Abstract
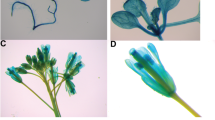
The FLOWERING TIME CONTROL PROTEIN FPA (FPA) gene encodes an RNA recognition motif (RRM) domain protein and plays an important role in flowering time control. Flowering responds to environmental conditions and developmental regulation through a network of signaling pathways. However, a little is known about the functions of autonomous pathway genes in Medicago truncatula. Here, we characterized the M. truncatula FPA (MtFPA) gene expression profiling through quantitative RT-PCR analysis, cellular localization, and functional analyses in transgenic plants. We cloned the FPA gene of M. truncatula based on its sequence similarity with Arabidopsis thaliana FPA. The quantitative RT-PCR analysis of MtFPA expression patterns showed that the MtFPA transcripts accumulated ubiquitously in the roots, leaves, stems, flowers, and pods of M. truncatula. The confocal image analysis of the fusion protein MtFPA:GFP revealed that MtFPA was localized in the nucleus. To examine the function of MtFPA, the 35S::MtFPA transgenic plants were generated in the Arabidopsis late-flowering mutant background, fpa-2. The overexpression of MtFPA accelerated flowering under long day conditions compared to the non-transgenic plants. In MtFPA transgenic lines, the expression of AtFLC was down-regulated, whereas that of the floral integrators, AtFT and AtSOC1, was up-regulated as compared to the control plants. These results suggest that MtFPA is a functional orthologue of Arabidopsis and plays an important role in the regulation of flowering time in legumes, especially in M. truncatula.






Similar content being viewed by others
References
Barker DG, Pfaff T, Moreau D, Groves E, Ruffel S, Lepetit M, Whitehand S, Maillet F, Nair RM, Journet EP (2006) Growing M. truncatula: choice of substrates and growth conditions. In: Mathesius U (ed) The Medicago truncatula handbook. Samuel Roberts Noble Foundation, Ardmore
Bäurle I, Smith L, Baulcombe DC, Dean C (2007) Widespread role for the flowering-time regulators FCA and FPA in RNA-mediated chromatin silencing. Science 318:109–112
Chen SY, Wang ZY, Cai XL (2007) OsRRM, a Spen-like rice gene expressed specifically in the endosperm. Cell Res 17:713–721
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Fernández V, Takahashi Y, Le Gourrierec J, Coupland G (2016) Photoperiodic and thermosensory pathways interact through CONSTANS to promote flowering at high temperature under short days. Plant J 86:426–440
Hecht V, Foucher F, Ferrándiz C, Macknight R, Navarro C, Morin J, Vardy ME, Ellis N, Beltrán JP, Rameau C, Weller JL (2005) Conservation of Arabidopsis flowering genes in model legumes. Plant Physiol 137:1420–1434
Jaudal M, Zhang L, Che C, Putterill J (2015) Three Medicago MtFUL genes have distinct and overlapping expression patterns during vegetative and reproductive development and 35S:MtFULb accelerates flowering and causes a terminal flower phenotype in Arabidopsis. Front Genet 6:50
Kim MY, Kang YJ, Lee T, Lee S-H (2013) Divergence of flowering-related genes in three legume species. Plant Genome 6(3). https://doi.org/10.3835/plantgenome2013.03.0008
Koornneef M, Hanhart CJ, van der Veen JH (1991) A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol Gen Genet 229:57–66
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Lee MO, Kim KP, Kim BG, Hahn JS, Hong CB (2009) Flooding stress-induced glycine-rich RNA-binding protein from Nicotiana tabacum. Mol Cells 27:47–54
Liu D, Cai X (2013) OsRRMh, a Spen-like gene, plays an important role during the vegetative to reproductive transition in rice. J Integr Plant Biol 55:876–887
Moon J, Lee H, Kim M, Lee I (2005) Analysis of flowering pathway integrators in Arabidopsis. Plant Cell Physiol 46:292–299
Mouradov A, Cremer F, Coupland G (2002) Control of flowering time: interacting pathways as a basis for diversity. Plant Cell 14:S111–S130
Ono N, Ishida K, Yamashino T, Nakanishi H, Sato S, Tabata S, Mizuno T (2010) Genome wide characterization of the light-responsive and clock-controlled output pathways in Lotus japonicus with special emphasis of its uniqueness. Plant Cell Physiol 51:1800–1814
Quesada V, Dean C, Simpson GG (2005) Regulated RNA processing in the control of Arabidopsis flowering. Int J Dev Biol 49:773–780
Sawa M, Kay SA (2011) GIGANTEA directly activates flowering locus T in Arabidopsis thaliana. Proc Natl Acad Sci U S A 108:11698–11703
Schomburg FM, Patton DA, Meinke DW, Amasino RM (2001) FPA, a gene involved in floral induction in Arabidopsis, encodes a protein containing RNA-recognition motifs. Plant Cell 13:1427–1436
Simpson GG, Quesada V, Henderson IR, Dijkwel PP, Macknight R, Dean C (2004) RNA processing and Arabidopsis flowering time control. Biochem Soc Trans 32:565–566
Sun TP (2011) The molecular mechanism and evolution of the GA–GID1–DELLA signaling module in plants. Curr Biol 21:R338-R345
Weller JL, Ortega R (2015) Genetic control of flowering time in legumes. Front Plant Sci 6:207
Yamashino T, Yamawaki S, Hagui E, Ishida K, Ueoka-Nakanishi H, Nakamichi N, Mizuno T (2013) Clock-controlled and FLOWERING LOCUS T (FT)-dependent photoperiodic pathway in Lotus japonicus II: characterization of a MicroRNA implicated in the control of flowering time. Biosci Biotechnol Biochem 77:1179–1185
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572
Acknowledgements
We thank Professor Ilha Lee for kindly providing the fpa-2 seeds. This study was supported by a grant from the title of “Novel gene isolation and utilization of rice insertional mutant population (Project No. SA00011643)” in Rural Development Administration’s Agend Program, Jeonju, Republic of Korea. This work was funded by a grant from the National Marine Biodiversity Institute of Korea Research Program (2018M01000).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hwang, HJ., Lim, H., Lee, M.O. et al. Functional conservation of MtFPA, a nucleus-localized RNA-recognition motif-binding protein that regulates flowering time in Medicago truncatula. Plant Biotechnol Rep 12, 39–46 (2018). https://doi.org/10.1007/s11816-018-0470-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-018-0470-2