Abstract
While the functions of vacuolar processing enzymes (VPE) in plant responses to pathogens are now better established, their possible roles against insect herbivores need validation. Inspired by an observation that βVPE and δVPE represent the most highly upregulated transcripts in leaves of Solanum bulbocastanum and its backcross progenies that were highly resistant to the phloem-feeding green peach aphids (Myzus persicae), we used a reverse genetic approach in Arabidopsis to investigate whether T-DNA insertion mutants for specific βVPE and δVPE loci would exhibit compromised defenses at the whole-plant, physiological, and molecular levels. We hypothesized that impaired expression of VPE genes would lead to hyper-susceptibility if VPE-mediated processes were essential for basal defenses, given that Arabidopsis is naturally susceptible. Compared to the wild-type, βvpe and δvpe mutants exhibited an early onset of leaf curling and necrosis, rapid aphid population build-up, and significant physiological tissue injuries as a result of herbivory. This indicates that basal defenses were compromised by the loss-of-functions of either VPE-encoding gene. VPE co-expression networks were comprised of genes involved in sucrose metabolism and transport (AtSUC5, AtSUS3, and invertases), efflux and oxidative detoxification proteins, and modulators of gene-for-gene-type disease response linked to sucrose metabolism (AtSWEET13 and AtSWEET15). These results suggest that VPEs play a role in basal defenses by mediating mechanisms of detoxification and metabolite availability in host tissues to counteract the rapid depletion caused by the phloem sucker. These processes might be important as first line of defense to delay the progression of perturbations caused by herbivory.







Similar content being viewed by others
References
Allie F, Pierce EJ, Okoniewski MJ, Rey C (2014) Transcriptional analysis of South African cassava mosaic virus-infected susceptible and tolerant landraces of cassava highlights differences in resistance, basal defense and cell wall associated genes during infection. BMC Genom 15:1006. doi:10.1186/1471-2164-15-1006
Ballou SM, Yun KY, Cheng C, de los Reyes BG (2007) Cold sensitivity gradient in tuber-bearing Solanum based on physiological and transcript profiles. Crop Sci 47:2027. doi:10.2135/cropsci2007.01.0039sc
Baxter A, Mittler R, Suzuki N (2014) ROS as key players in plant stress signaling. J Exp Bot 65:1229–1240. doi:10.1093/jxb/ert375
Bernoux M, Timmers T, Jauneau A, Brière C, de Wit PJ, Marco Y, Deslandes L (2008) RD19, an Arabidopsis cysteine protease required for RRS1-R-mediated resistance, is relocalized to the nucleus by the Ralstonia solanacearum PopP2 effector. Plant Cell 20:2252–2264. doi:10.1105/tpc.108.058685
Bowler C, Fluhr R (2000) The role of calcium and activated oxygens as signals for controlling cross-tolerance. Trends Plant Sci 5:241–246. doi:10.1016/S1360-1385(00)01628-9
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Wittwer CT (2009) The MIQE guidelines: minimum Information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. doi:10.1373/clinchem.2008.112797
Candat A, Paszkiewicz G, Neveu M, Gautier R, Logan DC, Avelange-Macherel MH, Macherel D (2014) The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26:1–20. doi:10.1105/tpc.114.127316
Carbon S, Ireland A, Mungall CJ, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group (2009) AmiGO: online access to ontology and annotation data. Bioinformatics 25:288–289
Chen HH, Li PH (1980) Characteristics of cold acclimation and deacclimation in tuber-bearing Solanum species. Plant Physiol 65:1146–1148. doi:10.1104/pp.65.6.1146
Chen Y, Chen P, de los Reyes BG (2006) Differential responses of cultivated and wild species of soybean to dehydration stress. Crop Sci 46:2041–2046. doi:10.2135/cropsci2005.12.0466
Chen LQ, Qu XQ, Hou BH, Sosso D, Osorio S, Fernie AR, Frommer WB (2012) Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335:207–211. doi:10.1126/science.1213351
Chen LQ, Lin IW, Qu XQ, Sosso D, McFarlane HE, Londoño A, Frommer WB (2015) A cascade of sequentially expressed sucrose transporters in the seed coat and endosperm provides nutrition for the Arabidopsis embryo. Plant Cell 27:607–619. doi:10.1105/tpc.114.134585
De Vos M, Kim JH, Jander G (2007) Biochemistry and molecular biology of Arabidopsis-aphid interactions. BioEssays 29:871–883. doi:10.1002/bies.20624
Del Pozo O, Lam E (1998) Caspases and programmed cell death in the hypersensitive response of plants to pathogens. Curr Biol 8:1129–1132. doi:10.1016/S0960-9822(98)70469-5
Dixon AFG (1998) Aphid ecology: an optimization approach, 2nd edn. Chapman and Hall, New York
Dutta A, Chan SHP, Pauli NT, Raina R (2015) HYPERSENSITIVE RESPONSE-LIKE LESIONS-1 codes for AtPPT1 and regulates accumulation of ROS and defense against bacterial pathogen Pseudomonas syringae in Arabidopsis thaliana. Antioxid Redox Signal 22:785–796. doi:10.1089/ars.2014.5963
Greenberg JT (1996) Review programmed cell death: a way of life for plants. Proc Natl Acad Sci USA 93:12094–12097
Hara-Nishimura I, Hatsugai N (2011) The role of vacuole in plant cell death. Cell Death Differ 18:1298–1304. doi:10.1038/cdd.2011.70
Hara-Nishimura I, Hatsugai N, Nakaune S, Kuroyanagi M, Nishimura M (2005) Vacuolar processing enzyme: an executor of plant cell death. Curr Opin Plant Biol 8:404–408. doi:10.1016/j.pbi.2005.05.016
Hatsugai N, Kuroyanagi M, Yamada K, Meshi T, Tsuda S, Kondo M, Nishimura M et al (2004) A plant vacuolar protease, VPE, mediates virus-induced hypersensitive cell death. Science 305:855–858. doi:10.1126/science.1099859
Hatsugai N, Yamada K, Goto-Yamada S, Hara-Nishimura I (2015) Vacuolar processing enzyme in plant programmed cell death. Front Plant Sci 6:1–11. doi:10.3389/fpls.2015.00234
Hayashi Y, Yamada K, Shimada T, Matsushima R, Nishizawa NK, Nishimura M, Hara-Nishimura I (2001) A proteinase-storing body that prepares for cell death or stresses in the epidermal cells of Arabidopsis. Plant Cell Physiol 42:894–899. doi:10.1093/pcp/pce144
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300. doi:10.1093/nar/27.1.297
Iakimova ET, Sobiczewski P, Michalczuk L, Wegrzynowicz-Lesiak E, Mikiciński A, Woltering EJ (2013) Morphological and biochemical characterization of Erwinia amylovora-induced hypersensitive cell death in apple leaves. Plant Physiol Biochem 63:292–305
Kerchev PI, Fenton B, Foyer CH, Hancock RD (2012) Plant responses to insect herbivory: interactions between photosynthesis, reactive oxygen species and hormonal signaling pathways. Plant Cell Environ 35:441–453. doi:10.1111/j.1365-3040.2011.02399.x
Kerchev PI, Karpińska B, Morris JA, Hussain A, Verrall SR, Hedley PE, Hancock RD (2013) Vitamin C and the abscisic acid-insensitive 4 transcription factor are important determinants of aphid resistance in Arabidopsis. Antioxid Redox Signal 18:2091–2105. doi:10.1089/ars.2012.5097
Kim NH, Hwang BK (2015) Pepper pathogenesis-related protein 4c is a plasma membrane-localized cysteine protease inhibitor that is required for plant cell death and defense signaling. Plant J 81:81–94. doi:10.1111/tpj.12709
Kinoshita T, Yamada K, Hiraiwa N, Kondo M, Nishimura M, Hara-Nishimura I (1999) Vacuolar processing enzyme is up-regulated in the lytic vacuoles of vegetative tissues during senescence and under various stressed conditions. Plant J 19(1):43–53. doi:10.1046/j.1365-313X.1999.00497.x
Kuroyanagi M, Yamada K, Hatsugai N, Kondo M, Nishimura M, Hara-Nishimura I (2005) Vacuolar processing enzyme is essential for mycotoxin-induced cell death in Arabidopsis thaliana. J Biol Chem 280:32914–32920. doi:10.1074/jbc.M504476200
Lam E (2004) Controlled cell death, plant survival and development. Nat Rev Mol Cell Biol 5:305–315. doi:10.1038/nrm1358
Lan A, Huang J, Zhao W, Peng Y, Chen Z, Kang D (2013) A salicylic acid-induced rice (Oryza sativa L.) transcription factor OsWRKY77 is involved in disease resistance of Arabidopsis thaliana. Plant Biol 15:452–461. doi:10.1111/j.1438-8677.2012.00664.x
Li Z, Yue H, Xing D (2012) MAP Kinase 6-mediated activation of vacuolar processing enzyme modulates heat shock-induced programmed cell death in Arabidopsis. New Phytol 195:85–96. doi:10.1111/j.1469-8137.2012.04131.x
Liu WX, Zhang FC, Zhang WZ, Song LF, Wu WH, Chen YF (2013) Arabidopsis Di19 functions as a transcription factor and modulates PR1, PR2, and PR5 expression in response to drought stress. Mol Plant 6:1487–1502. doi:10.1093/mp/sst031
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Llorca CM, Potschin M, Zentgraf U (2014) bZIPs and WRKYs: two large transcription factor families executing two different functional strategies. Front Plant Sci 5:169. doi:10.3389/fpls.2014.00169
Macho AP, Zipfel C (2014) Plant PRRs and the activation of innate immune signaling. Mol Cell 54:263–272. doi:10.1016/j.molcel.2014.03.028
Maffei MF, Mithofer A, Boland W (2007) Before gene expression: early events in plant–insect interaction. Trends Plant Sci 12:310–316
Martin B, Collar JL, Tjallingii WF, Fereres A (1997) Intracellular ingestion and salivation by aphids may cause the acquisition and inoculation of non-persistently transmitted plant viruses. J Gen Virol 78:2701–2705
Mittler R, Vanderauwera S, Suzuki N, Miller G, Tognetti VB, Vandepoele K, Van Breusegem F (2011) ROS signaling: the new wave? Trends Plant Sci 16:300–309. doi:10.1016/j.tplants.2011.03.007
Moran PJ, Thompson GA (2001) Molecular responses to aphid feeding in Arabidopsis in relation to plant defense pathways. Plant Physiol 125:1074–1085. doi:10.1104/pp.125.2.1074
Mukherjee A (2012) Interspecific introgression of aphid and late blight resistance mechanisms from Solanum bulbocastanum to Solanum tuberosum, Ph.D. Dissertation. University of Maine, Orono
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Noguero M, Atif RM, Ochatt S, Thompson RD (2013) The role of the DNA-binding One Zinc Finger (DOF) transcription factor family in plants. Plant Sci 209:32–45. doi:10.1016/j.plantsci.2013.03.016
Obayashi T, Kinoshita K, Nakai K, Shibaoka M, Hayashi S, Saeki M, Ohta H (2007) ATTED-II: a database of co-expressed genes and cis elements for identifying co-regulated gene groups in Arabidopsis. Nucleic Acids Res 35:4–6. doi:10.1093/nar/gkl783
Obayashi T, Okamura Y, Ito S, Tadaka S, Aoki Y, Shirota M, Kinoshita K (2014) ATTED-II in 2014: evaluation of gene coexpression in agriculturally important plants. Plant Cell Physiol 55:e6. doi:10.1093/pcp/pct178
Ohnuma T, Numata T, Osawa T, Mizuhara M, Lampela O, Juffer AH, Fukamizo T (2011) A class V chitinase from Arabidopsis thaliana: gene responses, enzymatic properties, and crystallographic analysis. Planta 234:123–137. doi:10.1007/s00425-011-1390-3
Olvera-Carrillo Y, Campos F, Reyes JL, Garciarrubio A, Covarrubias AA (2010) Functional analysis of the group-4 late embryogenesis abundant proteins reveals their relevance in the adaptive response during water deficit in Arabidopsis. Plant Physiol 154:373–390. doi:10.1104/pp.110.158964
Petrov V, Hille J, Mueller-Roeber B, Gechev TS (2015) ROS-mediated abiotic stress-induced programmed cell death in plants. Front Plant Sci 6:1–16. doi:10.3389/fpls.2015.00069
Rojo E, Martin R, Carter C, Zouhar J, Pan S, Plotnikova J, Jin H et al (2004) VPEγ exhibits a caspase-like activity that contributes to defense against pathogens. Proc Natl Acad Sci USA 14:1897–1906. doi:10.1016/j.cub.2004.09.056
Romeis T, Ludwig AA, Martin R, Jones JD (2001) Calcium-dependent protein kinases play an essential role in a plant defence response. EMBO J 20:5556–5567. doi:10.1093/emboj/20.20.5556
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108. doi:10.1038/nprot.2008.73
Schweizer F, Fernández-Calvo P, Zander M, Diez-Diaz M, Fonseca S, Glauser G, Reymond P (2013) Arabidopsis basic helix–loop–helix transcription factors MYC2, MYC3, and MYC4 regulate glucosinolate biosynthesis, insect performance, and feeding behavior. Plant Cell 25:3117–3132. doi:10.1105/tpc.113.115139
Seo PJ, Park JM, Kang SK, Kim SG, Park CM (2011) An Arabidopsis senescence-associated protein SAG29 regulates cell viability under high salinity. Planta 233:189–200. doi:10.1007/s00425-010-1293-8
Shimada T, Yamada K, Kataoka M, Nakaune S, Koumoto Y, Kuroyanagi M, Tabata S (2003) Vacuolar processing enzymes are essential for proper processing of seed storage proteins in Arabidopsis thaliana. J Biol Chem 278:32292–32299. doi:10.1074/jbc.M305740200
Streubel J, Pesce C, Hutin M, Koebnik R, Boch J, Szurek B (2013) Five phylogenetically close rice SWEET genes confer TAL effector-mediated susceptibility to Xanthomonas oryzae pv. oryzae. New Phytol 200:808–819. doi:10.1111/nph.12411
Sukuraman NP, Weiser CJ (1972) Freezing injury in potato leaves. Plant Physiol 50:564–567. doi:10.1104/pp.50.5.564
Thaler JS, Humphrey PT, Whiteman NK (2012) Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci 17:260–270. doi:10.1016/j.tplants.2012.02.010
The Gene Ontology Consortium (2015) Gene Ontology Consortium: going forward. Nucleic Acids Res 43:D1049–D1056. doi:10.1093/nar/gku1179
Van der Does D, Leon-Reyes A, Koornneef A, Van Verk MC, Rodenburg N, Pauwels L, Pieterse CMJ (2013) Salicylic acid suppresses jasmonic acid signaling downstream of SCFCOI1-JAZ by targeting GCC promoter motifs via transcription factor ORA59. Plant Cell 25:744–761. doi:10.1105/tpc.112.108548
Vitale A (2013) More players in the plant unfolded response. Proc Natl Acad Sci USA 110:19189–19190. doi:10.1073/pnas.1320254110
Wang Z, Tan X, Zhang Z, Gu S, Li G, Shi H (2012) Defense to Sclerotinia sclerotiorum in oilseed rape is associated with the sequential activations of salicylic acid signaling and jasmonic acid signaling. Plant Sci 184:75–82. doi:10.1016/j.plantsci.2011.12.013
Will T, van Bel AJE (2006) Physical and chemical interactions between aphids and plants. J Exp Bot 57:729–737. doi:10.1093/jxb/erj089
Will T, Vilcinskas A (2015) The structural sheath protein of aphids is required for phloem feeding. Insect Biochem Mol Biol 57:34–40. doi:10.1016/j.ibmb.2014.12.005
Will T, Tjallingii WF, Thönnessen A, van Bel AJE (2007) Molecular sabotage of plant defense by aphid saliva. Proc Natl Acad Sci USA 104:10536–10541. doi:10.1073/pnas.0703535104
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV, Provart NJ (2007) An “electronic fluorescent pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS One 2(8):e718. doi:10.1371/journal.pone.0000718
Wu S, Shan L, He P (2014) Microbial signature-triggered plant defense responses and early signaling mechanisms. Plant Sci 228:118–126. doi:10.1016/j.plantsci.2014.03.001
Xu Q, Zhang L (2009) Plant caspase-like proteases in plant programmed cell death. Plant Signal Behav 4:902–904. doi:10.1104/pp.108.125625
Yamada K, Shimada T, Nishimura M, Hara-Nishimura I (2005) A VPE family supporting various vacuolar functions in plants. Physiol Plant 123:369–375. doi:10.1111/j.1399-3054.2005.00464.x
Yanagisawa S (2002) The Dof family of plant transcription factors. Trends Plant Sci 7:555–560. doi:10.1093/pcp/pch055
Zhang H, Zheng X, Zhang Z (2010a) The role of vacuolar processing enzymes in plant immunity. Plant Signal Behav 5:1565–1567. doi:10.4161/psb.5.12.13809
Zhang H, Dong S, Wang M, Wang W, Song W, Dou X, Zheng X (2010b) The role of vacuolar processing enzyme (VPE) from Nicotiana benthamiana in the elicitor-triggered hypersensitive response and stomatal closure. J Exp Bot 61:3799–3812. doi:10.1093/jxb/erq189
Zhong R, Ye ZH (2012) MYB46 and MYB83 bind to the SMRE sites and directly activate a suite of transcription factors and secondary wall biosynthetic genes. Plant Cell Physiol 53:368–380. doi:10.1093/pcp/pcr185
Acknowledgements
The authors thank Myles Butler, Jason Cooney, and Siobhan Cusack for their technical assistance to this project. Promoter sequence analysis was performed using the supercomputing facilities at the National Institute of Genetics, Mishima, Japan. This project was supported by the Maine Agricultural and Forest Experiment Station, University of Maine (Publication 3412) and the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest in all aspects of the body of works included in this publication.
Additional information
Communicated by E. Kuzniak-Gebarowska.
Electronic supplementary material
Below is the link to the electronic supplementary material.
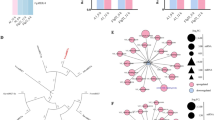
Fig. S1. Putative orthologs of βVPE and δVPE of Arabidopsis were among the most abundant transcripts in a suppression subtractive hybridization library obtained from the leaves of the aphid-resistant wild species of potato S. bulbocastanum (Acc. 243510) infested with M. persicae (data not shown). Differential expression of the putative βVPE and δVPE genes was confirmed by quantitative real-time PCR analysis across a comparison panel that included the resistant donor parent S. bulbocastanum (Sbu243510), susceptible recurrent parent S. tuberosum cv. Ivory Crisp, and two of their contrasting sibling backcross derivatives (Sbu8.5 = resistant; Sbu8.15 = susceptible). Backcross lines were derived from a somatic hybrid between S. tuberosum and S. bulbocastanum backcrossed for four generations to Ivory Crisp. Resistant Sbu8.5 exhibited the βVPE (A) and δVPE (B) expression profile of the donor parent Sbu243510. Susceptible Sbu8.15 exhibited the βVPE and δVPE expression profile of the recurrent parent Ivory Crisp. Relative expression values represent the mean + standard error (n = 3).
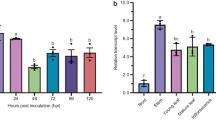
Fig. S2. Relative expression of βVPE and δVPE in the leaves of WT Arabidopsis plants in response to aphids (M. persicae) and other factors reported to affect their expression such as Sclerotinia sclerotiorum (pathogen) and various chemical signals such as salicylic acid (SA) and methyl jasmonate (JA). Relative expression values represent the mean + standard error (n = 4).
Rights and permissions
About this article
Cite this article
Alpuerto, J.B., Mukherjee, A., Kitazumi, A. et al. Impaired expressions of the beta and delta isoforms of vacuolar processing enzymes compromise the basal defenses of Arabidopsis thaliana against the phloem-feeding insect Myzus persicae . Acta Physiol Plant 39, 233 (2017). https://doi.org/10.1007/s11738-017-2529-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-017-2529-z