Abstract
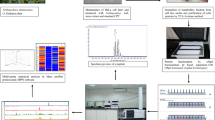
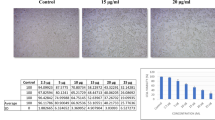
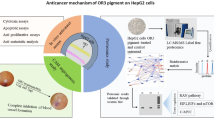
Search for ideal compounds with known pathways of anticancer mechanism is still a priority research focus for cancer, as it continues to be a major health challenge across the globe. Hence, in the present study, anticancer potential of a yellow pigment fraction, OR3, isolated from Streptomyces coelicolor JUACT03 was assessed on the breast cancer cell line MCF-7. TLC-fractionated OR3 pigment was subjected to HPLC and GC–MS analysis for characterization and identification of the bioactive component. MCF-7 cells were treated with IC50 concentration of OR3 and the molecular alterations were analyzed using mass spectrometry-based quantitative proteomic analysis. Bioinformatics tools such as STRING analysis and Ingenuity Pathway Analysis were performed to analyze proteomics data and to identify dysregulated signaling pathways. As per our obtained data, OR3 treatment decreased cell proliferation and induced apoptotic cell death due to significant dysregulation of protein expressions in MCF-7 cells. Altered expression included the ribosomal, mRNA processing and vesicle-mediated transport proteins as a result of OR3 treatment. Downregulation of MAPK proteins, NFkB, and estradiol signaling was identified in OR3-treated MCF-7 cells. Mainly eIF2, mTOR, and eIF4 signaling pathways were altered in OR3-treated cells. GC–MS data indicated the presence of novel compounds in OR3 fraction. It can be concluded that OR3 exhibits potent anticancer activity on the breast cancer cells mainly through altering the expression and affecting the signaling proteins which are involved in different cell proliferation/apoptotic pathways thereby causing inhibition of cancer cell proliferation, survival and metastasis.






Similar content being viewed by others
Data Availability
Raw MS files obtained in this experiment have been uploaded to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org), through the PRIDE partner repository with the dataset identifier PXD032999. Other datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Duggan, C., Dvaladze, A., Rositch, A. F., Ginsburg, O., Yip, C. H., Susan Horton, S., Rodriguez, R. C., Eniu, A., Mutebi, M., Bourque, J. M., Masood, M. S., Saldana, K. U., Cabanes, A., Carlson, R. W., Gralow, J. R., & Anderson, B. O. (2020). The breast health global initiative 2018 global summit on improving breast healthcare through resource-stratified phased implementation: Methods and overview. Cancer, 126(10), 2339–2352. https://doi.org/10.1002/cncr.32891
Fitzmaurice, C., Akinyemiju, T. F., Lami, F. H., et al. (2018). Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2016: A systematic analysis for the global burden of disease study. JAMA Oncology, 4(11), 1553–1568. https://doi.org/10.1001/jamaoncol.2018.2706
Allemani, C., Matsuda, T., Di Carlo, V., Harewood, R., Matz, M., Niksic, M., Bonaventure, A., Valkov, M., Johnson, C. J., Esteve, J., Ogunbiyi, O. J., Silva, G. A., Chen, W. Q., Eser, S., Engholm, G., Stiller, C. A., Monnereau, A., Woods, R. R., Visser, O., … MP. (2018). Global surveillance of trends in cancer survival 2000–14 (CONCORD-3): Analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries. Lancet, 391(10125), 1023–1075. https://doi.org/10.1016/s0140-6736(17)33326-3
Siegel, R. L., Miller, K. D., & Jemal, A. (2020). Cancer statistics. CA: A Cancer Journal for Clinicians, 70(1), 7–30. https://doi.org/10.3322/caac.21590
Law, J. W. F., Law, L. N. S., Letchumanan, V., Tan, L.T.-H., Wong, S. H., Chan, K.-G., Ab Mutalib, N.-S., & Lee, L.-H. (2020). Anticancer drug discovery from microbial sources: The unique mangrove streptomycetes. Molecules, 25(22), 5365. https://doi.org/10.3390/molecules25225365
Berdy, J. (2005). Bioactive microbial metabolites. Journal of Antibiotics, 58(1), 1–26. https://doi.org/10.1038/ja.2005.1
Harvey, A., Edrada-Ebel, R., & Quinn, R. (2015). The re-emergence of natural products for drug discovery in the genomics era. Nature Reviews. Drug Discovery, 14, 111–129. https://doi.org/10.1038/nrd4510
Newman, D. J., & Cragg, G. M. (2012). Natural products as sources of new drugs over the 30 years from 1981 to 2010. Journal of Natural Products., 75(3), 311–335. https://doi.org/10.1021/np200906s
Solanki, R., Khanna, M., & Lal, R. (2008). Bioactive compounds from marine actinomycetes. Indian J Microbiol., 48(4), 410–431. https://doi.org/10.1007/2Fs12088-008-0052-z
Olano, C., Mendez, C., & Salas, J. A. (2009). Antitumor compounds from marine actinomycetes. Marine Drugs, 7(2), 210–248. https://doi.org/10.3390/md7020210
Huang, M., Lu, J. J., & Ding, J. (2021). Natural products in cancer therapy: Past, present and future. Natural Products and Bioprospecting, 11, 5–13. https://doi.org/10.1007/s13659-020-00293-7
Zhang, R., Li, X., Zhang, X., Qina, H., & Xiao, W. (2020). Machine learning approaches for elucidating the biological effects of natural products. Natural Product Reports., 38,. https://doi.org/10.1039/D0NP00043D
Bruno, W. J., Socci, N. D., & Halpern, A. L. (2000). Weighted neighbor joining: A likelihood-based approach to distance-based phylogeny reconstruction. Molecular Biology and Evolution, 17(1), 189–197. https://doi.org/10.1093/oxfordjournals.molbev.a026231
Prashanthi, K., Suryan, S., Varalakshmi, K. N. (2015). In vitro anticancer property of yellow pigment from Streptomyces griseoaurantiacus JUACT 01. Brazilian Archives of Biology and Technology, 58(6). https://doi.org/10.1590/S1516-89132015060271.
Mosmann, T. (1983). Rapid colourimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. Journal of Immunological Methods, 65(1–2), 55–63. https://doi.org/10.1016/0022-1759(83)90303-4
Gondkar, K., Sathe, G., Joshi, N., Nair, B., Pandey, A., & Kumar, P. (2021). Integrated proteomic and phosphoproteomics analysis of DKK3 signaling reveals activated kinase in the most aggressive gallbladder cancer. Cells, 10(3), 511. https://doi.org/10.3390/cells10030511
Babu, N., Patil, S., Mohan, S. V., Subbannayya, T., Advani, J., Datta, K. K., Rajagopalan, P., Bhat, F. A., Sidransky, D., Gowda, H., & Chatterjee, A. (2021). Signaling alterations in oral keratinocytes in response to shisha and crude tobacco extract. Journal of Oral Pathology and Medicine, 50(5), 459–469. https://doi.org/10.1111/jop.13154
Advani, J., Verma, R., Chatterjee, O., Balaya, R. D. A., Najar, M. A., Ravishankara, N., Suresh, S., Pachori, P. K., Gupta, U. D., Pinto, S. M., Chauhan, D. S., Tripathy, S. P., Gowda, H., & Keshava Prasad, T. S. (2019). Rise of clinical microbial proteogenomics: A multiomics approach to nontuberculous mycobacterium-The case of Mycobacterium abscessus UC22. OMICS: A Journal of Integrative Biology, 23(1), 1–16. https://doi.org/10.1089/omi.2018.0116
Wisniewski, J. R., Zougman, A., Nagaraj, N., & Mann, M. (2009). Universal sample preparation method for proteome analysis. Nature Methods, 6(5), 359–362. https://doi.org/10.1038/nmeth.1322
Nirujogi, R.S., Muthusamy, B,. Kim, M.S., Sathe, G.J., P.T.V., Lakshmi, P.T.V., Kovbasnjuk, O.N., Keshava Prasad, T.S., Wade, M.,Jabbour, R.E. (2017). Secretome analysis of diarrhea-inducing strains of Escherichia coli. Proteomics, 17(6). https://doi.org/10.1002/pmic.201600299.
Kulak, N., Pichler, G., Paron, I., Nagaraj, N., & Mann, M. (2014). Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nature Methods, 11, 319–324. https://doi.org/10.1038/nmeth.2834
Dammalli, M., Murthy, K. R., Pinto, S. M., Murthy, K. B., Nirujogi, R. S., Madugundu, A. K., Dey, G., Nair, B., Gowda, H., & Keshava Prasad, T. S. (2017). Toward postgenomics ophthalmology: A proteomic map of the human choroid-retinal pigment epithelium tissue. OMICS: A Journal of Integrative Biology, 2, 114–122. https://doi.org/10.1089/omi.2016.0170
Palomba, A., Abbondio, M., Fiorito, G., Uzzau, S., Pagnozzi, D., & Tanca, A. (2021). Comparative evaluation of MaxQuant and proteome discoverer MS1-based protein quantification tools. Journal of Proteome Research, 20(7), 3497–3507. https://doi.org/10.1021/acs.jproteome.1c00143
Dammalli, M., Dey, G., Madugundu, A. K., Kumar, M., Rodrigues, B., Gowda, H., Siddaiah, B. G., Mahadevan, A., Shankar, S. K., & Prasad, T. S. K. (2017). Proteomic analysis of the human olfactory bulb. OMICS: A Journal of Integrative Biology, 21(8), 440–453. https://doi.org/10.1089/omi.2017.0084
Kumar, C., & Mann, M. (2009). Bioinformatics analysis of mass spectrometry-based proteomics data sets. FEBS Letters, 583(11), 1703–1712. https://doi.org/10.1016/j.febslet.2009.03.035
Ismail, G. A., Gheda, S. F., Abo-Shady, A. M., & Abdel-Karim, O. H. (2020). In vitro potential activity of some seaweeds as antioxidants and inhibitors of diabetic enzymes. Food Sci Technol, 40(3), 681–691.
El-Naggar, N. A. and El-Ewasy, S. (2017). Bioproduction, characterization, anticancer and antioxidant activities of extracellular melanin pigment produced by newly isolated microbial cell factories Streptomyces glaucescens NEAE-H. Scientific Report, 7(42129). https://doi.org/10.1038/srep42129.
Cheng, K. W., Lahad, J. P., Kuo, W. L., Lapuk, A., Yamada, K., Auersperg, N., Liu, J., Smith-McCune, K., Lu, K. H., Fishman, D., Gray, J. W., & Mills, G. B. (2004). The RAB25 small GTPase determines aggressiveness of ovarian and breast cancers. Nature Medicine, 10(11), 1251–1256. https://doi.org/10.1038/nm1125
Calderwood, S. K., & Gong, J. (2016). Heat shock proteins promote cancer: It’s a protection racket. Trends in Biochemical Sciences, 41(4), 311–323. https://doi.org/10.1016/j.tibs.2016.01.003
Wawrzynow, B., Zylicz, A., Zylicz, M. (2018). Chaperoning the guardian of the genome. The two-faced role of molecular chaperones in p53 tumor suppressor action. Biochimica et Biophysica Acta (BBA), 1869(2), 161–174. https://doi.org/10.1016/j.bbcan.2017.12.004.
Lagadec, C., Vlashi, E., Bhuta, S., Lai, C., Mischel, P., Werner, M., Henke, M., & Pajonk, F. (2014). Tumor cells with low proteasome subunit expression predict overall survival in head and neck cancer patients. BMC Cancer, 5(14), 152. https://doi.org/10.1186/1471-2407-14-152
Donna, L. D., Lagadec, C., & Pajonk, F. (2012). Radioresistance of prostate cancer cells with low proteasome activity. Prostate, 72(8), 868–874. https://doi.org/10.1002/pros.21489
Zou, Q., Jin, J., Hu, H., Li, H. S., Romano, S., Xiao, Y., Nakaya, M., Zhou, X., Cheng, X., Yang, P., Lozano, G., Zhu, C., Watowich, S. S., Ullrich, S. E., & Sun, S. C. (2014). USP15 stabilizes MDM2 to mediate cancer-cell survival and inhibit antitumor T cell responses. Nature Immunology, 15, 562–570. https://doi.org/10.1038/ni.2885
Kampen, K. R., Sulima, S. O., Vereecke, S., & De Keersmaecker, K. (2020). Hallmarks of ribosomopathies. Nucleic Acids Research, 48(3), 1013–1028.
Dai, M. S., Sears, R., & Lu, H. (2007). Feedback regulation of c-Myc by ribosomal protein L11. Cell Cycle, 6(22), 2735–2741. https://doi.org/10.4161/cc.6.22.4895
Liao, J. M., Zhou, X., Gatignol, A., & Lu, H. (2014). Ribosomal proteins L5 and L11 co-operatively inactivate c-Myc via RNA-induced silencing complex. Oncogene, 33(41), 4916–4923. https://doi.org/10.1038/onc.2013.430
Zhou, X., Hao, Q., Liao, J. M., Liao, P., & Lu, H. (2013). Ribosomal protein S14 negatively regulates c-Myc activity. Journal of Biological Chemistry, 288(30), 21793–21801. https://doi.org/10.1074/jbc.m112.445122
Desterro, J., Bak-Gordon, P., & Carmo-Fonseca, M. (2020). Targeting mRNA processing as an anticancer strategy. Nature Reviews. Drug Discovery, 19, 112–129. https://doi.org/10.1038/s41573-019-0042-3
Shrestha, N., Bahnan, W., Wiley, D. J., Barber, G., Fields, K. A., & Schesser, K. (2012). Eukaryotic initiation factor 2 (eIF2) signaling regulates proinflammatory cytokine expression and bacterial invasion. Journal of Biological Chemistry, 287(34), 28738–28744. https://doi.org/10.1074/jbc.M112.375915
Hinnebusch, A. G., Ivanov, I. P., & Sonenberg, N. (2016). Translational control by 5’-untranslated regions of eukaryotic mRNAs. Science, 352(6292), 1413–1416. https://doi.org/10.1126/science.aad9868
Quevedo, C., Salinas, M., & Alcázar, A. (2003). Initiation factor 2B activity is regulated by protein phosphatase 1, which is activated by the mitogen-activated protein kinase-dependent pathway in insulin-like growth factor 1-stimulated neuronal cells. Journal of Biological Chemistry, 278(19), 16579–16586. https://doi.org/10.1074/jbc.m212936200
Avila-Flores, A., Santos, T., Rincon, E., & Merida, I. (2005). Modulation of the mammalian target of rapamycin pathway by diacylglycerol kinase-produced phosphatidic acid. Journal of Biological Chemistry, 280(11), 10091–10099. https://doi.org/10.1074/jbc.m412296200
Liu, K., Xue, B., Bai, G., & Zhang, W. (2021). Downregulation of Diacylglycerol kinase zeta (DGKZ) suppresses tumorigenesis and progression of cervical cancer by facilitating cell apoptosis and cell cycle arrest. Bioengineered, 12(1), 1517–1529. https://doi.org/10.1080/21655979.2021.1918505
Roux, P. P., & Topisirovic, I. (2018). Signaling pathways involved in the regulation of mRNA translation. Molecular and Cellular Biology, 38(12), e00070-e118. https://doi.org/10.1128/MCB.00070-18
Moriyama, T., Yoneda, Y., Oka, M., Yamada, M. (2020). Transportin-2 plays a critical role in nucleocytoplasmic shuttling of oestrogen receptor-α. Scientific Report, 10(18640). https://doi.org/10.1038/s41598-020-75631-3.
Dubik, D., & Shiu, R. P. (1992). Mechanism of estrogen activation of c-myc oncogene expression. Oncogene, 7(8), 1587–1594.
Levin, E. R., & Hammes, S. R. (2016). Nuclear receptors outside the nucleus: Extranuclear signalling by steroid receptors. Nature Reviews Molecular Cell Biology, 17(12), 783–797. https://doi.org/10.1038/nrm.2016.122
Cavalcanti, F. N., Lucas, T. F., Lazari, M. F., & Porto, C. S. (2015). Estrogen receptor ESR1 mediates activation of ERK1/2, CREB, and ELK1 in the corpus of the epididymis. Journal of Molecular Endocrinology, 54(3), 339–349. https://doi.org/10.1530/jme-15-0086
Tecalco-Cruz, A. C., Perez-Alvarado, I. A., Ramirez-Jarquin, J. O., & Rocha-Zavaleta, L. (2017). Nucleo-cytoplasmic transport of estrogen receptor alpha in breast cancer cells. Cellular Signalling, 34, 121–132. https://doi.org/10.1016/j.cellsig.2017.03.011
Author information
Authors and Affiliations
Contributions
Study conception and design by Varalakshmi K Nadumane. Material preparation, data collection, and analysis were performed by Somasekhara D; Proteomics study and analysis by Manjunath Dammalli. The first draft of the manuscript was written by Somasekhara D; and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical Approval
Not applicable, as the study does not involve any human or animal samples.
Consent to Participate
Not applicable.
Consent to Publish
All authors agree mutually with the publication of this work and declare that this is an original research.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
D, S., Dammalli, M. & Nadumane, V.K. Proteomic Analysis of Human Breast Cancer MCF-7 Cells to Identify Cellular Targets of the Anticancer Pigment OR3 from Streptomyces coelicolor JUACT03. Appl Biochem Biotechnol 195, 236–252 (2023). https://doi.org/10.1007/s12010-022-04128-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-04128-8