Abstract
Purpose of the Review
Kidney disease is the major cause of morbidity and mortality in patients with diabetes. Poor glycemic control shows the strongest correlation with diabetic kidney disease (DKD) development. A period of poor glycemia increases kidney disease risk even after an extended period of improved glucose control—a phenomenon called metabolic memory. Changes in the epigenome have been proposed to mediate the metabolic memory effect, as epigenome editing enzymes are regulated by substrates of intermediate metabolism and changes in the epigenome can be maintained after cell division.
Recent Findings
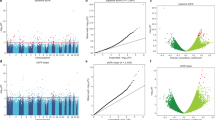
Epigenome-wide association studies (EWAS) have reported differentially methylated cytosines in blood and kidney samples of DKD subjects when compared with controls. Differentially methylated cytosines were enriched on regulatory regions and some correlated with gene expression. Methylation changes predicted the speed of kidney function decline. Site-specific methylome editing tools now can be used to interrogate the functional role of differentially methylated regions.
Summary
Methylome changes can be detected in blood and kidneys of patients with DKD. Methylation changes can predict future kidney function changes. Future studies shall determine their role in disease development.




Similar content being viewed by others
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Gluck C, Ko YA, Susztak K. Precision medicine approaches to diabetic kidney disease: tissue as an issue. Curr Diabetes Rep. 2017;17:30.
Reidy K, Kang HM, Hostetter T, Susztak K. Molecular mechanisms of diabetic kidney disease. J Clin Invest. 2014;124:2333–40.
Breyer MD, Susztak K. The next generation of therapeutics for chronic kidney disease. Nat Rev Drug Discov. 2016;15:568–88.
Pattaro C, et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nat Commun. 2016;7:10023.
Qiu C, Huang S, Park J, Park YS, Ko YA, Seasock MJ, et al. Renal compartment-specific genetic variation analyses identify new pathways in chronic kidney disease. Nat Med. 2018;24:1721–31.
D. E. R. Group, et al. Intensive diabetes therapy and glomerular filtration rate in type 1 diabetes. N Engl J Med. 2011;365:2366–76.
Barker DJ, et al. Type 2 (non-insulin-dependent) diabetes mellitus, hypertension and hyperlipidaemia (syndrome X): relation to reduced fetal growth. Diabetologia. 1993;36:62–7.
Jimenez-Chillaron JC, Isganaitis E, Charalambous M, Gesta S, Pentinat-Pelegrin T, Faucette RR, et al. Intergenerational transmission of glucose intolerance and obesity by in utero undernutrition in mice. Diabetes. 2009;58:460–8.
Oge A, Isganaitis E, Jimenez-Chillaron J, Reamer C, Faucette R, Barry K, et al. In utero undernutrition reduces diabetes incidence in non-obese diabetic mice. Diabetologia. 2007;50:1099–108.
Radford EJ, et al. In utero effects. In utero undernourishment perturbs the adult sperm methylome and intergenerational metabolism. Science. 2014;345:1255903.
Seki Y, Suzuki M, Guo X, Glenn AS, Vuguin PM, Fiallo A, et al. In utero exposure to a high-fat diet programs hepatic hypermethylation and gene dysregulation and development of metabolic syndrome in male mice. Endocrinology. 2017;158:2860–72.
• Chen Z, et al. Epigenomic profiling reveals an association between persistence of DNA methylation and metabolic memory in the DCCT/EDIC type 1 diabetes cohort. Proc Natl Acad Sci U S A. 2016;113:E3002–11. This study analyzed methylome changes in blood samples from the well-characterized DCCT cohort and identified methylation changes correlating with metabolic memory.
Cooper ME, El-Osta A. Epigenetics: mechanisms and implications for diabetic complications. Circ Res. 2010;107:1403–13.
Cooper ME, el-Osta A, Allen TJ, Watson AMD, Thomas MC, Jandeleit-Dahm KAM. Metabolic karma-the atherogenic legacy of diabetes: the 2017 Edwin Bierman Award Lecture. Diabetes. 2018;67:785–90.
Ko YA, Susztak K. Epigenomics: the science of no-longer-junk DNA. Why study it in chronic kidney disease? Semin Nephrol. 2013;33:354–62.
Pagliaroli L, Vető B, Arányi T, Barta C. From genetics to epigenetics: new perspectives in Tourette syndrome research. Front Neurosci. 2016;10:277.
Kato M, Natarajan R. Epigenetics and epigenomics in diabetic kidney disease and metabolic memory. Nat Rev Nephrol. 2019;15:327–45.
Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes Dev. 2011;25:1010–22.
Arányi T, et al. The tissue-specific methylation of the human tyrosine hydroxylase gene reveals new regulatory elements in the first exon. J Neurochem. 2005;94:129–39.
Mohtat D, Susztak K. Fine tuning gene expression: the epigenome. Semin Nephrol. 2010;30:468–76.
Li SY, Park J, Guan Y, Chung K, Shrestha R, Palmer MB, et al. DNMT1 in Six2 progenitor cells is essential for transposable element silencing and kidney development. J Am Soc Nephrol. 2019;30:594–609.
Rosen ED, Kaestner KH, Natarajan R, Patti ME, Sallari R, Sander M, et al. Epigenetics and epigenomics: implications for diabetes and obesity. Diabetes. 2018;67:1923–31.
Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–80.
Buenrostro JD, Wu B, Chang HY, Greenleaf WJ. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr Protocols Mol Biol. 2015;109:21 29 21–9.
Susztak K. Understanding the epigenetic syntax for the genetic alphabet in the kidney. J Am Soc Nephrol. 2014;25:10–7.
Brind'Amour J, et al. An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat Commun. 2015;6:6033.
Hyun BR, McElwee JL, Soloway PD. Single molecule and single cell epigenomics. Methods. 2015;72:41–50.
Tang F, Barbacioru C, Nordman E, Li B, Xu N, Bashkirov VI, et al. RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nat Protoc. 2010;5:516–35.
Clark SJ, Harrison J, Paul CL, Frommer M. High sensitivity mapping of methylated cytosines. Nucleic Acids Res. 1994;22:2990–7.
Feng L, Lou J. DNA methylation analysis. Methods Mol Biol. 2019;1894:181–227.
Schutsky EK, DeNizio JE, Hu P, Liu MY, Nabel CS, Fabyanic EB, et al. Nondestructive, base-resolution sequencing of 5-hydroxymethylcytosine using a DNA deaminase. Nat Biotechnol. 2018;36:1083–90.
Vető B, Szabó P, Bacquet C, Apró A, Hathy E, Kiss J, et al. Inhibition of DNA methyltransferase leads to increased genomic 5-hydroxymethylcytosine levels in hematopoietic cells. FEBS Open Bio. 2018;8:584–92.
Bibikova M, Barnes B, Tsan C, Ho V, Klotzle B, le JM, et al. High density DNA methylation array with single CpG site resolution. Genomics. 2011;98:288–95.
Pidsley R, Zotenko E, Peters TJ, Lawrence MG, Risbridger GP, Molloy P, et al. Critical evaluation of the Illumina MethylationEPIC BeadChip microarray for whole-genome DNA methylation profiling. Genome Biol. 2016;17:208.
Bell CG, Teschendorff AE, Rakyan VK, Maxwell AP, Beck S, Savage DA. Genome-wide DNA methylation analysis for diabetic nephropathy in type 1 diabetes mellitus. BMC Med Genet. 2010;3:33.
Swan EJ, Maxwell AP, McKnight AJ. Distinct methylation patterns in genes that affect mitochondrial function are associated with kidney disease in blood-derived DNA from individuals with type 1 diabetes. Diabet Med. 2015;32:1110–5.
Qiu C, Hanson RL, Fufaa G, Kobes S, Gluck C, Huang J, et al. Cytosine methylation predicts renal function decline in American Indians. Kidney Int. 2018;93:1417–31.
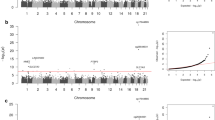
Chu AY, Tin A, Schlosser P, Ko YA, Qiu C, Yao C, et al. Epigenome-wide association studies identify DNA methylation associated with kidney function. Nat Commun. 2017;8:1286.
Ko YA, Mohtat D, Suzuki M, Park A, Izquierdo M, Han S, et al. Cytosine methylation changes in enhancer regions of core pro-fibrotic genes characterize kidney fibrosis development. Genome Biol. 2013;14:R108.
•• Gluck C, et al. Kidney cytosine methylation changes improve renal function decline estimation in patients with diabetic kidney disease. Nat Commun. 2019;10:2461. The authors identified DNA methylation changes in kidneys of patients with DKD. Methylation levels predicted kidney function deline.
•• J. Park et al., Functional methylome analysis of human diabetic kidney disease. JCI Insight, (2019). The authors reported whole genome base-resolution methylation changes in microdissected human DKD kidney samples. The authors demonstrated that dCAS9-based methylation editing of the TNF alpha locus leads to changes in gene expression.
Park J, Shrestha R, Qiu C, Kondo A, Huang S, Werth M, et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science. 2018;360:758–63.
Vojta A, Dobrinić P, Tadić V, Bočkor L, Korać P, Julg B, et al. Repurposing the CRISPR-Cas9 system for targeted DNA methylation. Nucleic Acids Res. 2016;44:5615–28.
Inoue K, Gan G, Ciarleglio M, Zhang Y, Tian X, Pedigo CE, et al. Podocyte histone deacetylase activity regulates murine and human glomerular diseases. J Clin Invest. 2019;129:1295–313.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Tamas Aranyi declares that he has no conflict of interest.
Katalin Susztak reports grant support from GSK, Regeneron, Boehringer Ingelheim, Merck, Bayer, Eli Lilly and Company, and Gilead; and consulting for Chemocentryx, Janssen, and Maze Bio. However, the work is not related to any of the studies supported by industry.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Pathogenesis of Type 2 Diabetes and Insulin Resistance
Rights and permissions
About this article
Cite this article
Aranyi, T., Susztak, K. Cytosine Methylation Studies in Patients with Diabetic Kidney Disease. Curr Diab Rep 19, 91 (2019). https://doi.org/10.1007/s11892-019-1214-6
Published:
DOI: https://doi.org/10.1007/s11892-019-1214-6