Abstract
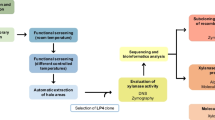
The genome of Dictyoglomus turgidum was sequenced and analyzed for carbohydrases. The broad range of carbohydrate substrate utilization is reflected in the high number of glycosyl hydrolases, 54, and the high percentage of CAZymes present in the genome, 3.09% of its total genes. Screening a random clone library generated from D. turgidum resulted in the discovery of five novel biomass-degrading enzymes with low homology to known molecules. Whole genome sequencing of the organism followed by bioinformatics-directed amplification of selected genes resulted in the recovery of seven additional novel enzyme molecules. Based on the analysis of the genome, D. turgidum does not appear to degrade cellulose using either conventional soluble enzymes or a cellulosomal degradation system. The types and quantities of glycosyl hydrolases and carbohydrate-binding modules present in the genome suggest that D. turgidum degrades cellulose via a mechanism similar to that used by Cytophaga hutchinsonii and Fibrobacter succinogenes.



Similar content being viewed by others
References
Svetlichnii, V. A., & Svetlichnii, T. P. (1988). Dictyoglomus turgidus sp. nov., a new extremely thermophilic eubacterium isolated from hot springs of the Uzon volcano caldera. Mikrobiologiya, 57, 435–441.
Saiki, A., Kobayashi, Y., Kawagoe, K., & Beppu, T. (1985). Dictyoglomus thermophilum gen. nov., sp. nov., a chemoorganotrophic, anaerobic, thermophilic bacterium. International Journal of Systematic Bacteriology, 35, 253–259.
Morris, D. D., Gibbs, M. D., Chin, C. W. J., Koh, M. H., Wong, K. K. Y., Allison, R. W., et al. (1998). Cloning of the xynB Gene from Dictyoglomus thermophilum Rt46B.1 and action of the gene product on kraft pulp. Applied and Environmental Microbiology, 64, 1759–1765.
Sambrook, J., Fritsch, E. F., & Maniatis, T. (1989). Molecular cloning: A laboratory manual, vol. 1, 2, 3. New York: Cold Spring Harbor Laboratory Press.
Brumm, P. J., Hochstein, B., Boyum, J., Magallanes, N., Desai, D., Hermersmann, N., et al. (2009). Mining Clostridium thermocellum for enzymatically active carbohydrases. Presented at The Thirty-First Symposium Biotechnology for Fuels and Chemicals.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410.
Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389–3402.
Thompson, J. D., Higgins, D. G., & Gibson, T. J. (1994). CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22, 4673–80.
Guindon, S., & Gascuel, O. (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Systems Biology, 52, 696–704.
Dereeper, A., Guignon, V., Blanc, G., Audic, S., Buffet, S., Chevenet, F., et al. (2008). Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res, 36, W465–9 (Web Server issue).
Qi, M., Jun, H., & ClW, F. (2007). Characterization and synergistic interactions of Fibrobacter succinogenes glycoside hydrolases. Applied and Environmental Microbiology, 73, 6098–6105.
Acknowledgements
This work was funded in part by DOE grant DOE DE-FG36-06GO16106, “Novel enzyme products for the conversion of defatted soybean meal to ethanol,” and funded in part by the DOE Great Lakes Bioenergy Research Center (DOE Office of Science BER DE-FC02-07ER64494). The work conducted by the U.S. Department of Energy Joint Genome Institute is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brumm, P., Hermanson, S., Hochstein, B. et al. Mining Dictyoglomus turgidum for Enzymatically Active Carbohydrases. Appl Biochem Biotechnol 163, 205–214 (2011). https://doi.org/10.1007/s12010-010-9029-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-010-9029-6