamous carcinoma. Oncol Res 31(1):35–61
Wang X, Wang Y, Sun F, Xu Y, Zhang Z, Yang C et al (2022) Novel LncRNA ZFHX4-AS1 as a potential prognostic biomarker that affects the immune microenvironment in ovarian cancer. Front Oncol 12:945518
Cheng Y, Wang X, Qi P, Liu C, Wang S, Wan Q et al (2021) Tumor microenvironmental competitive endogenous RNA network and immune cells act as robust prognostic predictor of acute myeloid leukemia. Front Oncol 11:584884
Guo R, Qin Y (2020) LEMD1-AS1 suppresses ovarian cancer progression through regulating miR-183-5p/TP53 axis. Onco Targets Ther 13:7387–98
Zhang Z, Wang J, Zhang X, Ran B, Wen J, Zhang H (2023) TYMSOS-miR-101-3p-NETO2 axis promotes osteosarcoma progression. Mol Cell Probes 67:101887
Gu Y, Wan C, Zhou G, Zhu J, Shi Z, Zhuang Z (2021) TYMSOS drives the proliferation, migration, and invasion of gastric cancer cells by regulating ZNF703 via sponging miR-4739. Cell Biol Int 45(8):1710–9
Funding
This research was supported by Guangxi Medical high-level and sub backbone talents training “139” program (No. G202003005), the Key Research and Development Program of Guangxi (No. GuikeAB18126056) and Liuzhou Science and Technology Bureau Project (No. 2018BJ10301, No. 2019BJ10604), Guangxi Science and Technology Plan Project (Guangxi Clinical Research Center for Obstetrics and Gynecology, GuiKe AD22035223) and Guangxi Self-Financing Research Program of Guangxi Region Health Commission (No. Z20210518, No. Z-B20221595).
Author information
Authors and Affiliations
Contributions
All authors contributed to this present work: [NL] and [KY] designed the study; [DLH] and [SL] acquired the data; [DYZ] drafted the manuscript; [JJL] and [LF] revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, N., Yu, K., Huang, D. et al. Molecular Characterization of Cuproptosis-related lncRNAs: Defining Molecular Subtypes and a Prognostic Signature of Ovarian Cancer. Biol Trace Elem Res 202, 1428–1445 (2024). https://doi.org/10.1007/s12011-023-03780-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12011-023-03780-3