Abstract
Small cysteine-rich peptides (CRPs) are important in plant responses to biotic and abiotic stresses. OsDT11, encoding an 88 amino acid CRP-type protein, has been reported to enhance tolerance to drought in rice (Oryza sativa L.) without negatively affecting other agronomic traits. However, the molecular mechanisms of OsDT11-mediated drought tolerance are still unclear. Here, we performed RNA-Seq analysis to compare the transcriptome profiles between wild-type (WT) and OsDT11-overexpressing (OE) rice lines under drought stress or under control (non-drought) conditions. A total of 1570 and 1421 differentially expressed genes were identified in the OE lines and the WT under drought treatment, respectively, compared to non-drought conditions. Gene ontology analysis indicated that the 430 up-regulated genes in common to both OE and the WT lines were induced for functions related to responses to water deprivation and to abscisic acid (ABA). More than half of these genes had higher expression in the OE than in the WT under drought stress. In the OE, but not in the WT, 294 genes were specifically up-regulated under drought stress and were functionally enriched in starch and sucrose biosynthetic processes and in response to stress. This implies that OsDT11 not only triggers strongly response to drought stress, but also alters several metabolic processes to enhance drought tolerance. Gene expression profiling suggests that OsDT11 confers drought tolerance by mediating an enhanced response to drought stress in an ABA-dependent signaling pathways.







Similar content being viewed by others
References
Chien PS, Nam HG, Chen YR (2015) A salt-regulated peptide derived from the CAP superfamily protein negatively regulates salt-stress tolerance in Arabidopsis. J Exp Bot 66:5301–5313
Cui Y, Li M, Yin X, Song S, Xu G, Gang M, Li C, Peng C, Xia X (2018) OsDSSR1, a novel small peptide, enhances drought tolerance in transgenic rice. Plant Sci 270:85–96
De Hoon MJ, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454
Du H, Wang N, Cui F, Li X, Xiao J, Xiong L (2010) Characterization of the beta-carotene hydroxylase gene DSM2 conferring drought and oxidative stress resistance by increasing xanthophylls and abscisic acid synthesis in rice. Plant Physiol 154:1304–1318
Fujii H, Chinnusamy V, Rodrigues A, Rubio S, Antoni R, Park SY, Cutler SR, Sheen J (2009) In vitro reconstitution of an abscisic acid signalling pathway. Nature 462:660–664
Furihata T, Maruyama K, Fujita Y, Umezawa T, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2006) Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc Natl Acad Sci USA 103:1988–1993
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
Joo J, Lee YH, Song SI (2014) Overexpression of the rice basic leucine zipper transcription factor OsbZIP12 confers drought tolerance to rice and makes seedlings hypersensitive to ABA. Plant Biotechnol Rep 8:431–441
Jeong JS, Kim YS, Baek KH, Jung H, Ha SH, Do Choi Y, Kim M, Reuzeau C, Kim JK (2010) Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol 153:185–197
Jin X, Xue Y, Wang R, Xu R, Bian L, Zhu B, Han H, Pen R, Yao Q (2013) Transcription factor OsAP21 gene increases salt/drought tolerance in transgenic Arabidopsis thaliana. Mol Biol Rep 40:1743–1752
Ju Y, Tian H, Zhang R, Zuo L, Jin G, Xu Q, Ding X, Li X, Chu Z (2017) Overexpression of OsHSP18.0-CI enhances resistance to bacterial leaf streak in rice. Rice 10:12
Kim JS, Mizoi J, Yoshida T, Fujita Y, Nakajima J, Ohori T, Todaka D, Nakashima K, Hirayama T (2011) An ABRE promoter sequence is involved in osmotic stress-responsive expression of the DREB2A gene, which encodes a transcription factor regulating drought-inducible genes in Arabidopsis. Plant Cell Physiol 52:2136–2146
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Lee HY, Jang G, Um T, Kim JK, Lee JS, Choi YD (2015) The soluble ABA receptor PYL8 regulates drought resistance by controlling ABA signaling in Arabidopsis. Plant Biotechnol Rep 9:319–330
Lemoine R, La Camera S, Atanassova R, Dedaldechamp F, Allario T, Pourtau N, Bonnemain JL (2013) Source-to-sink transport of sugar and regulation by environmental factors. Front Plant Sci 4:272
Li X, Han H, Chen M, Yang W, Liu L, Li N, Ding X, Chu Z (2017) Overexpression of OsDT11, which encodes a novel cysteine-rich peptide, enhances drought tolerance and increases ABA concentration in rice. Plant Mol Biol 93:21–34
Li N, Wei ST, Chen J, Yang FF, Kong LG, Chen CX, Ding XH, Chu ZH (2018) OsASR2 regulates the expression of a defence-related gene, Os2H16, by targeting the GT-1 cis-element. Plant Biotechnol J 16(3):771–783
Matsukura S, Mizoi J, Yoshida T, Todaka D, Ito Y, Maruyama K, Shinozaki K, Yamaguchi-Shinozaki K (2010) Comprehensive analysis of rice DREB2-type genes that encode transcription factors involved in the expression of abiotic stress-responsive genes. Mol Genet Genom 283:185–196
Ostrowski M, Kowalczyk S (2015) Plant signaling peptides. Cysteine-rich peptides. Postepy Biochem 61(1):79–92
Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071
Saldanha AJ (2004) Java Treeview—extensible visualization of microarray data. Bioinformatics 20:3246–3248
Santiago J, Dupeux F, Round A, Antoni R, Park SY, Jamin M, Cutler SR, Rodriguez PL (2009) The abscisic acid receptor PYR1 in complex with abscisic acid. Nature 462:665–668
Sun L, Huang L, Hong Y, Zhang H, Song F, Li D (2015) Comprehensive analysis suggests overlapping expression of rice ONAC transcription factors in abiotic and biotic stress responses. Int J Mol Sci 16:4306–4326
Todaka D, Shinozaki K, Yamaguchi SK (2015) Recent advances in the dissection of drought-stress regulatory networks and strategies for development of drought-tolerant transgenic rice plants. Front Plant Sci 6:84
Wang JC, Xu H, Zhu Y, Liu QQ, Cai XL (2013) OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. J Exp Bot 64:3453–3466
Weiner JJ, Peterson FC, Volkman BF, Cutler SR (2010) Structural and functional insights into core ABA signaling. Curr Opin Plant Biol 13:495–502
Xiang Y, Tang N, Du H, Ye H, Xiong L (2008) Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol 148:1938–1952
Xu W, Cui K, Xu A, Nie L, Huang J, Peng S (2015) Drought stress condition increases root to shoot ratio via alteration of carbohydrate partitioning and enzymatic activity in rice seedlings. Acta Physiol Plant 37:2
Xu Y, Yu Z, Zhang D, Huang J, Wu C, Yang G, Yan K, Zhang S, Zheng C (2018) CYSTM, a novel non-secreted cysteine-rich peptide family, involved in environmental stresses in Arabidopsis thaliana. Plant Cell Physiol 59:423–438
Yoon S, Dong KL, In JY, Youn SK, Yang DC, Ju KK (2017) Overexpression of the OsbZIP66 transcription factor enhances drought tolerance of rice plants. Plant Biotechnol Rep 11(1):53–62
Zhao L, Hu Y, Chong K, Wang T (2010) ARAG1, an ABA-responsive DREB gene, plays a role in seed germination and drought tolerance of rice. Ann Bot-london 105:401–409
Zhao Y, Chan Z, Xing L, Liu X, Hou YJ, Chinnusamy V, Wang P, Duan C, Zhu JK (2013) The unique mode of action of a divergent member of the ABA-receptor protein family in ABA and stress signaling. Cell Res 23:1380–1395
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zong W, Tang N, Yang J, Peng L, Ma S, Xu Y, Li G, Xiong L (2016) Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought-resistance-related genes. Plant Physiol 171:2810–2825
Acknowledgements
This study was supported by the National Natural Science Foundation of China (31700219, 31771748, 31872925), the Natural Science Fund for Outstanding Young Scholars of Shandong Province (JQ201807), the Shandong Modern Agricultural Technology and Industry system (SDAIT-17-06), and Shandong provincial key research and development plan (2019GNC106152).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11816_2020_637_MOESM1_ESM.tif
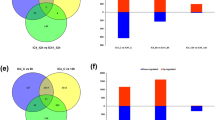
Fig. S1 Venn diagrams of differentially expressed genes (DEGs) in OE and WT under normal and drought stress. DEGs were identified with adjusted p-value <0.01. Venn diagrams representing distribution of up-regulated and down-regulated genes in OE or WT under normal (referred to as OE and WT, respectively) and drought stress (referred to as OED and WTD, respectively) (TIF 99 kb)
11816_2020_637_MOESM2_ESM.tif
Fig. S2. The correlation coefficient between RNA-seq and qRT-PCR data. The fold changes in gene expression by qRT-PCR were transformed to log2 scale. The correlation of the qRT-PCR log2 values (x-axis) with RNA-seq data log2-values (y-axis) of (A) 12 common changed genes and (B) 10 specially changed genes in OE. (TIF 65 kb)
11816_2020_637_MOESM3_ESM.tif
Fig. S3. Negative control that is not regulated by OsDT11. GFP fluorescence assay of expanded ayammetrical NB leaves infiltrated with OsDT11 and the constucts(p4, pLOC_Os04g37950 :: GFP) at 48 hpi. Bars = 200 μm. (TIF 113 kb)
Rights and permissions
About this article
Cite this article
Zhao, M., Ju, Y., Zhao, B. et al. Transcriptional profiling analysis of OsDT11-mediated ABA-dependent signal pathway for drought tolerance in rice. Plant Biotechnol Rep 14, 613–626 (2020). https://doi.org/10.1007/s11816-020-00637-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-020-00637-2