Abstract
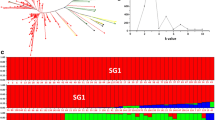
Ethiopia is the center of origin and diversity for Arabica coffee with high morphological diversity between accessions as compared to commercial cultivars. Coffee germplasm collection and molecular characterization are crucial steps towards its conservation, breeding, and development of superior genotypes for various end uses. Hence, this study was initiated with the objective of studying the genetic diversity of Coffea arabica accessions collected from different regions of Ethiopia, using SSR markers. A total of 20 SSR markers were used to genotype 86 accessions and produced a total of 112 alleles, with an average of 5.6 alleles per locus. All the loci across the entire populations were found to be highly polymorphic with a mean of 0.6 PIC value. Average observed heterozygosity and allelic richness across all populations ranged from 0.22 to 0.27 and 3.52–4.26, with a mean of 2.43 and 3.97, respectively. AMOVA showed high variation within population based on geographical origin. The smaller Fst (0.037) observed indicates the presence of lower population genetic differentiation as a result of higher gene flow (Nm = 2.45) across populations and the lowest mean genetic distance (0.21) observed between populations. The UPGMA, PCoA, and structure analysis poorly grouped the individuals into distinct clusters indicating the presence of population admixture. The observed higher genetic variability in all populations indicates that the country has huge coffee genetic diversity which can be used for future coffee improvement. Our results revealed an unexploited highly diverse genetic resource particularly from Omo, Ilubabor, and Benchi Maji that should be considered in future coffee breeding program and germplasm conservation.



Similar content being viewed by others
Availability of data and materials
Full passport data of the 86 coffee samples representing the 10 populations used in the present study is provided in Table 1. The other datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Aerts R, Berecha G, Gijbels P, Hundera K, Van Glabeke S, Vandepitte K, Muys B, Roldan Ruiz I, Honnay O (2013) Genetic variation and risks of introgression in the wild Coffea arabica gene pool in south-western Ethiopian montane rain forests. Evol Appl 6(2):243–252. https://doi.org/10.1111/j.1752-4571.2012.00285.x
Aga E, Bryngelsson T, Bekele E, Balemi S (2003) Genetic diversity of forest arabica coffee (Coffea arabica L.) in Ethiopia as revealed by random amplified polymorphic DNA (RAPD) analysis. Hereditas 138:36–46. https://doi.org/10.1034/j.1601-5223.2003.01636.x
Al-Abdulkader AM, Al-Namazi AA, Al Turki TA, Al-Khuraish MM, Al-Dakhil AI (2018) Optimizing coffee cultivation and its impact on economic growth and export earnings of the producing countries: the case of Saudi Arabia. Saudi J Biol Sci 25(4):776–782. https://doi.org/10.1016/j.sjbs.2017.08.016
Al-Murish TM, Elshafei AA, Al-Doss AA, Barakat MN (2013) Genetic diversity of coffee (Coffea arabica L) in Yemen via SRAP, TRAP and SSR markers. J Food Agric Environ 11(2):411–416. https://doi.org/10.1234/4.2013.4315
Anthony F, Combes MC, Astorga C, Bertrand B, Graziosi G, Lashermes P (2002) the origin of cultivated Coffea arabica L. varieties revealed by AFLP and SSR markers. Theor Appl Genet 104:894–900. https://doi.org/10.1007/s00122-001-0798-8
Balemi S (2007) Genetic diversity analysis of the wild Coffea arabica L. populations from Harenna forest, Bale Mountains of Ethiopia, using Inter Simple Sequence Repeats (ISSR) marker. MSc thesis, Addis Ababa University, Addis Ababa, Ethiopia. P 89
Benti T (2017) Progress in Arabica coffee breeding in Ethiopia: achievements, challenges and prospects. Int J Sci Basic Appl Res 33(2):15–25
Bigirimana J, Njoroge K, Muthomi JW, Gahakwa D, Phiri NA, Gichuru EK, Walyaro DJ (2013) Genetic diversity among disease resistant coffee varieties and cultivars in Rwanda based on RAPD and SSR markers. J Renew Agric 1(6):106–112. https://doi.org/10.12966/jra.09.01.2013
Bunn C, Läderach P, Ovalle-Rivera O, Kirschke D (2015) a bitter cup: climate change profile of global production of Arabica and Robusta coffee. Clim Change 129:89–101. https://doi.org/10.1007/s10584-014-1306-x
Combes MC, Anderzejewski F, Anthony B, Bertrand P, Rovelli G, Graziosi and Lashermes, (2000) characterization of microsatellite loci in Coffea arabica and related coffee species. Mol Ecol 9:1178–1180. https://doi.org/10.1046/j.1365-294x.2000.00954-5.x
Cubry PP, Musali H, Legnate D, Pot FD, Bellis V, Poncet F, Anthony M, Dufour M, Leroy T (2008) diversity in Coffee assessed with SSR markers: structure of the genus Coffea and perspectives for breeding. Genome 51:50–63. https://doi.org/10.1139/G07-096
da Silva BSR, Sant’Ana GC, Chaves CL, Androcioli LG, Ferreira RV, Sera GH, Charmetant P, Leroy T, Pot D, Domingues DS, Pereira LFP (2019) population structure and genetic relationships between Ethiopian and Brazilian Coffea arabica genotypes revealed by SSR markers. Genetica 147(2):205–216. https://doi.org/10.1007/s10709-019-00064-4
Davis AP, Tosh J, Ruch N, Fay MF (2011) growing coffee: Psilanthus (Rubiaceae) subsumed on the basis of molecular and morphological data; implications for the size, morphology, distribution and evolutionary history of Coffea. Bot J Linn Soc 167(4):357–377. https://doi.org/10.1111/j.1095-8339.2011.01177.x
Diniz LEC, Ruas CDF, Carvalho VDP, Torres FM, Ruas EA, Santos MDO, Sera T, Ruas PM (2005) Genetic diversity among forty coffee varieties assessed by RAPD markers associated with restriction digestion. Braz Arch Biol Technol 48(4):511–521. https://doi.org/10.1590/S1516-89132005000500002
Earl DA, Von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conser Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Gadissa F, Tesfaye K, Dagne K, Geleta M (2018) Genetic diversity and population structure analyses of Plectranthus edulis (Vatke) Agnew collections from diverse agro-ecologies in Ethiopia using newly developed EST-SSRs marker system. BMC Genet 19(1):1–15. https://doi.org/10.1186/s12863-018-0682-z
Gebreselassie H, Atinafu G, Degefa M, Ayano A (2018) Arabica Coffee (Coffea arabica L) Hybrid genotypes evaluation for growth characteristics and yield performance under Southern Ethiopian growing condition. Acad Res J Agric Sci Res 6(2):89–96. https://doi.org/10.14662/ARJASR2017.071
Geleta M, Herrera I, Monzón A, Bryngelsson T (2012) Genetic diversity of Arabica coffee (Coffea arabica L.) in Nicaragua as estimated by simple sequence repeat markers. Sci World J 2012:1–11. https://doi.org/10.1100/2012/939820
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity Assessment in crop plants and its recent advances: an overview of its analytical perspectives. A review article. Genet Res Int 2015:1–14. https://doi.org/10.1155/2015/431487
Hussein MAA, Al-Azab AAA, Habib SS, El Sherif FM, El-Garhy HA (2017) Genetic diversity, structure and dna fingerprint for developing molecular ids of Yemeni Coffee (Coffea arabica L) germplasm assessed by SSR markers. Egypt J Plant Breed 21(4):713–736
Kalinowski ST (2005) HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189. https://doi.org/10.1111/j.1471-8286.2004.00845.x
Khanuja SP, Shasany AK, Darokar MP, Kumar S (1999) Rapid isolation of DNA from dry and fresh samples of plants producing large amounts of secondary metabolites and essential oils. Plant Mol Biol Rep 17(1):74–74. https://doi.org/10.1023/A:1007528101452
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I (2015) CLUMPAK: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15:1179–1191. https://doi.org/10.1111/1755-0998.12387
Lashermes P, Combes MC, Cros J, Trouslot P, Anthony F, Charrier A (1995). Origin and genetic diversity of Coffea arabica L. based on DNA molecular markers. In: 16th Conference of ASIC, Kyoto (Japan), pp 528–535
Leberg PL (1991) Influence of fragmentation and bottlenecks on genetic divergence of wild turkey populations. Conserv Biol 5(4):522–530. https://doi.org/10.1111/j.1523-1739.1991.tb00359.x
Liu K, Muse SV (2005) Power marker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21(9):2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Loveless MD, Hamrick JL (1984) Ecological determinants of genetic structure in plant populations. Annu Rev Ecol Syst 15:65–95
Maluf MP, Silvestrini M, Ruggiero LM, Guerreiro Filho O, Colombo C (2005) Genetic diversity of cultivated Coffea arabica inbred lines assessed by RAPD, AFLP and SSR marker systems. Scientia Agricola 62(4):366–373. https://doi.org/10.1590/S010390162005000400010
Marulanda ML, Lopez AM, Isuzu L, Lopez P (2014) Microsatellite isolation and characterization for Colletotrichum spp, causal agent of anthracnose in Andean black berry. Genet Mol Res 13:7673–7685. https://doi.org/10.4238/2014.September.26.5
Moncada P, Couch M (2004) Simple sequence repeats diversity in diploid and tetraploid Coffea species. Genome 47:501–509. https://doi.org/10.1139/g03-129
Motta LB, Soares TCB, Ferrao MAG, Caixeta ET, Lorenzoni RM, Souza Neto JDD (2014) Molecular characterization of arabica and Conilon coffee plants genotypes by SSR and ISSR markers. Braz Arch Biol Technol 57(5):728–735. https://doi.org/10.1590/S1516-8913201402071
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/
Pestana KN, Capucho AS, Caixeta ET, de Almeida DP, Zambolim EM, Cruz CD, Zambolim L, Pereira AA, de Oliveira ACB, Sakiyama NS (2015) Inheritance study and linkage mapping of resistance loci to Hemileia vastatrix in Híbrido de Timor UFV 443–03. Tree Genet Genomes 11(4):72. https://doi.org/10.1007/s11295-015-0903-9
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Rovelli P, Mettulio R, Anthony F, Anzueto F, Lashermes P, Graziosi G (2000) Microsatellites in Coffea arabica L. In: Sera T, Soccol CR, Pandey A, Roussos S (eds) Coffee biotechnology and quality. Springer, Dordrecht, pp 123–133
Sant’Ana GC, Pereira LF, Pot D, Ivamoto ST, Domingues DS, Ferreira RV, Pagiatto NF, da Silva BSR, Nogueira LM, Kitzberger CS, Scholz MB, de Oliveira FF, Sera GH, Padilha LL, Labouisse JP, Guyot R, Charmetant P, Leroy T (2018) Genome-wide association study reveals candidate genes influencing lipids and diterpenes contents in Coffea arabica L. Sci Rep 8(1):465. https://doi.org/10.1038/s41598-017-18800-1
Silvarolla MB, Mazzafera P, Fazuoli LC (2004) Plant biochemistry: a naturally decaffeinated Arabica coffee. Nature 429(6994):826. https://doi.org/10.1038/429826a
Slatkin M (1985) Rare alleles as indicators of gene flow. Evolution 39(1):53–65. https://doi.org/10.1111/j.1558-5646.1985.tb04079.x
Sousa TV, Caixeta ET, Alkimim ER, de Oliveira ACB, Pereira AA, Zambolim L, Sakiyama NS (2017) Molecular markers useful to discriminate Coffea arabica L cultivars with high genetic similarity. Euphytica 213(3):75. https://doi.org/10.1007/s10681-017-1865-9
Steiger D, Nagai C, Moore P, Morden C, Osgood R, Ming R (2002) AFLP analysis of genetic diversity within and among Coffea arabica cultivars. Theor Appl Genet 105(2–3):209–215. https://doi.org/10.1007/s00122-002-0939-8
Tadele S, Mekbib F, Tesfaye K (2014) Genetic diversity of coffee (Coffea arabica L.) landraces from southern Ethiopia as revealed by inter simple sequence repeat marker. Glob Adv Res J Agric Sci 3(1):24–34
Teressa A, Crouzillat D, Petiard V, Brouhan P (2010) Genetic diversity of Arabica coffee (Coffea arabica L) collections. EJAST 1(1):63–79
Tesfaye K (2006) Genetic diversity of wild Coffea arabica populations in Ethiopia as a contribution to conservation and use planning. Ecology and development series. University of Bonn, Germany
Tesfaye K, Govers K, Bekele E, Borsch T (2014) ISSR fingerprinting of Coffea arabica throughout Ethiopia reveals high variability in wild populations and distinguishes them from landraces. Plant Syst Evol 300(5):881–897. https://doi.org/10.1007/s00606-013-0927-2
Tran HT, Lee LS, Furtado A, Smyth H, Henry RJ (2016) Advances in genomics for the improvement of quality in coffee. J Sci Food Agric 96:3300–3312. https://doi.org/10.1002/jsfa.7692
Vieira ESN, von Pinho EVR, Carvalho MG, Esselink GD, Vosman B (2010) Development of microsatellite markers for the identification of Brazilian Coffea arabica varieties. Genet Mol Biol 33(3):507–514. https://doi.org/10.1590/S1415-47572010005000055
Waples RS (1987) A multi species approach to the analysis of gene flow in marine shore fishes. Evolution 41(2):385–400. https://doi.org/10.1111/j.1558-5646.1987.tb05805.x
White Head and Peakall (2015) GENALEX 6.502: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Wright S (1951) The genetic structure of populations. Annu Eugen 15:323–354. https://doi.org/10.1111/j.1469-1809.1949.tb02451.x
Yeh FC, Yang R (1999) POPGENE VERSION 1.31: Microsoft Window-based Freeware for Population Genetic Analysis. Department of Renewable Resources University of Alberta, Canada. pp 1–29
Acknowledgements
The study was conducted by financial support provided by Ethiopian Institute of Agricultural Research Institute, National Agricultural Biotechnology Research Center. The authors are grateful to the Ethiopian Biodiversity Institute for providing the coffee germplasm.
Funding
This work was supported by grants from the Ethiopian Institute of Agricultural Research Institute, National Agricultural Biotechnology Research Center.
Author information
Authors and Affiliations
Contributions
Material preparation, data collection, and analysis were performed by GD and TD. The first draft of the manuscript was written by GD and all authors commented on previous versions of the manuscript. All authors participated in designing the experiment, interpreting the data, drafting and revising the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethics approval and consent to participate
Not Applicable.
Consent for publication
Not Applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dida, G., Bantte, K. & Disasa, T. Molecular characterization of Arabica Coffee (Coffea arabica L.) germplasms and their contribution to biodiversity in Ethiopia. Plant Biotechnol Rep 15, 791–804 (2021). https://doi.org/10.1007/s11816-021-00721-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-021-00721-1