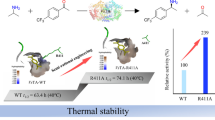
Abstract
Reliable kinetic parameters of enzymes are of paramount importance for a precise understanding of catalytic performance, which is essential for enzyme engineering and process optimization. Here, we developed a simple and convenient method to determine intrinsic kinetic parameters of R-selective ω-transaminases (ω-TAs) with a minimal set of kinetic data. Using (R)-α-methylbenzylamine ((R)-α-MBA) and pyruvate as a substrate pair, two R-selective ω-TAs from Arthrobacter sp. and Aspergillus fumigatus were subjected to kinetic measurements. In contrast to S-selective ω-TAs, both R-selective ω-TAs were observed to be devoid of substrate inhibition by pyruvate. Double reciprocal plot analysis was carried out with two sets of kinetic data obtained at varying concentrations of (R)-α-MBA under a fixed concentration of pyruvate and vice versa, leading to the determination of three intrinsic kinetic parameters, i.e., one kcat and two KM values, using three regression constants. The validity of the kinetic parameters was verified by a self-consistency test using a regression constant left out in the kinetic parameter determination, showing that deviations of calculated regression constants from the experimental ones were less than 15%. Because the kinetic parameters for (R)-α-MBA and pyruvate are not apparent but intrinsic, a cosubstrate substitution method enabled rapid determination of intrinsic parameters for a new substrate pair using just one set of kinetic data. Eventually, computational modeling of kinetic resolution of rac-α-MBA was carried out and showed a good agreement with experimental reaction progresses.





Similar content being viewed by others
References
Ghislieri, D., & Turner, N. J. (2014). Biocatalytic approaches to the synthesis of enantiomerically pure chiral amines. Topics in Catalysis, 57, 284–300.
Kelly, S. A., Pohle, S., Wharry, S., Mix, S., Allen, C. C. R., Moody, T. S., & Gilmore, B. F. (2018). Application of ω-transaminases in the pharmaceutical industry. Chemical Reviews, 118, 349–367.
Patil, M. D., Grogan, G., Bommarius, A., & Yun, H. (2018). Oxidoreductase-catalyzed synthesis of chiral amines. ACS Catalysis, 8, 10985–11015.
Grogan, G. (2018). Synthesis of chiral amines using redox biocatalysis. Current Opinion in Chemical Biology, 43, 15–22.
Malik, M. S., Park, E. S., & Shin, J. S. (2012). Features and technical applications of ω-transaminases. Applied Microbiology and Biotechnology, 94(5), 1163–1171.
Guo, F., & Berglund, P. (2017). Transaminase biocatalysis: optimization and application. Green Chemistry, 19, 333–360.
Savile, C. K., Janey, J. M., Mundorff, E. C., Moore, J. C., Tam, S., Jarvis, W. R., Colbeck, J. C., Krebber, A., Fleitz, F. J., Brands, J., Devine, P. N., Huisman, G. W., & Hughes, G. J. (2010). Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science, 329(5989), 305–309.
Simon, R. C., Richter, N., Busto, E., & Kroutil, W. (2013). Recent developments of cascade reactions involving ω-transaminases. ACS Catalysis, 4, 129–143.
Höhne, M., Schätzle, S., Jochens, H., Robins, K., & Bornscheuer, U. T. (2010). Rational assignment of key motifs for function guides in silico enzyme identification. Nature Chemical Biology, 6(11), 807–813.
Esparza-Isunza, T., González-Brambila, M., Gani, R., Woodley, J. M., & López-Isunza, F. (2015). The coupling of ω-transaminase and Oppenauer oxidation reactions via intra-membrane multicomponent diffusion - a process model for the synthesis of chiral amines. Chemical Engineering Journal, 259, 221–231.
Milker, S., Fink, M. J., Oberleitner, N., Ressmann, A. K., Bornscheuer, U. T., Mihovilovic, M. D., & Rudroff, F. (2017). Kinetic modeling of an enzymatic redox cascade in vivo reveals bottlenecks caused by cofactors. ChemCatChem, 9, 3420–3427.
Shin, J. S., & Kim, B. G. (1998). Kinetic modeling of ω-transamination for enzymatic kinetic resolution of α-methylbenzylamine. Biotechnology and Bioengineering, 60(5), 534–540.
Shin, J. S., & Kim, B. G. (2002). Substrate inhibition mode of ω-transaminase from Vibrio fluvialis JS17 is dependent on the chirality of substrate. Biotechnology and Bioengineering, 77(7), 832–837.
Park, E. S., & Shin, J. S. (2013). ω-Transaminase from Ochrobactrum anthropi is devoid of substrate and product inhibitions. Applied and Environmental Microbiology, 79(13), 4141–4144.
Leipold, L., Dobrijevic, D., Jeffries, J. W. E., Bawn, M., Moody, T. S., Ward, J. M., & Hailes, H. C. (2019). The identification and use of robust transaminases from a domestic drain metagenome. Green Chemistry, 21(1), 75–86.
Heuson, E., Charmantray, F., Petit, J. L., de Berardinis, V., & Gefflaut, T. (2019). Enantioselective synthesis of D- and L-α-amino acids by enzymatic transamination using glutamine as smart amine donor. Advanced Synthesis & Catalysis, 361, 778–785.
Han, S. W., & Shin, J. S. (2016). A facile method to determine intrinsic kinetic parameters of ω-transaminase displaying substrate inhibition. Journal of Molecular Catalysis B: Enzymatic, 133, S500–S507.
Shin, J. S., & Kim, B. G. (2001). Comparison of the ω-transaminases from different microorganisms and application to production of chiral amines. Bioscience, Biotechnology, and Biochemistry, 65, 1782–1788.
Park, E. S., Kim, M., & Shin, J. S. (2012). Molecular determinants for substrate selectivity of ω-transaminases. Applied Microbiology and Biotechnology, 93(6), 2425–2435.
Park, E. S., Malik, M. S., Dong, J. Y., & Shin, J. S. (2013). One-pot production of enantiopure alkylamines and arylalkylamines of opposite chirality catalyzed by ω-transaminase. ChemCatChem, 5, 1734–1738.
Park, E. S., Dong, J. Y., & Shin, J. S. (2014). Active site model of (R)-selective ω-transaminase and its application to the production of D-amino acids. Applied Microbiology and Biotechnology, 98, 651–660.
Han, S. W., Kim, J., Cho, H. S., & Shin, J. S. (2017). Active site engineering of ω-transaminase guided by docking orientation analysis and virtual activity screening. ACS Catalysis, 7, 3752–3762.
Schätzle, S., Höhne, M., Redestad, E., Robins, K., & Bornscheuer, U. T. (2009). Rapid and sensitive kinetic assay for characterization of ω-transaminases. Analytical Chemistry, 81(19), 8244–8248.
Bhushan, R., & Brückner, H. (2004). Marfey’s reagent for chiral amino acid analysis: a review. Amino Acids, 27(3-4), 231–247.
Han, S. W., Park, E. S., Dong, J. Y., & Shin, J. S. (2015). Mechanism-guided engineering of ω-transaminase to accelerate reductive amination of ketones. Advanced Synthesis & Catalysis, 357, 1732–1740.
Shin, G., Mathew, S., & Yun, H. (2015). Kinetic resolution of amines by (R)-selective omega-transaminase from Mycobacterium vanbaalenii. Journal of Industrial and Engineering Chemistry, 23, 128–133.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (MSIT) (2019R1F1A1062845). Dr. S.-W. Han was financially supported by the Initiative for Biological Function & Systems under the BK21 PLUS program of the Korean Ministry of Education.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no competing interests.
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Han, SW., Shin, JS. Kinetic Analysis of R-Selective ω-Transaminases for Determination of Intrinsic Kinetic Parameters and Computational Modeling of Kinetic Resolution of Chiral Amine. Appl Biochem Biotechnol 191, 92–103 (2020). https://doi.org/10.1007/s12010-020-03240-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-020-03240-x