Abstract
Background
Coronary artery disease (CAD) originates from the blockage of the inner walls of the coronary arteries due to a plaque buildup. Circular RNA (circRNA) circ_0001445 has been reported to be downregulated in patients with a higher coronary atherosclerotic burden. This study is designed to explore the role and mechanism of circ_0001445 on the oxidized low-density lipoprotein (ox-LDL)-induced endothelial cell damage.
Methods
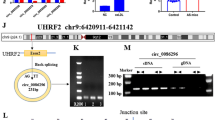
Circ_0001445, microRNA-208b-5p (miR-208b-5p), and ATP-binding cassette sub-family G member 1 (ABCG1) levels were detected by real-time quantitative polymerase chain reaction (RT-qPCR). Inflammatory cytokines levels, cell viability, proliferation, migration were detected by Enzyme-linked immunosorbent assay (ELISA) kits, Cell Counting Kit-8 (CCK-8), 5-ethynyl-2’-deoxyuridine (EdU), and transwell assays, respectively. Protein levels were determined by western blot assay. The binding between miR-208b-5p and circ_0001445 or ABCG1 was predicted by circBank or TargetScan, and then verified by a dual-luciferase reporter, RNA Immunoprecipitation (RIP), and RNA pull-down assays.
Results
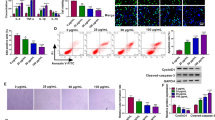
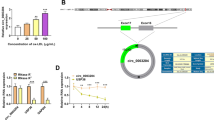
Circ_0001445 and ABCG1 were decreased, and miR-208b-5p was increased in CAD patients and ox-LDL-treated HAECs. Also, circ_0001445 overexpression could weaken ox-LDL-triggered HAEC injury by boosting proliferation, migration, and repressing inflammation and extracellular matrix (ECM). Mechanically, circ_0001445 directly targeted miR-208b-5p. Furthermore, miR-208b-5p mediated the modulation of circ_0001445 in ox-LDL-induced HAEC injury. ABCG1 acted as a direct target of miR-208b-5p, and the downregulation of miR-208b-5p relieved ox-LDL-induced HAEC damage by interacting with ABCG1. Additionally, circ_0001445 regulated ABCG1 expression by sponging miR-208b-5p.
Conclusion
Circ_0001445 could abate ox-LDL-mediated HAEC damage by the miR-208b-5p/ABCG1 axis, providing a novel insight into the pathogenesis and treatment of CAD.







Similar content being viewed by others
Data availability
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
References
Malakar AK, Choudhury D, Halder B, et al. A review on coronary artery disease, its risk factors, and therapeutics. J Cell Physiol. 2019;234(10):16812–23. https://doi.org/10.1002/jcp.28350.
McCullough PA. Coronary artery disease. Clin J Am Soc Nephrol. 2007;2(3):611–6. https://doi.org/10.2215/cjn.03871106.
Li X, Wu C, Lu J, et al. Cardiovascular risk factors in China: a nationwide population-based cohort study. Lancet Public Health. 2020;5(12):e672–81. https://doi.org/10.1016/s2468-2667(20)30191-2.
Liu L. Cardiovascular diseases in China. Biochem Cell Biol. 2007;85(2):157–63. https://doi.org/10.1139/o07-004.
Bentzon JF, Otsuka F, Virmani R, et al. Mechanisms of plaque formation and rupture. Circ Res. 2014;114(12):1852–66. https://doi.org/10.1161/circresaha.114.302721.
Ishida M, Sakuma H. Magnetic resonance of coronary arteries: assessment of luminal narrowing and blood flow in the coronary arteries. J Thorac Imaging. 2014;29(3):155–62. https://doi.org/10.1097/rti.0000000000000081.
Maiolino G, Rossitto G, Caielli P, et al. The role of oxidized low-density lipoproteins in atherosclerosis: the myths and the facts. Mediators Inflamm. 2013. https://doi.org/10.1155/2013/714653.
Yang H, Mohamed AS, Zhou SH. Oxidized low density lipoprotein, stem cells, and atherosclerosis. Lipids Health Dis. 2012;11:85. https://doi.org/10.1186/1476-511x-11-85.
Zhao X, Zhang HW, Sun D, et al. Relation of oxidized-low-density lipoprotein and high-density lipoprotein subfractions in non-treated patients with coronary artery disease. Prostaglandins Other Lipid Mediat. 2019;144: 106345. https://doi.org/10.1016/j.prostaglandins.2019.106345.
Zhang YH, He K, Shi G. Effects of microRNA-499 on the inflammatory damage of endothelial cells during coronary artery disease via the targeting of PDCD4 through the NF-Κβ/TNF-α signaling pathway. Cell Physiol Biochem. 2017;44(1):110–24. https://doi.org/10.1159/000484588.
Zhang C, Ge S, Gong W, et al. LncRNA ANRIL acts as a modular scaffold of WDR5 and HDAC3 complexes and promotes alteration of the vascular smooth muscle cell phenotype. Cell Death Dis. 2020;11(6):435. https://doi.org/10.1038/s41419-020-2645-3.
Peng Y, Calin GA. Crucial role of non-coding RNAs in disease. Cancer Lett. 2018. https://doi.org/10.1016/j.canlet.2018.02.001.
Santosh B, Varshney A, Yadava PK. Non-coding RNAs: biological functions and applications. Cell Biochem Funct. 2015;33(1):14–22. https://doi.org/10.1002/cbf.3079.
Birney E, Stamatoyannopoulos JA, Dutta A, et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447(7146):799–816. https://doi.org/10.1038/nature05874.
Kristensen LS, Andersen MS, Stagsted LVW, et al. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20(11):675–91. https://doi.org/10.1038/s41576-019-0158-7.
Barrett SP, Wang PL, Salzman J. Circular RNA biogenesis can proceed through an exon-containing lariat precursor. Elife. 2015;4: e07540. https://doi.org/10.7554/eLife.07540.
Patop IL, Wüst S, Kadener S. Past, present, and future of circRNAs. Embo J. 2019;38(16):e100836. https://doi.org/10.1525/embj.2018100836.
Han B, Chao J, Yao H. Circular RNA and its mechanisms in disease: From the bench to the clinic. Pharm Ther. 2018;187:31–44. https://doi.org/10.1016/j.pharmthera.2018.01.010.
Altesha MA, Ni T, Khan A, et al. Circular RNA in cardiovascular disease. J Cell Physiol. 2019;234(5):5588–600. https://doi.org/10.1002/jcp.27384.
Pan RY, Zhao CH, Yuan JX, et al. Circular RNA profile in coronary artery disease. Am J Transl Res. 2019;11(11):7115–25.
Wang L, Shen C, Wang Y, et al. Identification of circular RNA Hsa_circ_0001879 and Hsa_circ_0004104 as novel biomarkers for coronary artery disease. Atherosclerosis. 2019;286:88–96. https://doi.org/10.1016/j.atherosclerosis.2019.05.006.
Gao Y, Li G, Fan S, et al. Circ_0093887 upregulates CCND2 and SUCNR1 to inhibit the ox-LDL-induced endothelial dysfunction in atherosclerosis by functioning as a miR-876–3p sponge. Clin Exp Pharm Physiol. 2021. https://doi.org/10.1111/1440-1681.13504.
Vilades D, Martínez-Camblor P, Ferrero-Gregori A, et al. Plasma circular RNA hsa_circ_0001445 and coronary artery disease: performance as a biomarker. Faseb j. 2020;34(3):4403–14. https://doi.org/10.1096/fj.201902507R.
Cai Y, Xu L, Xu C, et al. Hsa_circ_0001445 inhibits ox-LDL-induced HUVECs inflammation, oxidative stress and apoptosis by regulating miRNA-640. Perfusion. 2020. https://doi.org/10.1177/0267659120979472.
Su Q, Lv X. Revealing new landscape of cardiovascular disease through circular RNA-miRNA-mRNA axis. Genomics. 2020;112(2):1680–5. https://doi.org/10.1016/j.ygeno.2019.10.006.
Panda AC. Circular RNAs Act as miRNA Sponges. Adv Exp Med Biol. 2018;1087:67–79. https://doi.org/10.1007/978-981-13-1426-1_6.
Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384–8. https://doi.org/10.1038/nature11993.
Wang W, Li T, Gao L, et al. Plasma miR-208b and miR-499: potential biomarkers for severity of coronary artery disease. Dis Mark. 2019. https://doi.org/10.1155/2019/9842427.
Zhou Q, Schötterl S, Backes D, et al. Inhibition of miR-208b improves cardiac function in titin-based dilated cardiomyopathy. Int J Cardiol. 2017;230:634–41. https://doi.org/10.1016/j.ijcard.2016.12.171.
López-Jiménez E, Rojas AM, Andrés-León E. RNA sequencing and prediction tools for circular RNAs analysis. Adv Exp Med Biol. 2018;1087:17–33. https://doi.org/10.1007/978-981-13-1426-1_2.
Sekar S, Geiger P, Cuyugan L, et al. Identification of circular RNAs using RNA sequencing. J Vis Exp. 2019. https://doi.org/10.3791/59981.
Santer L, Bär C, Thum T. Circular RNAs: a novel class of functional RNA molecules with a therapeutic perspective. Mol Ther. 2019;27(8):1350–63. https://doi.org/10.1016/j.ymthe.2019.07.001.
Ojha R, Nandani R, Chatterjee N, et al. Emerging role of circular RNAs as potential biomarkers for the diagnosis of human diseases. Adv Exp Med Biol. 2018;1087:141–57. https://doi.org/10.1007/978-981-13-1426-1_12.
Zhang S, Wang W, Wu X, et al. Regulatory roles of circular rnas in coronary artery disease. Mol Ther Nucleic Acids. 2020. https://doi.org/10.1016/j.omtn.2020.05.024.
Bach LA. Endothelial cells and the IGF system. J Mol Endocrinol. 2015;54(1):R1-13. https://doi.org/10.1530/jme-14-0215.
Suna G, Wojakowski W, Lynch M, et al. Extracellular matrix proteomics reveals interplay of aggrecan and aggrecanases in vascular remodeling of stented coronary arteries. Circulation. 2018;137(2):166–83. https://doi.org/10.1161/circulationaha.116.023381.
Nielsen SH, Mygind ND, Michelsen MM, et al. Accelerated collagen turnover in women with angina pectoris without obstructive coronary artery disease: an iPOWER substudy. Eur J Prev Cardiol. 2018;25(7):719–27. https://doi.org/10.1177/2047487318758750.
Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505(7483):344–52. https://doi.org/10.1038/nature12986.
Corsten MF, Dennert R, Jochems S, et al. Circulating microRNA-208b and microRNA-499 reflect myocardial damage in cardiovascular disease. Circ Cardiovasc Genet. 2010;3(6):499–506. https://doi.org/10.1161/circgenetics.110.957415.
Liu X, Yuan L, Chen F, et al. Circulating miR-208b: a potentially sensitive and reliable biomarker for the diagnosis and prognosis of acute myocardial infarction. Clin Lab. 2017;63(1):101–9. https://doi.org/10.7754/Clin.Lab.2016.160632.
Goldberg L, Tirosh-Wagner T, Vardi A, et al. Circulating microRNAs: a potential biomarker for cardiac damage, inflammatory response, and left ventricular function recovery in pediatric viral myocarditis. J Cardiovasc Transl Res. 2018;11(4):319–28. https://doi.org/10.1007/s12265-018-9814-0.
Pandzic E, Gelissen IC, Whan R, et al. The ATP binding cassette transporter, ABCG1, localizes to cortical actin filaments. Sci Rep. 2017. https://doi.org/10.1038/srep42025.
Hutchins PM, Heinecke JW. Cholesterol efflux capacity, macrophage reverse cholesterol transport and cardioprotective HDL. Curr Opin Lipidol. 2015;26(5):388–93. https://doi.org/10.1097/mol.0000000000000209.
Rafiei A, Ferns GA, Ahmadi R, et al. Expression levels of miR-27a, miR-329, ABCA1, and ABCG1 genes in peripheral blood mononuclear cells and their correlation with serum levels of oxidative stress and hs-CRP in the patients with coronary artery disease. IUBMB Life. 2021;73(1):223–37. https://doi.org/10.1002/iub.2421.
Rosenson RS, Brewer HB Jr, Ansell BJ, et al. Dysfunctional HDL and atherosclerotic cardiovascular disease. Nat Rev Cardiol. 2016;13(1):48–60. https://doi.org/10.1038/nrcardio.2015.124.
Acknowledgements
Not applicable.
Funding
No funding was received.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
The present study was approved by the ethical review committee of 920 Hospital of Joint Logistics Support Force. Written informed consent was obtained from all enrolled patients.
Consent for publication
Patients agree to participate in this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11748_2022_1799_MOESM2_ESM.tif
Supplementary file2 (TIF 1504 KB) Figure S1 circ_0001445 expression was reduced in CAD patients. (A) RT-qPCR analysis of circ_0001445 expression in serum samples from 26 healthy volunteers and 44 CAD patients (24 Angina pectoris, 6 Myocardial infarction, 9 Asymptomatic myocardial ischemia, 7 Ischemic cardiomyopathy, 4 Sudden death coronary heart disease). (B) ROC curve analysis for the diagnostic value evaluation of circ_0001445 in CAD patients. **P <0.01, ***P <0.001. Figure S2 The effects of circ_0001445 knockdown ox-LDL-mediated inflammatory response, proliferation, migration, and ECM in HAECs. (A) RT-qPCR assay was used to detect circ_0001445 level in HAECs transfected with sh-NC or sh-circ_0001445. (B-G) HAEC were treated with control, ox-LDL, ox-LDL+sh-NC, and ox-LDL+sh-circ_0001445. (B and C) ELISA was adopted for the secretions of TNF-α and IL-6 in treated HAECs. (D and E) Cell proliferative ability was assessed using CCK-8 assay and EdU assay in treated HAECs. (F) Migration ability was detected using Transwell assay in treated HAECs. (G) Protein levels of Collagen type II, MMP13, ADAMTS4 in treated HAECs were determined using western blot assay. **P <0.01, ***P <0.001.
Rights and permissions
About this article
Cite this article
Yang, Z., Liang, X. & Yang, L. Circular RNA circ_0001445 alleviates the ox-LDL-induced endothelial injury in human primary aortic endothelial cells through regulating ABCG1 via acting as a sponge of miR-208b-5p. Gen Thorac Cardiovasc Surg 70, 779–792 (2022). https://doi.org/10.1007/s11748-022-01799-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11748-022-01799-2