Abstract
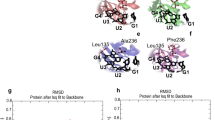
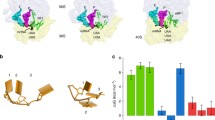
The C domain of eRF1 interacts with the C domain of eRF3, and the binding of both factors is essential for fast kinetics of the termination of protein translation. Analysis by computational simulation demonstrated that several peptides involved in Eo-eRF1/Eo-eRF3 interaction directly. Among these peptides, the two motifs GVEDT and GFGG were highly conserved, while the fragment aa338-346 of Eo-eRF1a/b was variable. In additional, I290 and D293 of Eo-eRF1 were also highly conserved. By the site-directed mutagenesis and pull-down analysis, the amino acid D293 in Eo-eRF1bC domain was conformed playing an important role in eRF1–eRF3 interaction. Eo-eRF1a and Eo-eRF1b may select different manners to interact with Eo-eRF3. These studies contribute to the better understanding the mode of eRF1–eRF3 interaction.




Similar content being viewed by others
Abbreviations
- eRF1:
-
Class I polypeptide release factor in eukaryotes
- eRF3:
-
Class II polypeptide release factor in eukaryotes
- Bj-eRF1:
-
Class I polypeptide release factor in Blepharisma japonicum
- Eo-eRF1a:
-
Class I polypeptide release factor a in Euplotes octocarinatus
- Eo-eRF1b:
-
Class I polypeptide release factor b in E. octocarinatus
- Hs-eRF1:
-
Class I polypeptide release factor in Homo sapiens
- Bj-eRF1C:
-
C domain of Bj-eRF1 (274–436)
- Eo-eRF1aC:
-
C domain of Eo-eRF1a (270–446)
- Eo-eRF1bC:
-
C domain of Eo-eRF1b (270–437)
- Eo-eRF3:
-
Class II polypeptide release factor in E. octocarinatus
- Eo-eRF3Cm6:
-
Truncated peptide of Eo-eRF3 (640–723)
- Bj-eRF1C/Eo-eRF3Cm6:
-
Heterodimeric complex of Bj-eRF1C and Eo-eRF3Cm6
- Eo-eRF1aC/Eo-eRF3Cm6:
-
Heterodimeric complex of Eo-eRF1aC and Eo-eRF3Cm6
- Eo-eRF1bC/Eo-eRF3Cm6:
-
Heterodimeric complex of Eo-eRF1bC and Eo-eRF3Cm6
- MD:
-
Molecular dynamics
- RLS:
-
Resonance light scattering
References
Frolova, L., Le Goff, X., Rasmussen, H. H., Cheperegin, S., Drugeon, G., Kress, M., et al. (1994). Nature, 372, 701–703.
Song, H., Mugnier, P., Das, A. K., Webb, H. M., Evans, D. R., Tuite, M. F., et al. (2000). Cell, 100, 311–321.
Kononenko, A. V., Dembo, K. A., Kiselev, L. L., & Volkov, V. V. (2004). Molecular Biology (Mosk), 38, 303–311.
Merkulova, T. I., Frolova, L. Y., Lazar, M., Camonis, J., & Kisselev, L. L. (1999). FEBS Letters, 44, 341–347.
Fan-Minogue, Hua, Du, Ming, Pisarev, A. V., Kallmeyer, A. K., Salas-Marco, J., Keeling, K. M., et al. (2008). Molecular Cell, 30, 599–609.
Salas-Marco, J., Bedwell, D. M. (2004). Molecular Cellular Biology. Sept. 7769–7778.
Alkalaeva, E. Z., Pisarev, A. V., Frolova, L. Y., Kisselev, L. L., & Pestova, T. V. (2006). Cell, 125, 1125–1136.
Lozupone, C. A., Knight, R. D., & Landweber, L. F. (2001). Current Biology, 11, 65–74.
Liang, A., Brünen-Nieweler, C., Muramatsu, T., Kuchino, Y., Beier, H., & Heckmann, K. (2001). Gene, 262, 161–168.
Kim, O., Yura, K., Go, N., & Harumoto, T. (2005). Gene, 346, 277–286.
Vorob'ev, I., Kiselev, L. L. (2008). Molecular Biology (Mosk), 42341–351.
Wang, Y., Chai, B. F., Wang, W., & Liang, A. H. (2010). Bioscience Reports, 30, 425–431.
Song, L., Chai, B. F., Wang, W., & Liang, A. H. (2006). Research in Microbiology, 157, 842–850.
Mantsyzov, A. B., Ivanova, E. V., Birdsall, B., Alkalaeva, E. Z., Kryuchkova, P. N., Kelly, G., et al. (2010). FEBS Journal, 277, 2611–2627.
Cheng, Z. H., Saito, K., & Pisarev, A. V. (2009). Genes & Development, 23, 1106–1118.
Arnold, K., Bordol, L. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics. 22195–201.
Schwede, T., & Kopp, J. (2003). Nucleic Acids Research, 31, 3381–3385.
Guex, N., & Peitsch, M. C. (1997). Electrophoresis, 18, 2714–2723.
Sambrook, J., Fritsch, E. F., & Maniatis, T. (2005). Proteins, 60, 207–221.
Wang, W., Chai, B. F., Heckmann, K., & Liang, A. H. (2004). Biotechnology Letters, 26, 959–963.
Acknowledgments
This work was supported by grants from the National Natural Scientific Foundation of China (Nos. 31071294, 30770294, and 30770295).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Table S1
The parameters of hydrogen bond in Eo-eRF1bC/Eo-eRF3Cm6 complex (DOCX 12 kb)
Table S2
The parameters of hydrogen bond in Eo-eRF1aC/Eo-eRF3Cm6 complex (DOCX 12 kb)
Table S3
The parameters of hydrogen bond in Bj-eRF1C/Eo-eRF3Cm6 complex (DOCX 11 kb)
Figure S1
The conformers of minidomain and the site D297 in eRF1C/Eo-eRF3Cm6 complexes (DOC 1977 kb)
Rights and permissions
About this article
Cite this article
Chen, J., Fu, Yj., Yang, Bs. et al. The Binding Sites of Class I Release Factor (eRF1) Toward Class II Release Factor (eRF3) in Euplotes octocarinatus . Appl Biochem Biotechnol 165, 1507–1518 (2011). https://doi.org/10.1007/s12010-011-9371-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-011-9371-3