Abstract
The study was undertaken as an attempt at explaining interrelations between the presently found much diversified isoforms of ACC synthase, a key enzyme of the ethylene synthesis pathway in higher plants. The results of our study analysed on a background of current knowledge implied some not yet discussed, physiological and evolutionary aspects of the plant ACS isozymes.
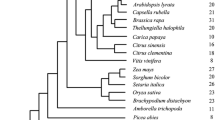
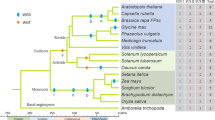
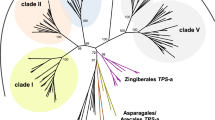
The computer methods applied are based on analysis of phenetic data. The subjects of the study were 159 synthases from higher plants and the only one known from the fungus Penicillium citrinum. The phenograms of 95 full-length sequence ACC synthases from higher plants and the synthase from Penicillium citrinum were made for cDNA and polypeptides by the two different techniques: UPGMA and Kitsch’s method and were almost identical. They indicate the presence of three significantly different types of ACS plant isozymes denoted as type A, type B and A3 group. A number of arguments are given showing that the synthases denoted as A1 and A2 group, hitherto treated by many authors as separate evolutionary lineages, are of the same type A. The presence of a new poorly recognised and probably evolutionary separate group of synthases, denoted as A3, is for the first time evidenced. The type B isozymes are shown to comprise two distinct groups B1 and B2, and B1 group can be divided into subgroups. A comparison of the proportion of genes encoding different type synthases in taxa distinguished by molecular systematicians has shown that similar sets and numbers of genes of type A and A3 group synthases are conserved in the genomes of eudicots and noneudicots. In the genomes of noneudicots the genes of B1 (B1a and B1b) group synthaseswere not found. The genes of B1(B1a and B1b) group in full diversity were established to occur in Rosidae and as B1a subgroup in Asteridae, the subclasses representing eudicots.
It is evidenced that the C-terminal region of the enzyme, hitherto treated as highly variable, and the last aa residue are conserved within the B type and A1 synthases. The acidic character of the polypeptides and the lack of conservation of the last aa residue in the synthases of A2 group are explained by the loss of the 3′-terminal fragment by some A type genes.
The C-terminal region of B type and A1 group synthases was found to contain a fragment similar to the so-called MAPK-docking domain, which suggests that these isozymes are controlled by the processes of phosphorylation.
The aa residues forming the catalytically important three-dimensional structure called the hydrophobic pocket for the adenine ring of SAM can differ slightly in the particular type or groups of ACS. Moreover, A3 group differs from the other synthase groups by the aa residues required for correct orientation of PLP in the active site. Probably, particular ACS groups can differ in the kinetic features and the half-life.
Similar content being viewed by others
Abbreviations
- aa:
-
amino acid
- ACC:
-
1-aminocyclopropane-1-carboxylic acid
- ACO:
-
1-aminocyclopropane-1-carboxylate oxidase
- ACS:
-
1-aminocyclopropane-1-carboxylate synthase
- MACC:
-
1-(malonylamino) cyclopropane-1-carboxylic acid
- MAPK:
-
mitogen-activated protein kinase
- MAPKK:
-
MAPK kinase
- PLP:
-
pyridoxal 5′-phosphate
- SAM:
-
S-adenosyl-L-methionine
- UPGMA:
-
unweighted pair group method with arithmetic averages
- Ach :
-
Actinidia chinensis
- Acher :
-
Annona cherimola
- AD :
-
Actinidia deliciosa
- Ama :
-
Antirrhinum majus
- AS :
-
Asparagus officinalis
- AT :
-
Arabidopsis thaliana
- Av :
-
Averrhoa carambola
- BJ :
-
Brassica juncea
- BO :
-
Brassica oleracea
- BP :
-
Betula pendula
- CA :
-
Capsicum annum
- Cpap :
-
Carica papaya
- Csi :
-
Citrus sinensis
- CM :
-
Cucubita maxima
- Cme :
-
Cucumis melo
- CP :
-
Cucurbita pepo
- Cs :
-
Cucumis sativus
- Dc :
-
Dianthus caryophyllus
- DCu :
-
Dendrobium crumenatum
- DK :
-
Diospyros kaki
- Ds :
-
Doritaneopsis
- Fs :
-
Fagus sylvatica
- GM :
-
Glycine max
- LA :
-
Lupinus albus
- LE :
-
Lycopersicon esculentum
- Ls :
-
Lactuca sativa
- MA :
-
Musa acuminata
- Md :
-
Malus domestica
- Mi :
-
Mangifera indica
- Mtru :
-
Medicago truncatula
- NG :
-
Nicotiana glutinosa
- NT :
-
Nicotiana tabacum
- OS :
-
Oryza sativa
- PA :
-
Persea americana
- PC :
-
Pyrus communis
- Pci :
-
Penicillium citrinum
- PE :
-
Passiflora edulis
- PH :
-
Petunia hybrida
- Ph :
-
Phalenopsis sp.
- Phort :
-
Pelargonium hortorum
- Ped :
-
Phyllostachys edulis
- Pmu :
-
Prunus mume
- PP :
-
Prunus persica
- PPa :
-
Prunus armeniaca
- pPP :
-
Pyrus pyrifolia
- POP :
-
Populus euroamericana
- POPe :
-
Populus euphratica
- PS :
-
Pisum sativum
- Pvu :
-
Phaseolus vulgaris
- RP :
-
Rumex palustris
- Sa :
-
Sinapis arvensis
- SH :
-
Striga hermontica
- Sm :
-
Solanum melongena
- SL :
-
Stellaria longipes
- ST :
-
Solanum tuberosum
- TA :
-
Triticum aestivum
- VR :
-
Vigna radiata
References
Asai T., Tena G., Plotnikova J., Willmann M.R., Chiu W.L., Gomez-Gomez L., Boller T., Ausubel F.M., Sheen J. 2002. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature, 415: 977–983.
Capitani G., Hohenester E., Feng L., Storici P., Kirsch J.F., Jansonius J.N. 1999. Structure of 1-aminocyclopropane-1-carboxylate synthase, a key enzyme in the biosynthesis of the plant hormone ethylene. J. Mol. Biol., 294: 745–756.
Destefano-Beltran L.J.C., Caeneghem W.V., Gielen J., Richard L., Van Montagu M., Van Der Straeten D. 1995. Characterization of three members of the ACC synthase gene family in Solanum tuberosum. Mol. Gen. Genet., 246: 496–508.
Dong JG., Kim WT., Yip WK., Thompson GA., Li L., Bennett AB., Yang SF. 1991. Cloning of a cDNA encoding 1-aminocyclopropane-1-carboxylate synthase and expression of its mRNA in ripening apple fruit. Planta, 185: 38–45.
Eliot AC., Kirsch JF. 2002. Modulation of the internal aldimine pK(a)’s of 1-aminocyclopropane-1-carboxylate synthase and aspartate aminotransferase by specific active site residues. Biochemistry, 41: 3836–3842.
Felsenstein J. 1995 and 2001. PHYLIP (Phylogeny Interference package) v.3.57c and v.3.6a2. University of the Washington.
Feng L., Geck M.K., Eliot A.C., Kirsch J.F. 2000. Aminotransferase activity and bioinformatic analysis of 1-aminocyclopropane-1-carboxylate synthase. Biochemistry, 39: 15242–15249.
Galanis A., Yang S.H., Sharrocks A.D. 2001. Selective targeting of MAPKs to the ETS domain transcription factor SAP-1. J. Biol. Chem., 12: 965–973.
Huai Q., Xia Y., Chen Y., Callahan B., Li N., Ke H. 2001. Crystal Structures of 1-aminocyclopropane-1-carboxylate (ACC) synthase in complex with aminoethoxyvinylglycine and pyridoxal-5′-phosphate provide new insight into catalytic mechanisms. J. Biol. Chem., 276: 38210–38216.
Jackson B., Summers J.E., Voesenek L.A.C.J. 1997. Potamogeton pectinatus: a vascular plant that makes no ethylene. Biology and Biotechnology of the Plant Hormone Ethylene. 1997 Kluwer Academic Publishers. NATO ASI Series. Edited by Kanellis A.K., Chang C., Kende H. and Grierson D.
Jansonius J.N. 1998. Structure, evolution and action of vitamin B6-dependent enzymes. Current Opinion in Structural Biology, 8: 759–769.
Jia Y-J., Kakuta Y., Sugawara M., Igarashi T., Oki N., Kisaki M., Shoji T., Kanetuna Y., Horita T., Matsui H., Honma M. 1999. Synthesis and degradation of 1-aminocyclopropane-1-carboxylic acid by Penicillium citrinum. Biosci. Biotech. Biochem., 63: 542–549.
Jonak C., Okresz L., Bogre L., Hirt H. 2002. Complexity cross talk and integration of plant MAP kinase signalling. Current Opinion in Plant Biology, 5: 415–424.
Kakuta Y., Igarashi T., Murakami T., Ito H., Matsui H., Matsui H., Honma M. 2001. 1-Aminocyclopropane-1-carboxylate synthase of Penicillium primary structure and expression in Escherichia coli and S. cerevisiae. Biosci Biotech. Biochem., 65: 1511–1518.
Kende H. 1993. Ethylene biosynthesis. Ann. Rev. Plant Physiol. Plant Mol. Biol., 44: 283–307.
Klintborg A., Eklund L., Little C.H.A. 2002. Ethylene metabolism in Scots pine (Pinus silvestris) shoots during the year. Tree Physiology, 22: 59–66.
Koch K.A., Capitani G., Gruetter M.G., Kirsch J.F. 2001. The human cDNA for a homologue of the plant enzyme 1-aminocyclopropane-1-carboxylate synthase encodes a protein activity. Gene, 272: 75–84.
Li N., Mattoo A. 1994. Deletion of the carboxyl-terminal region of 1-aminocyclopropane-1-carboxylic acid synthase, a key protein in the biosynthesis of ethylene, results in catalitically hyperactive, monomeric enzyme. J. Biol. Chem., 269: 6908–6917.
Li Y., Feng L., Kirsch JF. 1997. Kinetic and spectroscopic investigations of wild-type and mutant forms of apple 1-aminocyclopropane-1-carboxylate synthase. Biochemistry, 36: 15477–15480.
Liang X., Abel S., Keller J.A., Shen N.F., Theologis A. 1992. The 1-aminocyclopropane-1-carboxylate synthase gene family of Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA, 89: 11046–11050.
Lincoln J.E., Campbell A.D., Oetiker J., Rottmann W.H., Oeller P.W., Shen N.F., Theologis A. 1993. LE-ACS4, a fruit ripening and woud-induced 1-aminocyclopropanne-1-carboxylate synthase gene of tomato (Lycopersicon esculentum). J. Biol. Chem., 268: 19422–19430.
McCarthy DL., Capitani G., Feng L., Gruetter MG., Kirsch JF. (2001) Glutamate 47 in 1-aminocyclopropane-1-carboxylate synthase is a major specifity determinant. Biochemistry, 40: 12276–12284.
Matsuoka D., Nanmori T., Sato K., Fukami Y., Kikkawa U., Yasuda T. 2002. Activation of AtMEK1, an Arabidopsis mitogen-activated protein kinase kinase, in vitro and in vivo: analysis of active mutants expressed in E.coli and generation of the active form in stress response in seedlings. Plant J., 29: 637–647.
Mattoo A.K., Baker J.E., and Moline H.E. 1985. Induction by copper ions of ethylene production in Spirodela oligorrhiza: evidence for a pathway independent of 1-aminocyclopropane-1-carboxylic acid. J. Plant Physiol., 123: 193–202.
Osborne D.J., Walters J., Milborrow B.V., Norville A., Stange L.M.C. 1996. Evidence for a non-ACC-dependent ethylene biosynthesis pathway in lower plants. Phytochemistry, 42: 51–60.
Page R.D.M. 1996. TreeView: An application to display phylogenetic trees on personal computers. Computer Applications in the Biological Sciences 12: 357–358.
Peixoto B.R., Mikawa Y., Brenner S. 2001. Characterization of the recombinase activating gene-1 and 2 locus in the Japanese pufferfish, Fugu rubripes. Gene, 246: 275–283.
Ren D., Yang H., Zhang S. 2000. Cell death mediated by MAPKs is associated with hydrogen peroxide production in Arabidopsis. J. Biol. Chem., 277: 559–565.
Rohwer F. and Bopp M. 1984. Ethylene Synthesis in Moss Protonema. J. Plant Physiol., 117: 331–338.
Satoh S., Esashi Y. 1986. Inactivation of 1-aminocyclopropane-1-carboxylic acid synthase of etiolated mung bean hypocotyl segments by its substrate, S-adenosyl-L-methionine. Plant Cell Physiol., 91: 1036–1039.
Satoh S., Mori H., Imaseki H. 1993. Monomeric and dimeric forms and the mechanism-based inactivation of 1-aminocyclopropane-1-carboxylate synthase. Plant Cell Physiol., 34: 753–760.
Savolainen V., Chase M.W., Hoot S.B., Morton C.M., Soltis D.E., Bayer C.F., de Bruijn A.Y., Sullivan S., Qiu Y.L. 2000. Phylogenetic of flowering plants based on combined analysis plastid atp B and rbcL gene sequences. Syst. Biol., 49: 306–362.
Spanu P., Grosskopf D.G., Felix G., Boller T. 1994. The apparent turnover of 1-aminocyclopropane-1-carboxylate synthase in tomato cells is regulated by protein phosphorylation and dephosphorylation. Plant Physiol., 106: 529–535.
Shiu O. Y., Oetiker J.H., Yip W.K., Yang S. F. 1998. The promoter of LE-ACS7, an early flooding-induced 1-aminocyclopropane-1-carboxylate synthase gene of the tomato, is tagged by a Sol3 transposon. Proc. Natl. Acad. Sci USA, 95: 9796–9801.
Tarun A.S., Theologis A. 1998. Complementation analysis of mutants of 1-aminocyclopropane-1-carboxylate synthase reveals the enzyme is a dimer with shared active sites. J. Biol. Chem., 273: 12509–12514.
Tarun A.S., Lee J.S., Theologis A. 1998. Random mutagenesis of 1-aminocyclopropane-1-carboxylate synthase: a key enzyme in ethylene biosynthesis. Proc. Natl. Acad. Sci. USA, 95: 9796–9801.
Thompson J.D., Gibson T.J., Plewniak F., Jeanmougin F., Higgins D.G. 1997. The ClustalX windows interface: flexible strategies for multiple sequences alignment aided by quality analysis tools. Nucleic Acids Res., 24: 4876–4882.
Tittle F.L. 1987. Auxin-stimulated ethylene production in fern gametophytes and sporophytes. Physiol. Plantarum, 70: 499–502.
Vanden Driessche T., Kevers C., Collet M., Gaspar T. 1988. Acetabularia mediterranea and Ethylene: Production in Relation with Development, Circadian Rhythms in Emission, and Response to External Application. J. Plant Physiol., 133: 635–639.
Yang S.F., Hoffman N.E. 1984. Ethylene biosynthesis and its regulation in higher plants. Ann. Rev. Plant Physiol., 35: 155–189.
Zarembinski T. I., Theologis A. 1994. Ethylene biosynthesis and action: case of conservation. Plant Mol. Biol., 26: 1579–1597.
Zhou H., Wang HW., Zhu K., Sui SF., Xu P., Yang SF., Li N. 1999. The multiple roles of conserved arginine 286 of 1-aminocyclopropane-1-carboxylate synthase. Coenzyme binding, substrate binding, and beyond. Plant Physiol., 121: 913–919.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Jakubowicz, M., Pacak, A. Relationships between ACC synthase isozymes calculated using phenetic data. The catalytical and evolutionary aspects. Acta Physiol Plant 26, 5–27 (2004). https://doi.org/10.1007/s11738-004-0040-9
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11738-004-0040-9