Abstract
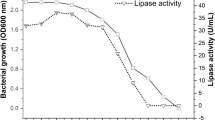
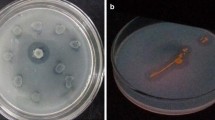
Cold-active enzymes are extremozymes produced by psychrophiles, and have attracted much attention as biocatalysts due to their capacity to resist extreme reaction conditions in industrial processes. This study aimed to isolate bacterial strains from Arctic fjord sediments and screen them for lipase enzyme production. Out of 73 isolates, 8 were identified as good lipase producers. Bacillus cereus I13 exhibited the highest activity of 11.42 U/mL on the 4th day of incubation at 20 °C, along with the large zone of clearance of 27 mm. Optimum conditions for lipase production were identified as pH 7.0, a temperature of 20 °C, and 96 h of incubation, with glucose as a carbon source, yeast as a nitrogen source, and olive oil as a substrate. Bacillus cereus I13 has the potential to be used for the industrial production of cold-active lipase, and this study provides evidence that potent isolates can effectively increase lipase production.
Graphical Abstract











Similar content being viewed by others
Data Availability
The data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Abbreviations
- BLAST:
-
Basic Local Alignment Search Tool
- CAGR:
-
Compound Annual Growth Rate
- DNA:
-
Deoxyribonucleic acid
- IU:
-
International Unit
- MTCC:
-
Microbial Type Culture Collection
- mM:
-
Millimolar
- PCR:
-
Polymerase Chain Reaction
- ROA:
-
Rhodamine Olive oil Agar
- POA:
-
Phenol red Olive oil Agar
- p-NPP:
-
Para-nitro phenyl palmitate
- rRNA:
-
Ribosomal Ribonucleic acid
- UV:
-
Ultra violet
References
Abbas N, Javed J, Abbas Z, Choudry S, Ali S (2017) Lipase Production from Bacillus subtilis using various Agricultural waste. Int J Adv Eng Manag Sci 3(5):239830. https://doi.org/10.24001/ijaems.3.5.1
Adlercreutz P (2017) Comparison of lipases and glycoside hydrolases as catalysts in synthesis reactions. Appl Microbiol Biotechnol 101(2):513–519. https://doi.org/10.1007/s00253-016-8055-x
Akhter K, Karim I, Aziz B, Bibi A, Khan J, Akhtar T (2022) Optimization and characterization of alkaliphilic lipase from a novel Bacillus cereus NC7401 strain isolated from diesel fuel polluted soil. PLoS ONE 17(8):e0273368. https://doi.org/10.1371/journal.pone.0273368
Al-Ghanayem AA, Joseph B (2020) Current prospective in using cold-active enzymes as eco-friendly detergent additive. Appl Microbiol Biotechnol 104(7):2871–2882. https://doi.org/10.1007/s00253-020-10429-x
Alhamdani MA, Alkabbi HJJ (2016) Isolation and identification of lipase producing bacteria from oil-contaminant soil. J Biol, Agric Healthcare 6(20):1–7
Al-Zazaee MMA, Abdu MM, Mahmoud DA, Abdu AAAM (2022) Extraction and Characterization of Lipase Enzymes from Bacillus cereus (MS6) and their Medical & Industrial Applications. Int J Innov Sci Res Technol Special Issue-(2nd ICTSA–2022)
Arifin AR, Kim SJ, Yim JH, Suwanto A, Kim HK (2013) Isolation and biochemical characterization of Bacillus pumilus lipases from the Antarctic. J Microbiol Biotechnol 23(5):661–667. https://doi.org/10.4014/jmb.1212.12040
Arpigny JL, Lamotte J, Gerday C (1997) Molecular adaptation to cold of an Antarctic bacterial lipase. J Mol Catal B Enzym 3(1–4):29–35. https://doi.org/10.1016/S1381-1177(96)00041-0
Awad GE, Mostafa H, Danial EN, Abdelwahed NA, Awad HM (2015) Enhanced production of thermostable lipase from Bacillus cereus ASSCRC-P1 in waste frying oil based medium using statistical experimental design. J Appl Pharmaceut Sci 5(9):007–015. https://doi.org/10.7324/JAPS.2015.50902
Balaji L, Chittoor JT, Jayaraman G (2020) Optimization of extracellular lipase production by halotolerant Bacillus sp. VITL8 using factorial design and applicability of enzyme in pretreatment of food industry effluents. Prep Biochem Biotechnol 50(7):708–716. https://doi.org/10.1080/10826068.2020.1734936
Bharathi D, Rajalakshmi G, Komathi S (2019) Optimization and production of lipase enzyme from bacterial strains isolated from petrol spilled soil. J King Saud Univ-Sci 31(4):898–901. https://doi.org/10.1016/j.jksus.2017.12.018
Bhavani M, Chowdary GV, David M, Archana G (2012) Screening, isolation and biochemical characterization of novel lipase producing bacteria from soil samples. Int J Biol Eng 2(2):18–22. https://doi.org/10.5923/j.ijbe.20120202.03
Choo DW, Kurihara T, Suzuki T, Soda K, Esaki N (1998) A cold-adapted lipase of an Alaskan psychrotroph Pseudomonas sp. strain B11–1: gene cloning and enzyme purification and characterization. Appl Environ Microbiol 64(2):486–491. https://doi.org/10.1128/AEM.64.2.486-491.1998
Das S, Singha R, Rai C, Roy A (2014) Isolation and characterization of bacteria with spoilage potential from some refrigerated foods of West Bengal, India. Int J Curr Microbiol App Sci 3(9):630–639
De Pascale D, Cusano AM, Parrilli E, di Prisco G, Marino G, Tutino ML (2008) The cold-active Lip1 lipase from the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125 is a member of a new bacterial lipolytic enzyme family. Extremophiles 12(3):311–323. https://doi.org/10.1007/s00792-008-0163-9
Demirkan E, Çetinkaya AA, Abdou M (2021) Lipase from new isolate Bacillus cereus ATA179: optimization of production conditions, partial purification, characterization and its potential in the detergent industry. Turk J Biol 45(3):287–300. https://doi.org/10.3906/biy-2101-22
El-Bendary MA, Moharam ME, Mahmoud DA (2015) Economic production of polyethylene modifying lipase enzyme under solid state fermentation using banana peels and sand. BioTechnol: Indian J 11(3):94–101
Fact C, Chandra P, Singh R, Arora PK (2020) Microbial lipases and their industrial applications: a comprehensive review. BioMed Central 1–42. https://doi.org/10.1186/s12934-020-01428-8
Feller G, Thiry M, Gerday C (1991) Nucleotide sequence of the lipase gene lip2 from the Antarctic psychrotroph Moraxella TA 144 and site-specific mutagenesis of the conserved serine and histidine residues. DNA Cell Biol 10(5):381–388. https://doi.org/10.1089/dna.1991.10.381
Ghaima KK, Mohamed AI, Mohamed MM (2014) Effect of some factors on lipase production by Bacillus cereus isolated from diesel fuel polluted soil. Int J Sci Res Publ 4(8):416–420
Ghori MI, Iqbal MJ, Hameed A (2011) Characterization of a novel lipase from Bacillus sp isolated from tannery wastes. Braz J Microbiol 42(1):22–9. https://doi.org/10.1590/S1517-83822011000100003
Guncheva M, Zhiryakova D (2011) Catalytic properties and potential applications of Bacillus lipases. J Mol Catal B Enzym 68:1–2. https://doi.org/10.1016/j.molcatb.2010.09.002
Hassan SW, Abd El Latif HH, Ali SM (2018) Production of cold-active lipase by free and immobilized marine Bacillus cereus HSS: application in wastewater treatment. Front Microbiol 9:2377. https://doi.org/10.3389/fmicb.2018.02377
Ilesanmi OI, Adekunle AE, Omolaiye JA, Olorode EM, Ogunkanmi AL (2020) Isolation, optimization and molecular characterization of lipase producing bacteria from contaminated soil. Scientific African 8:e00279. https://doi.org/10.1016/j.sciaf.2020.e00279
IsiakaAdetunji A, OlufolahanOlaniran A (2018) Optimization of culture conditions for enhanced lipase production by an indigenous Bacillus aryabhattai SE3-PB using response surface methodology. Biotechnol Biotechnol Equipment 32(6):1514–1526. https://doi.org/10.1080/13102818.2018.1514985
Jadhav VV, Pote SS, Yadav A, Shouche YS, Bhadekar RK (2013) Extracellular cold active lipase from the psychrotrophic Halomonas sp. BRI 8 isolated from the Antarctic sea water. Songklanakarin J Sci Technol 35(6)
Javed S, Azeem F, Hussain S, Rasul I, Hussnain M, Riaz M, Afzal M (2018) Bacterial lipases : A review on purification and characterization. Progress in Biophysics and Molecular Biology 132: 23–34. Elsevier Ltd. https://doi.org/10.1016/j.pbiomolbio.2017.07.014
Joseph B, Upadhyaya S, Ramteke P (2011) Production of cold-active bacterial lipases through semisolid state fermentation using oil cakes. Enzyme research 2011. https://doi.org/10.4061/2011/796407
Katiyar P, Pratibha VH, Baghel VS (2017) Isolation, partial purification and characterization of a cold active lipase from Pseudomonas sp. isolated from Satopanth Glacier of Western Himalaya, India. Int J Sci Res Manag (IJSRM) 5(7):6106–6112. https://doi.org/10.18535/ijsrm/v5i7.37
Kuddus M (2018) Cold-active enzymes in food biotechnology : An updated mini-review. J Appl Biol Biotechnol 6:58–63. https://doi.org/10.7324/JABB.2018.60310
Kulakovaa L, Galkin A, Nakayama T, Nishino T, Esaki N (2004) Cold-active esterase from Psychrobacter sp. Ant300: gene cloning, characterization, and the effects of Gly→ Pro substitution near the active site on its catalytic activity and stability. Biochimica et Biophysica Acta (BBA)-Proteins Proteomics 1696(1):59–65. https://doi.org/10.1016/j.bbapap.2003.09.008
Lanka S, Latha JNL (2015) A Short Review on Various Screening Methods to Isolate Potential Lipase Producers : Lipases-the Present and Future Enzymes of Biotech Industry. Int J Biol Chem 9:207–219. https://doi.org/10.17311/ijbc.2015.207.219. (Science Alert)
Lee LP, Karbul HM, Citartan M, Gopinath SC, Lakshmipriya T, Tang TH (2015) Lipase-secreting Bacillus species in an oil-contaminated habitat: promising strains to alleviate oil pollution. BioMed Res Int 2015. https://doi.org/10.1155/2015/820575
Lindsay D, Brözel VS, Mostert JF, Von Holy A (2000) Physiology of dairy associated Bacillus spp. over a wide pH range. Int J Food Microbiol 54(1–2):49–62. https://doi.org/10.1016/S0168-1605(99)00178-6
Mazhar H, Abbas N, Ali S, Sohail A, Hussain Z, Ali SS (2017) Optimized production of lipase from Bacillus subtilis PCSIRNL-39. Afr J Biotech 16(19):1106–1115. https://doi.org/10.5897/AJB2017.15924
Melani NB, Tambourgi EB, Silveira E (2020) Lipases: from production to applications. Sep Purif Rev 49(2):143–158. https://doi.org/10.1080/15422119.2018.1564328
Mondal M, Biswas JK, Tsang YF, Sarkar B, Sarkar D, Rai M, Hooda PS (2019) A wastewater bacterium Bacillus sp. KUJM2 acts as an agent for remediation of potentially toxic elements and promoter of plant (Lens culinaris) growth. Chemosphere 232:439–452. https://doi.org/10.1016/j.chemosphere.2019.05.156
Mukhtar S, Rashid N, Haque MFU, Malik KA (2022) Metagenomic approach for the isolation of novel extremophiles. In Microbial Extremozymes (pp. 55–66). Academic Press. https://doi.org/10.1016/B978-0-12-822945-3.00010-5
RD Nadaf, SC Thimmappa (2020) Optimization of the media components by one factor at a time methodology to enhance lipase production by Bacillus Substilis KUBT4. Int J Sci Technol Res 9(01)
Neethu CS, Rahiman KM, Rosmine E, Saramma AV, Hatha AM (2015) Utilization of agro-industrial wastes for the production of lipase from Stenotrophomonas maltophilia isolated from Arctic and optimization of physical parameters. Biocatal Agric Biotechnol 4(4):703–709. https://doi.org/10.1016/j.bcab.2015.09.002
Novototskaya-Vlasova KA, Petrovskaya LE, Rivkina EM, Dolgikh DA, Kirpichnikov MP (2013) Characterization of a cold-active lipase from Psychrobacter cryohalolentis K5T and its deletion mutants. Biochem Mosc 78(4):385–394. https://doi.org/10.1134/S000629791304007X
Patel GB, Shah KR, Shindhal T, Rakholiya P, Varjani S (2021) Process parameter studies by central composite design of response surface methodology for lipase activity of newly obtained actinomycete. Environ Technol Innov 23:101724. https://doi.org/10.1016/j.eti.2021.101724
Pham VHT, Kim J, Chang S, Chung W (2021) Investigation of Lipolytic-Secreting Bacteria from an Artificially Polluted Soil Using a Modified Culture Method and Optimization of Their Lipase Production. Microorganisms 9(12):2590. https://doi.org/10.3390/microorganisms9122590
Prasad MP, Sethi R (2013) Comparative Studies on the production of Lipase by Bacillus species under various growth parameters. Int J Curr Microbiol Appl Sci 2(11):179–185. https://doi.org/10.12691/jaem-6-1-2
Pulicherla KK, Ghosh M, Kumar PS, Rao KRSS (2011) Psychrozymes-the next generation industrial enzymes. J Marine Sci Res Dev 1(2). https://doi.org/10.4172/2155-9910.1000102
Ramle Z, Rahim RA (2016) Psychrophilic lipase from Arctic bacterium. Trop Life Sci Res 27(supp1):151. https://doi.org/10.21315/2Ftlsr2016.27.3.21
Rampelotto PH (2014) Polar microbiology: Recent advances and future perspectives. Biology 3(1):81–84. https://doi.org/10.3390/biology3010081
Rasol R, Rashidah AR, Nazuha RSN, Smykla J, Maznah WW, Alias SA (2014) Psychrotrophic lipase producers from Arctic soil and sediment samples. Pol J Microbiol 63(1):75
Remonatto D, Miotti RH Jr, Monti R, Bassan JC, de Paula AV (2022) Applications of immobilized lipases in enzymatic reactors: A review. Process Biochem 114:1–20. https://doi.org/10.1016/j.procbio.2022.01.004
Saleena SK, Johnson JI, Joseph JK, Padinchati KK, Abdulla MH (2022) Production and optimization of l-asparaginase by Streptomyces koyangensis SK4 isolated from Arctic sediment. J Basic Microbiol 63(3–4):417–426. https://doi.org/10.1002/jobm.202200116
Salwoom L, Raja Abd Rahman RNZ, Salleh AB, MohdShariff F, Convey P, Pearce D, Mohamad Ali MS (2019) Isolation, characterisation, and lipase production of a cold-adapted bacterial strain Pseudomonas sp. LSK25 isolated from Signy Island Antarctica. Molecules 24(4):715. https://doi.org/10.3390/molecules24040715
Saraswat R, Bhushan I, Gupta P, Kumar V, Verma V (2018) Production and purification of an alkaline lipase from Bacillus sp. for enantioselective resolution of (±)-Ketoprofen butyl ester. 3 Biotech 8:1–12. 10.1007/2Fs13205–018–1506–6
Shaheen M, Ullah I, Rafiq M, Maqsood Ur Rehman M, Shah AA, Hasan F (2020) Purification and characterization of lipase from psychrophilic bacteria Pseudomonas Mandelii htb2 from Batura glacier Pakistan. Appl Ecol Environ Res 18(3):4103. https://doi.org/10.15666/aeer/1803_41034114
Sharma AK, Sharma V, Saxena J (2016) A review on applications of microbial lipases. Int J Biotech Trends Technol 19(1):1–5. https://doi.org/10.14445/22490183/IJBTT-V19P601
Siddiqui KS (2015) Some like it hot, some like it cold: Temperature dependent biotechnological applications and improvements in extremophilic enzymes. Biotechnol Adv 33(8):1912–1922. https://doi.org/10.1016/j.biotechadv.2015.11.001
Sirisha E, Rajasekar N, Narasu ML (2010) Isolation and optimization of lipase producing bacteria from oil contaminated soils. Adv Biol Res 4(5):249–252
Stoytcheva M, Montero G, Zlatev RA, Leon J, Gochev V (2012) Analytical methods for lipases activity determination: a review. Curr Anal Chem 8(3):400–407
Verma S, Meghwanshi GK, Kumar R (2021) Current perspectives for microbial lipases from extremophiles and metagenomics. Biochimie 182:23–36. https://doi.org/10.1016/j.biochi.2020.12.027
Wang J, Li K, He Y, Wang Y, Han X, Yan Y (2019) Enhanced performance of lipase immobilized onto Co2+-chelated magnetic nanoparticles and its application in biodiesel production. Fuel 255:115794. https://doi.org/10.1016/j.fuel.2019.115794
Wi AR, Jeon SJ, Kim S, Park HJ, Kim D, Han SJ, Yim JH, Kim HW (2014) Characterization and a point mutational approach of a psychrophilic lipase from an arctic bacterium. Bacillus Pumilus Biotechnol Lett 36(6):1295–1302. https://doi.org/10.1007/s10529-014-1475-8
Willerding AL, Oliveira LAD, Moreira FW, Germano MG, Chagas AF (2011) Lipase activity among bacteria isolated from Amazonian soils. Enzyme Research 2011. https://doi.org/10.4061/2011/720194
Yasemin S, Arabac N, Güvenmez HK (2017) Production of Cold Active Lipase from Bacillus sp. J Appl Biol Sci 11(2):24–27
Zhang J, Lin S, Zeng R (2007) Cloning, expression, and characterization of a cold-adapted lipase gene from an Antarctic deep-sea psychrotrophic bacterium, psychrobacter sp. 7195. J Microbiol Biotechnol 17(4):604–610
Zhu B, Connolly PJ, Zhang YM, McDonnell ME, Bian H, Lin SC, Macielag MJ (2020) The discovery of azetidine-piperazine di-amides as potent, selective and reversible monoacylglycerol lipase (MAGL) inhibitors. Bioorg Med Chem Lett 30(14):127243. https://doi.org/10.1016/j.bmcl.2020.127243
Acknowledgment
The first author is grateful to the Council of Scientific and Industrial Research (CSIR) for the awarded fellowship. The authors wish to thank the Department of Marine Biology, Microbiology & Biochemistry, Cochin University of Science and Technology for providing the necessary facilities to carry out the work. The authors are thankful to the Director, National Centre for Antarctic and Ocean Research, Ministry of Earth Sciences, for logistic support and facilities for sample, collection, and analysis.
Funding
The Council of Scientific and Industrial Research (CSIR) supported the first author with grant no.09/239(0534)2018-EMR-1.
Author information
Authors and Affiliations
Contributions
HK designed and performed the experiments, analyzed the data, and prepared the manuscript. VS helped to perform the experiments. SK helped in reviewing and finalizing the manuscript. AP helped to analyze the data and in reviewing the manuscript. KKP helped with logistical support and funding. MH conceptualized the project, designed the experiments, and reviewed and finalized the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declared that they have no competing interests.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kattatheyil, H., Sajeela, V., Kabeer, S.S. et al. Screening, optimization, and molecular characterization of cold-active lipase producing Bacillus cereus I13 from Arctic sediments. Biologia 79, 1041–1055 (2024). https://doi.org/10.1007/s11756-024-01610-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-024-01610-y