Abstract
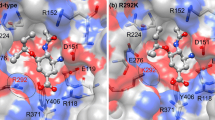
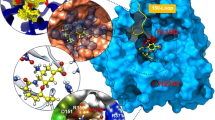
Oseltamivir (Tamiflu) is the preferred anti-viral drug employed to fight the flu virus in infected individuals. The principal target for this drug is a virus surface glycoprotein, neuraminidase (NA), which facilitates the release of nascent virus and thus spreads infections. Until recently, only a low prevalence of neuraminidase inhibitors (NAIs) resistance (<1 %) had been detected in circulating viruses. However, there have been reports of significant numbers of A (H1N1) influenza strains with a H274Y neuraminidase mutation that was highly resistant to the NAI, oseltamivir. In this study, we highlight the effect of point mutation-induced oseltamivir resistance in H1N1 subtype neuraminidases by molecular docking and molecular dynamics simulation approach. Our results suggested that wild-type NA could be more indispensable for the oseltamivir binding, as characterized by minimum number of H-bonds, high flexibility and largest binding affinity than mutant-type NA. This study throws light on the possible effects of drug-resistant mutations on the large functionally important collective motions in biological systems.













Similar content being viewed by others
References
Mukhtar, M. M., Rasool, S. T., Song, D., Zhu, C., Hao, Q., Zhu, Y., & Wu, J. (2007). Journal of General Virology, 88, 3094–3099.
Shirvan, A. N., Moradi, M., Aminian, M., & Madani, R. J. (2007). Vetenary and Animal Science, 31, 219–223.
Takeda, M., Leser, G. P., Russell, C. J., & Lamb, R. A. (2003). Proceedings of the National Academy of Sciences, 100, 14610–14617.
McKimm-Breschkin, J. L. (2000). Antiviral Research, 47, 1–17.
Du, Q. S., Wang, S. Q., & Chou, K. C. (2007). Biophysica Research Communication, 354, 634–640.
Von Itzstein, M., Wu, W. Y., Kok, G. B., Pegg, M. S., Dyason, J. C., Jin, B., Van Phan, T., Smythe, M. L., White, H. F., & Oliver, S. W. (1993). Nature, 363, 418–423.
Kim, C. U., Lew, W., Williams, M. A., Liu, H., Zhang, L., Swaminathan, S., Bischofberger, N., Chen, M. S., Mendel, D. B., & Tai, C. Y. (1997). Journal of the American Chemical Society, 119, 681–690.
Gubareva, L. V., Kaiser, L., & Hayden, F. G. (2000). Lancet, 355, 827–835.
Bucher, D. J., & Kilbourne, E. D. (1972). Journal of Virology, 10, 60–66.
Baum, S. G. (2009). Clinical Infectious Diseases, 49, 1836–1837.
Centers for Disease Control & Prevention (CDC). (2008). MMWR Morb Mortal Wkly Rep, 57, 1329–1332.
Gubareva, L. V. (2004). Virus Research, 103, 199–203.
Lander, G. C., Evilevitch, A., Jeembaeva, M., Potter, C. S., Carragher, B., & Johnson, J. E. (2008). Structure, 16, 1399–1406.
Berman, H. M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T. N., Weissig, H., Shindyalov, I. N., & Bourne, P. E. (2000). Nucleic Acids Research, 28, 235–242.
Russell, R. J., Haire, L. F., Stevens, D. J., Collins, P. J., Lin, Y. P., Blackburn, G. M., Hay, A. J., Gamblin, S. J., & Skehel, J. J. (2006). Nature, 443, 45–49.
Collins, P. J., Haire, L. F., Lin, Y. P., Liu, J., Russell, R. J., Walker, P. A., Skehel, J. J., Martin, S. R., Hay, A. J., & Gamblin, S. J. (2008). Nature, 453, 1258–1261.
Feldman, J., Snyder, K. A., Ticoll, A., Pintilie, G., & Hogue, C. W. (2006). FEBS Letters, 580, 1649–1653.
Gasteiger, J., Rudolph, C., & Sadowski, J. (1990). Tetrahedron. Compututer, Method, 3, 537–547.
Lopez, G., Valencia, A., & Tress, M. L. (2007). Nucleic Acids Research, 35, 573–577.
Yuan, Z., Bailey, T. L., & Teasdale, R. D. (2005). Proteins, 58, 905–912.
Ringe, D., & Petsko, G. A. (1986). Methods in Enzymology, 131, 389–433.
Parthasarathy, S., & Murthy, M. (2000). r. Protein Engineering, 13, 9–13.
Carlson, H. A., & McCammon, J. A. (2000). Molecular Pharmacology, 57, 213–218.
Hinkle, A., & Tobacman, L. S. (2003). Journal of Biological Chemistry, 278, 506–513.
Rajasekaran, R., George Priya Doss, C., Sudandiradoss, C., Ramanathan, K., Purohit, R., & Sethumadhavan, R. (2008). Comptes Rendus Biologies, 331, 409–417.
Suhre, K., & Sanejouand, Y. H. (2004). Nucleic Acids Research, 32, 610–614.
Thompson M. A. (2004). ArgusLab 4.0.1. Planaria Software, LLC, Seattle, WA, http://www.ArgusLab.com.
Thompson, M. A. (2004). Poster presentation: molecular docking using arguslab: an efficient shape-based search algorithm & the AScore scoring function. Philadelphia: Fall ACS meeting.
Hess, B., Kutzner, C., Spoel, D., & Lindahl, E. (2008). J Chem Theory Comput, 4, 435–447.
Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E., & Berendsen, H. J. (2005). J Chem Theory Comput, 26, 1701–1718.
Meagher, K. L., & Carlson, H. A. (2005). Proteins: Structural Functions Bioinformatics, 58, 119–125.
Darden, T., Perera, L., Li, L., & Pedersen, L. (1999). Structure, 7, 55–60.
Van Gunsteren, W. F., & Berendsen, H. J. (1977). Molecular Physics, 34, 1311–1327.
Wallace, A. C., Laskowski, R. A., & Thornton, J. M. (1995). Protein Engineering, 8, 127–134.
Colman, P. M., Varghese, J. N., & Laver, W. G. (1983). Nature, 303, 41–44.
Rungrotmongkol, T., Udommaneethanakit, T., Malaisree, M., Nunthaboot, N., Intharathep, P., Sompornpisut, P., & Hannongbua, S. (2009). Biophysical Chemistry, 145, 29–36.
Cavasotto, C. N., Kovacs, J. A., & Abagyan, R. A. (2005). Journal of the American Chemical Society, 26, 9632–9640.
Teague, S. J. (2003). Nature Reviews. Drug Discovery, 2, 527–541.
Durrant, J. D., & McCammon, J. A. (2011). BMC Biology, 9, 71–79.
Acknowledgments
The authors express deep sense of gratitude to the management of Vellore Institute of Technology for all the support, assistance, and constant encouragements to carry out this work. The authors also thank Professor M.A. Mohamed Sahul Hameed, English division, for English editing and grammar corrections in our manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Karthick, V., Shanthi, V., Rajasekaran, R. et al. Exploring the Cause of Oseltamivir Resistance Against Mutant H274Y Neuraminidase by Molecular Simulation Approach. Appl Biochem Biotechnol 167, 237–249 (2012). https://doi.org/10.1007/s12010-012-9687-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-012-9687-7