Abstract
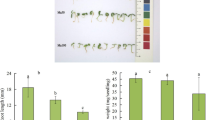
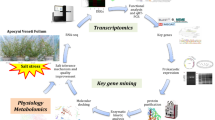
Flavonoids contained in ramie (Boehmeria nivea) root are important resources for officinal use. The plant seedling can grow optimally and develop an easy-harvesting adventitious root system in a hydroponic system. This research was carried out to unravel the regulation mechanism of flavonoid biosynthesis under hydroponic culturing condition, which hitherto remained unknown. Root samples of the ramie plants grown in soil and hydroponic conditions at different developmental stages were collected and pooled for transcriptome sequencing, and the expression levels of flavonoid biosynthesis-related genes and total flavonoid content were quantified. A total of 43,541 unigenes were finally annotated and the expression of 9 key unigenes were confirmed by qRT-PCR. The flavonoid biosynthesis-related gene BnF3H was significantly up-regulated in the roots of hydroponic ramie, and higher total flavonoids content in the hydroponic ramie roots was also observed. The large dataset of transcripts and unigenes provided abundant genetic information for identifying flavonoid biosynthesis genes in ramie roots. Our results suggest flavonoid biosynthesis in ramie roots is positively affected in aquatic environment, and hydroponic culturing could have a potential for enhancing total flavonoids content in ramie roots.





Similar content being viewed by others
References
André CM, Schafleitner R, Legay S, Lefèvre I, Aliaga CAA, Nomberto G, Hoffmann L, Hausman JF, Larondelle Y, Evers D (2009) Gene expression changes related to the production of phenolic compounds in potato tubers grown under drought stress. Phytochemistry 70(9):1107–1116
Bakar MFA, Mohamed M, Rahmat A, Fry J (2009) Phytochemicals and antioxidant activity of different parts of bambangan (Mangifera pajang) and tarap (Artocarpus odoratissimus). Food Chem 113(2):479–483
Beno-Moualem D, Vinokur Y, Prusky D (2001) Cytokinins increase epicatechin content and fungal decay resistance in avocado fruits. J Plant Growth Regul 20(1):95–100
Boudet AM (2007) Evolution and current status of research in phenolic compounds. Phytochemistry 68(22–24):2722–2735
Castellarin SD, Matthews MA, Gaspero GD, Gambetta GA (2007) Water deficits accelerate ripening and induce changes in gene expression regulating flavonoid biosynthesis in grape berries. Planta 227(1):101–112
Chappelle EW, Kim MS, Iii MM (1992) Ratio analysis of reflectance spectra (RARS): an algorithm for the remote estimation of the concentrations of chlorophyll A, chlorophyll B, and carotenoids in soybean leaves. Remote Sens Environ 39(3):239–247
Chen Y, Wang G, Hong W, Cheng C, Zang G, Guo X, Rui HL (2014) Phytochemical profiles and antioxidant activities in six species of ramie leaves. PLoS ONE 9(9):e108140
Conesa A, Götz S (2008) Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genomics 2008(2008):619832
Gao G, Xiong H, Chen J, Chen K, Chen P, Yu C, Zhu A (2018) Hydroponic method for ramie and removal of nitrogen and phosphorus from livestock wastewater. Int J Phytorem 20(6):545–551
Goda K, Sreekala MS, Gomes A, Kaji T, Ohgi J (2006) Improvement of plant based natural fibers for toughening green composites—effect of load application during mercerization of ramie fibers. Compos A Appl Sci Manuf 37(12):2213–2220
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29(7):644
Huang J, Liang X, Xuan Y, Geng C, Li Y, Lu H, Qu S, Mei X, Chen H, Yu T (2017) A reference human genome dataset of the BGISEQ-500 sequencer. Gigascience 6(5):1–9
Jaini R, Wang P, Dudareva N, Chapple C, Morgan J (2017) Targeted metabolomics of the phenylpropanoid pathway in Arabidopsis thaliana using reversed phase liquid chromatography coupled with tandem mass spectrometry. Phytochem Anal 28(4):267–276
Kitamura S (2006) Transport of flavonoids: from cytosolic synthesis to vacuolar accumulation. In: Grotewold E (ed) The science of flavonoids. Springer, New York, pp 123–146
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10(5):236–242
Koes RE, Quattrocchio F, Mol JNM (1994) The flavonoid biosynthetic pathway in plants: function and evolution. BioEssays 16(2):123–132
Lee DG, Cho S, Lee J, Yang S, Jung YS, Kim HB, Cho EJ, Lee S (2015) Quantitative analysis of the flavonoid content in the leaves of Boehmeria nivea and related commercial products. Nat Prod Sci 21(1):66–70
Lee H, Joo N (2012) Optimization of pan bread prepared with ramie powder and preservation of optimized pan bread treated by gamma irradiation during storage. Prev Nutr Food Sci 17(1):53–63
Lin CC, Yen MH, Lo TS, Lin CF (1997) The antiinflammatory and liver protective effects of Boehmeria nivea and B. nivea subsp. nippononivea in rats. Phytomed Int J Phytother Phytopharmacol 4(4):301
Liu M, Li X, Liu Y, Cao B (2013) Regulation of flavanone 3-hydroxylase gene involved in the flavonoid biosynthesis pathway in response to UV-B radiation and drought stress in the desert plant, Reaumuria soongorica. Plant Physiol Biochem Ppb 73(6):161
Morales LO, Tegelberg R, Brosche M, Keinanen M, Lindfors A, Aphalo PJ (2009) Effects of solar UV-A and UV-B radiation on gene expression and phenolic accumulation in Betula pendula leaves. Comp Biochem Physiol A Mol Integr Physiol 153:2–923
Madhava Rao KV, Sresty TV (2000) Antioxidative parameters in the seedlings of pigeonpea (Cajanus cajan (L.) Millspaugh) in response to Zn and Ni stresses. Plant Sci 157(1):113–128
Mak SST, Gopalakrishnan S, Caroe C, Geng C, Liu S, Sinding MS, Kuderna LFK, Zhang W, Fu S, Vieira FG, Germonpre M, Bocherens H, Fedorov S, Petersen B, Sicheritz-Ponten T, Marques-Bonet T, Zhang G, Jiang H, Gilbert MTP (2017) Comparative performance of the BGISEQ-500 vs Illumina HiSeq2500 sequencing platforms for palaeogenomic sequencing. Gigascience 6(8):1–13
Morozova O, Hirst M, Marra MA (2009) Applications of new sequencing technologies for transcriptome analysis. Annu Rev Genomics Hum Genet 10(10):135
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621
Nishihara M, Nakatsuka T, Yamamura S (2005) Flavonoid components and flower color change in transgenic tobacco plants by suppression of chalcone isomerase gene. FEBS Lett 579(27):6074–6078
Park JS, Choung MG, Kim JB, Hahn BS, Kim JB, Bae SC, Roh KH, Kim YH, Cheon CI, Sung MK (2007) Genes up-regulated during red coloration in UV-B irradiated lettuce leaves. Plant Cell Rep 26(4):507–516
Peng M, Shahzad R, Gul A, Subthain H, Shen S, Lei L, Zheng Z, Zhou J, Lu D, Wang S, Nishawy E, Liu X, Tohge T, Fernie AR, Luo J (2017) Differentially evolved glucosyltransferases determine natural variation of rice flavone accumulation and UV-tolerance. Nat Commun 8(1):1975
Quettierdeleu C, Gressier B, Vasseur J, Dine T, Brunet C, Luyckx M, Cazin M, Cazin JC, Bailleul F, Trotin F (2000) Phenolic compounds and antioxidant activities of buckwheat (Fagopyrum esculentum Moench) hulls and flour. J Ethnopharmacol 72(1–2):35
Sparvoli F, Martin C, Scienza A, Gavazzi G, Tonelli C (1994) Cloning and molecular analysis of structural genes involved in flavonoid and stilbene biosynthesis in grape (Vitis vinifera L.). Plant Mol Biol 24(5):743
Strauss SY, Whittal JB (2006) Non-pollinator agents of selection on floral traits. Ecol Evol Flowers 208:120–139
Von Wettberg EJ, Stanton ML, Whittall JB (2010) How anthocyanin mutants respond to stress: the need to distinguish between stress tolerance and maximal vigour. Evol Ecol Res 12(4):457–476
WinkelShirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126(2):485–493
Yao L, Wang J, Li B, Meng Y, Ma X, Si E, Ren P, Yang K, Shang X, Wang H et al (2017) Transcriptome sequencing and comparative analysis of differentially-expressed isoforms in the roots of Halogeton glomeratus under salt stress. Gene 649:159–168
Zhang Y, Xia H, Yuan M, Zhao C, Li A, Wang X (2012) Cloning and expression analysis of peanut (Arachis hypogaea L.) CHI gene. Electron J Biotechnol 15(15):5–5
Acknowledgements
This work was supported by the Agricultural Science and Technology Innovation Project of Chinese under Grant Number CAAS-ASTIP-2018, and China Agricultural Research System under Grant Number CARS-16 and National Natural Science Foundation of China under Grant Number 31701477.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by P. Wojtaszek.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.



Rights and permissions
About this article
Cite this article
Gao, G., Chen, P., Chen, J. et al. Expression profile of genes involved in ramie flavonoids biosynthesis pathway and regulation of flavanone 3-hydroxylase (BnF3H) in response to aquatic environment. Acta Physiol Plant 42, 60 (2020). https://doi.org/10.1007/s11738-020-03036-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-020-03036-w