Abstract
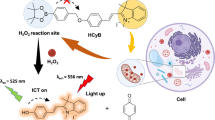
Mitochondrial DNA has a special structure that is prone to damage resulting in many serious diseases, such as genetic diseases and cancers. Therefore, the rapid and specific monitoring of mitochondrial DNA damage is urgently needed for biological recognition. Herein, we constructed an in situ hydrophobic environment-triggering reactive fluorescence probe named MBI-CN. The fluorophore was 2-styrene-1H-benzo[d]imidazole, and malononitrile was introduced as a core into a molecule to initiate the hydrolysis reaction in the specific environment containing damaged mitochondrial DNA. In this design, MBI-CN conjugates to mitochondrial DNA without causing additional damages. Thus, MBI-CN can be hydrolyzed to generate MBI-CHO in an in situ hydrophobic environment with mitochondrial DNA damage. Meanwhile, MBI-CHO immediately emitted a significative fluorescence signal changes at 437 and 553 nm within 25 s for the damaged mitochondria DNA. Give that the specific and rapid response of MBI-CN does not cause additional damages to mitochondrial DNA, it is a potentially effective detection tool for the real-time monitoring of mitochondrial DNA damage during cell apoptosis and initial assessment of cell apoptosis.

Similar content being viewed by others
References
Kuchlyan J, Martinez-Fernandez L, Mori M, Gavvala K, Ciaco S, Boudier C, Richert C, Didier P, Tor Y, Improta R, Mély Y. What makes thienoguanosine an outstanding fluorescent DNA probe. Journal of the American Chemical Society, 2020, 142(40): 16999–17014
Gao F L, Li L J, Fan J L, Cao J F, Li Y Q, Chen L Y, Peng X J. An off-on two-photon carbazole-based fluorescent probe: highly targeting and super-resolution imaging of mtDNA. Analytical Chemistry, 2019, 91(5): 3336–3341
Yakes F M, Van Houten B. Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proceedings of the National Academy of Sciences of the United States of America, 1997, 94 (2): 514–519
Vladimir B S, Evgeny E, Natalia A, Maria A A, Sergey I N, Georgii A B, Igor A, Shamil R S. Error-prone bypass of DNA lesions during lagging-strand replication is a common source of germline and cancer mutations. Nature Genetics, 2019, 51(1): 36–41
Anderson S, Bankier A T, Barrell B G, De Bruijn M H, Coulson A R, Drouin J, Eperon I C, Nierlich D P, Roe B A, Sanger F, Schreier P H, Smith A J H, Staden R, Young I G. Sequence and organization of the human mitochondrial genome. Nature, 1981, 290(5806): 457–465
Bibb M J, Van Etten R A, Wright C T, Walberg M W, Clayton D A. Sequence and gene organization of mouse mitochondrial DNA. Cell, 1981, 26(2): 167–180
Assaf C B, Maayan R, Yifat S O, Michael M I, Dan S, Malka C, Aaron B, Gideon Z, Donna S S, Batsheva K. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell, 2011, 145(3): 435–446
Makoto R H, Jeffrey J K, Erin J W, Sudarshan R, Ryan T S, Wayne G, Aaron J T, Barbara W, Christopher M L, Kunhong X, et al. A stress response pathway regulates DNA damage through β,2-adrenoreceptors and β-arrestin-1. Nature, 2011, 477(7364): 349–353
Furda A M, Marrangoni A M, Lokshin A, Van Houten B. Oxidants and not alkylating agents induce rapid mtDNA loss and mitochondrial dysfunction. DNA Repair, 2012, 11(8): 684–692
Feng B D, Wang K, Liu J W, Mao G J, Cui J Q, Xuan X P, Jiang K, Zhang H. Ultrasensitive apurinic/apyrimidinic site-specific ratio fluorescent rotor for real-time highly selective evaluation of mtDNA oxidative damage in living cells. Analytical Chemistry, 2019, 91 (21): 13962–13969
Neiman M, Taylor D R. The causes of mutation accumulation in mitochondrial genomes. Proceedings. Biological Sciences, 2009, 276(1660): 1201–1209
Furda A M, Marrangoni A M, Lokshin A, Van Houten B. Oxidants and not alkylating agents induce rapid mtDNA loss and mitochondrial dysfunction. DNA Repair, 2012, 11(8): 684–692
Schon E A, DiMauro S, Hirano M. Human mitochondrial DNA: roles of inherited and somatic mutations. Nature Reviews. Genetics, 2012, 13(12): 878–890
Schuermann D, Scheidegger S P, Weber A R, Bjoras M, Leumann C J, Schar P. 3CAPS—a structural AP-site analogue as a tool to investigate DNA base excision repair. Nucleic Acids Research, 2016, 44(5): 2187–2198
Bender A, Krishnan K J, Morris C M, Taylor G A, Reeve A K, Perry R H, Jaros E, Hersheson J S, Betts J, Klopstock T, Taylor R W, Turnbull D M. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and parkinson disease. Nature Genetics, 2006, 38(5): 515–517
Rebbeck C A, Leroi A M, Burt A. Mitochondrial capture by a transmissible cancer. Science, 2011, 331(6015): 303
Ascaso F J, Lopez-Gallardo E, DelPrado E, Ruiz-Pesini E, Montoya J. Macular lesion resembling adult-onset vitelliform macular dystrophy in Kearns-Sayre syndrome with multiple mtDNA deletions. Clinical & Experimental Ophthalmology, 2010, 38(8): 812–816
Badura-Stronka M, Wawrocka A, Zawieja K, Silska S, Krawczynski M R. Severe manifestation of Leber’s hereditary optic neuropathy due to 11778G > A mtDNA mutation in a female with hypoestrogenism due to perrault syndrome. Mitochondrion, 2013, 13(6): 831–834
Sun W, Li M, Fan J L, Peng X J. Activity-based sensing and theranostic probes based on photoinduced electron transfer. Accounts of Chemical Research, 2019, 52(10): 2818–2831
Sun W, Guo S G, Hu C, Fan J L, Peng X J. Recent development of chemosensors based on cyanine platforms. Chemical Reviews, 2016, 116(14): 7768–7817
Lin F, Zhou Y F, Li Q S, Zhou X S, Shao Y, Haber-meyer B, Wang H, Shi X H, Xu Z A. Prototropically allosteric probe for superbly selective DNA analysis. Analytical Chemistry, 2017, 89(17): 9299–9306
Zou X X, Shi Y L, Zhu R, Han J H, Han S F. Organelle-redirected chameleon sensor-enabled live cell imaging of mitochondrial DNA. Analytical Chemistry, 2019, 91(24): 15899–15907
Abeywickrama C S, Bertman K A, Plescia C B, Stahelin R V, Pang Y. Structural effect on the cellular selectivity of an NIR-emitting cyanine probe: from lysosome to simultaneous nucleus and mitochondria selectivity with potential for monitoring mitochondria dysfunction in cells. ACS Applied Bio Materials, 2019, 2(11): 5174–5181
Briggs C, Jones M. SYBR Green I-induced fluorescence in cultured immune cells: a comparison with acridine orange. Acta Histochemica, 2005, 107(4): 301–312
Davis S K, Bardeen C J. Cross-linking of histone proteins to DNA by UV illumination of chromatin stained with hoechst 33342. Photochemistry and Photobiology, 2003, 77(6): 675–679
Pfeifer G P, You Y H, Besaratinia A. Mutations induced by ultraviolet light. Mutation Research, 2005, 571(1–2): 19–31
Smith P J, Blunt N, Wiltshire M, Hoy T, Teesdale-Spittle P, Craven M R, Watson J V, Amos W B, Errington R J, Patterson L H. Characteristics of a novel deep red/infrared fluorescent cell-permeant DNA probe, DRAQ5, in intact human cells analyzed by flow cytometry, confocal and multiphoton microscopy. Cytometry, 2000, 40(4): 280–291
Burke C S, Byrne A, Keyes T E. Highly selective mitochondrial targeting by a ruthenium(II) peptide conjugate: imaging and photoinduced damage of mitochondrial DNA. Angewandte Chemie International Edition, 2018, 57(38): 12420–12424
Cao J J, Zheng Y, Wu X W, Tan C P, Chen M H, Wu N, Ji L N, Mao Z W. Anticancer cyclometalated iridium(III) complexes with planar ligands: mitochondrial DNA damage and metabolism disturbance. Journal of Medicinal Chemistry, 2019, 62(7): 3311–3322
Zheng Y, Zhang D Y, Zhang H, Cao J J, Tan C P, Ji L N, Mao Z W. Photodamaging of mitochondrial DNA to overcome cisplatin resistance by a Ru II-Pt II bimetallic complex. Chemistry (Weinheim an der Bergstrasse, Germany), 2018, 24(71): 18971–18980
Feng B D, Wang K, Yang Y G, Wang G, Zhang H, Liu Y F, Jiang K. Ultrasensitive recognition of AP sites in DNA at the single-cell level: one molecular rotor sequentially self-regulated to form multiple different stable conformations. Chemical Science (Cambridge), 2019, 10(44): 10373–10380
Peng X J, Wu T, Fan J L, Wang J Y, Zhang S, Song F L, Sun S G. An effective minor groove binder as a red fluorescent marker for live-cell DNA imaging and quantification. Angewandte Chemie International Edition, 2011, 50(18): 4180–4183
Friedberg E C. DNA damage and repair. Nature, 2003, 421(6921): 436–440
Song Y L, Tian T, Shi Y Z, Liu W L, Zou Y, Khajvand T, Wang S L, Zhu Z, Yang C Y. Enrichment and single-cell analysis of circulating tumor cells. Chemical Science (Cambridge), 2017, 8(3): 1736–1751
Leung C H, Zhong H J, He H Z, Lu L, Chan D S H, Ma D L. Luminescent oligonucleotide-based detection of enzymes involved with DNA repair. Chemical Science (Cambridge), 2013, 4(10): 3781–3795
Liu C X, Wang Y F, Zhang X, Wu F, Yang W, Zou G G, Yao Q, Wang J Q, Chen Y Q, Wang S R, Zhou X. Enrichment and fluorogenic labelling of 5-formyluracil in DNA. Chemical Science (Cambridge), 2017, 8(6): 4505–4510
Visvardis E E, Tassiou A M, Piperakis S M. Study of DNA damage induction and repair capacity of fresh and cryopreserved lymphocytes exposed to H2O2 and γ-irradiation with the alkaline comet assay. Mutation Research, 1997, 383(1): 71–80
Berova N, Nakanishi K, Woody R W. Circular Dichroism: Principles and Applications. New York: Wiley-VCH, 2000, 703–718
Monnot M, Mauffret O, Lescot E, Fermandjian S. Probing intercalation and conformational effects of the anticancer drug 2-methyl-9-hydroxyellipticinium acetate in DNA fragments with circular dichroism. European Journal of Biochemistry, 1992, 204 (3): 1035–1039
Karami K, Shirani-Sarmazeh Z, Hosseini-Kharat M, Lipkowski J, Saeidifar M. Synthesis, spectral characterization, crystal structure and in vitro DNA/protein binding studies of phosphorous ylide palladacyclic complexes containing azide group. Journal of Photochemistry and Photobiology. B, Biology, 2015, 144: 11–19
Baase W A, Johnson W C Jr. Circular dichroism and DNA secondary structure. Nucleic Acids Research, 1979, 6(2): 797–814
Sarkar D, Das P, Basak S, Chattopadhyay N. Binding interaction of cationic phenazinium dyes with calf thymus DNA: a comparative study. Journal of Physical Chemistry B, 2008, 112(30): 9243–9249
Poulsen B C, Estalayo-Adrián S, Blasco S, Bright S A, Kelly J M, Williams D C, Gunnlaugsson T. Luminescent ruthenium polypyridyl complexes with extended ‘dppz’ like ligands as DNA targeting binders and cellular agents. Dalton Transactions (Cambridge, England), 2016, 45(45): 18208–18220
Fishel M L, Seo Y R, Smith M L, Kelley M R. Imbalancing the DNA base excision repair pathway in the mitochondria; targeting and overexpressing N-methylpurine DNA glyeosylase in mitochondria leads to enhanced cell killing. Cancer Research, 2003, 63(3): 608–615
Scaduto R C Jr, Grotyohann L W. Measurement of mitochondrial membrane potential using fluorescent rhodamine derivatives. Biophysical Journal, 1999, 76(1): 469–477
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 21722501 and 22004028), Henan Special Support for High-level Talents Central Plains Science and Technology Innovation Leading Talents (Grant No. 204200510006), Key Project of Science and Technology of Henan Province (Grant No. 202102310139).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Feng, B., Niu, H., Zhai, H. et al. In-situ hydrophobic environment triggering reactive fluorescence probe to real-time monitor mitochondrial DNA damage. Front. Chem. Sci. Eng. 16, 92–102 (2022). https://doi.org/10.1007/s11705-021-2063-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11705-021-2063-9