Abstract
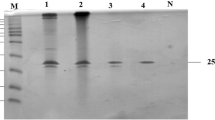
To obtain extracellular and high-level expression of the Dictyoglomus thermophilum Rt46B.1 xylanase B gene, this gene was integrated into the α-amylase gene site of a host strain of Bacillus subtilis WB800. The extreme thermophile xylanase gene was successfully integrated and expressed in the host, measured at 24 ± 0.4 XUs/mL in the Luria broth medium supernatant. The recombinant enzyme was purified by ammonium sulfate precipitation, anion exchange chromatography, and gel filtration. The molecular mass and pI value of xylanase were estimated to be 24 kDa and 4.3, respectively. The optimal pH level and temperature of the purified enzyme were 6.5 and 85 °C, respectively. Xylanase showed reasonable activity at temperatures up to 95 °C and remained stable at 4 °C for 1 week. The purified enzyme retained most of its activity in 1 mM ethylenediaminetetraacetic acid or dithiothreitol and 0.1% Tween-20 or Triton X-100. However, strong inhibition was observed in the presence of 5 mM Mn2+, 0.5% sodium dodecyl sulfate, Tween-20, or Triton X-100; a strong stimulating effect was also observed in the presence of Fe2+. The K m and V max values of the recombinant xylanase for birchwood xylan were calculated to be 2.417 ± 0.36 mg/mL and 325 ± 41 µmol/min mg, respectively. Xylanase was found to be useful in the prebleaching process of paper pulps.




Similar content being viewed by others
References
Biely, P. (1985). Trends in Biotechnology, 3, 286–290. doi:10.1016/0167-7799(85)90004-6.
Collins, T., Gerday, C., & Feller, G. (2005). FEMS Microbiology Reviews, 29, 3–23. doi:10.1016/j.femsre.2004.06.005.
Beg, Q. K., Kapoor, M., Mahajan, L., & Hoondal, G. S. (2001). Applied Microbiology and Biotechnology, 56, 326–338. doi:10.1007/s002530100704.
Subramaniyan, S., & Prema, P. (2002). Critical Reviews in Biotechnology, 22, 33–64. doi:10.1080/07388550290789450.
Yin, Y. L., Baidoo, S. K., Jin, L. Z., Liu, Y. G., Schulze, H., & Simmins, P. H. (2001). Livestock Production Science, 71, 109–120. doi:10.1016/S0301-6226(01)00215-9.
Kantelinen, A., Rantanen, T., Buchert, J., & Vikari, L. (1993). Journal of Biotechnology, 28, 219–228. doi:10.1016/0168-1656(93)90171-I.
Viikari, L., Kantelinen, A., Sundquist, J., & Linko, M. (1994). FEMS Microbiology Reviews, 13, 335–350. doi:10.1111/j.1574-6976.1994.tb00053.x.
Prade, R. A. (1996). Biotechnology & Genetic Engineering Reviews, 13, 101–131.
Georis, J., Giannotta, F., Buyl, E. D., Granier, B., & Frere, J. M. (2000). Enzyme and Microbial Technology, 26, 178–186. doi:10.1016/S0141-0229(99)00141-6.
Haarhoff, J., Moes, C. J., Cerff, C., Wyk, W. J. V., Gerischer, G., & Janse, B. J. H. (1999). Biotechnology Letters, 21, 415–420.
Ximenes, F. A., Sousa, M. V., Puls, J., Silva, F. G., & Filho, E. X. F. (1999). Current Microbiology, 38, 18–21. doi:10.1007/PL00006765.
Kulkarni, N., Shendye, A., & Rao, M. (1999). FEMS Microbiology Reviews, 23, 411–456. doi:10.1111/j.1574-6976.1999.tb00407.x.
Maheshwari, R., Bharadwaj, G., & Bhat, M. K. (2000). Microbiology and Molecular Biology Reviews, 64, 461–488. doi:10.1128/MMBR.64.3.461-488.2000.
Singh, S., Madlala, A. M., & Prior, B. A. (2003). FEMS Microbiology Reviews, 27, 3–16. doi:10.1016/S0168-6445(03)00018-4.
Dhillon, A., Gupta, J. K., Jauhari, B. M., & Khanna, S. (2000). Bioresource Technology, 73, 273–277. doi:10.1016/S0960-8524(99)00116-9.
Lama, L., Calandrelli, V., Gambacorta, A., & Nicolaus, B. (2004). Research in Microbiology, 155, 283–289. doi:10.1016/j.resmic.2004.02.001.
Reeves, R. A., Gibbs, M. D., Morris, D. D., Griffiths, K. R., Saul, D. J., & Bergquist, P. L. (2000). Applied and Environmental Microbiology, 66, 1532–1537. doi:10.1128/AEM.66.4.1532-1537.2000.
Winterhalter, C., & Liebl, W. (1995). Applied and Environmental Microbiology, 61(5), 1810–1815.
Sambrook, J., Fritsch, E. F., & Maniatis, T. (1989). Molecular cloning: a laboratory manual (2nd ed.). Cold Spring Harbor, NY: Cold Spring Harbor Laboratory.
Anagnostopoulos, C., & Spizizen, J. (1961). Journal of Bacteriology, 81, 741–746.
Shimotsu, H., & Henner, D. J. (1986). Gene, 43, 85–94. doi:10.1016/0378-1119(86)90011-9.
Lever, M. (1973). Biochemical Medicine, 7, 274–281. doi:10.1016/0006-2944(73)90083-5.
Baily, M. J., Biely, P., & Poutanen, K. (1992). Journal of Biotechnology, 23, 257–270. doi:10.1016/0168-1656(92)90074-J.
Shah, A. R., & Madamwar, D. (2005). Process Biochemistry, 40, 1763–1771. doi:10.1016/j.procbio.2004.06.041.
Laemmli, U. K. (1970). Nature, 227, 680–685. doi:10.1038/227680a0.
Smith, P. K., Krohn, R. I., Hermanson, G. T., Mallia, A. K., Gartner, F. H., Provenzano, M. D., et al. (1985). Analytical Biochemistry, 150, 76–85. doi:10.1016/0003-2697(85)90442-7.
Lineweaver, H., & Burk, D. (1934). Journal of the American Chemical Society, 57, 685–66.
Morris, D. D., Gibbs, M. D., Chin, C. W., Koh, M. H., Wong, K. Y., Allison, R. W., et al. (1998). Applied and Environmental Microbiology, 64, 1759–1765.
Charles, C. L., Rena, E. K., Michael, R. S., Kurt, W., William, J. O., & Dominic, W. S. W. (2008). Current Microbiology, 57, 301–305. doi:10.1007/s00284-008-9193-x.
Gessesse, A. (1998). Applied and Environmental Microbiology, 64, 3533–3535.
Johnvesly, B., Virupakshi, S., Patil, G.N., Ramalingam, & Naik, G.R. (2002). J of Microbiology and Biotechnology, 12, 153–156.
Acknowledgements
This work was financially supported by the High Technology Foundation (200515124) of Xinjiang Province, People's Republic of China. We thank Dr Satoshi Nakamula and Dr. Xing Xin-hui for providing the material of integration vector pDG364 and the plasmid of pET-xynB.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, W., Lou, K. & Li, G. Expression and Characterization of the Dictyoglomus thermophilum Rt46B.1 Xylanase Gene (xynB) in Bacillus subtilis . Appl Biochem Biotechnol 160, 1484–1495 (2010). https://doi.org/10.1007/s12010-009-8634-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-009-8634-8