Abstract
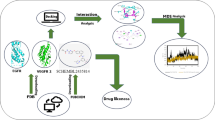
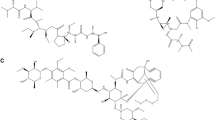
The coumarin nucleus is a simple privileged scaffold distributed in many plants. It has recently gained attention for its diverse biological activities and interactions with enzymes and receptors. The vascular endothelial growth factor receptor-2 (VEGFR-2), a receptor tyrosine kinase, is a crucial cancer target as it is involved in angiogenesis. This study employs virtual screening, molecular docking, and molecular simulation studies to identify potential coumarin candidates against VEGFR-2 from the COCONUT database. After thorough docking studies, CNP0056360, CNP0340213, and CNP0366287 were identified as final hits. Molecular dynamics simulation studies revealed strong stability and better binding energies for CNP0056360 and CNP0340213, outperforming lenvatinib; CNP0366287 showed comparable behaviour. The identified coumarins exhibited good in-silico pharmacokinetics and demonstrated low toxicity.
Graphical Abstract










Similar content being viewed by others
Data availability
The files associated with the research could be freely assessed on “https://zenodo.org/records/10646050”.
References
Abolibda TZ, Fathalla M, Farag B, Zaki MEA, Gomha SM (2023) Synthesis and molecular docking of some novel 3-Thiazolyl-coumarins as inhibitors of VEGFR-2 Kinase. Molecules [Online], 28
Alshabanah LA, Al-Mutabagani LA, Gomha SM, Ahmed HA (2022) Three-component synthesis of some new coumarin derivatives as anticancer agents. Front Chem 9:762248
Al-Warhi T, Sabt A, Elkaeed EB, Eldehna WM (2020) Recent advancements of coumarin-based anticancer agents: an up-to-date review. Bioorg Chem 103:104163
Annunziata F, Pinna C, Dallavalle S, Tamborini L, Pinto A (2020) An overview of coumarin as a versatile and readily accessible scaffold with broad-ranging biological activities. Int J Mol Sci [Online] 21
Aziz MA, Serya RAT, Lasheen DS, Abdel-Aziz AK, Esmat A, Mansour AM, Singab ANB, Abouzid KAM (2016) Discovery of potent VEGFR-2 inhibitors based on furopyrimidine and thienopyrimidne scaffolds as cancer targeting agents. Sci Rep 6:24460
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucl Acids Res 28:235–242
Berthold MR, Cebron N, Dill F, Gabriel TR, Kötter T, Meinl T, Ohl P, Sieb C, Thiel K, Wiswedel B (2008) KNIME: the Konstanz information miner. In: Data analysis, machine learning and applications. Springer, Berlin pp 319–326
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717
Davis IW, Leaver-Fay A, Chen VB, Block JN, Kapral GJ, Wang X, Murray LW, Arendall WB III, Snoeyink J, Richardson JS, Richardson DC (2007) MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucl Acids Res 35:W375–W383
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: an automated pipeline for the setup of Poisson–Boltzmann electrostatics calculations. Nucl Acids Res 32:W665–W667
Durrant JD, McCammon JA (2011) Molecular dynamics simulations and drug discovery. BMC Biol 9:71
Ganeshpurkar A, Singh R, Gore PG, Kumar D, Gutti G, Kumar A, Singh SK (2020) Structure-based screening and molecular dynamics simulation studies for the identification of potential acetylcholinesterase inhibitors. Mol Simul 46:169–185
Ghosh P, Singh R, Ganeshpurkar A, Swetha R, Kumar D, Singh SK, Kumar A (2022) Identification of potential death-associated protein kinase-1 (DAPK1) inhibitors by an integrated ligand-based and structure-based computational drug design approach. J Biomol Struct Dyn 1–13
Gomha SM, Abdel-Aziz HM, El-Reedy AAM (2018) Facile synthesis of Pyrazolo[3,4-c]pyrazoles bearing coumarine ring as anticancer agents. J Heterocycl Chem 55:1960–1965
Hao Z, Wang P (2020) Lenvatinib in management of solid tumors. Oncologist 25:e302–e310
Hata H, Phuoc Tran D, MarzoukSobeh M, Kitao A (2021) Binding free energy of protein/ligand complexes calculated using dissociation parallel cascade selection molecular dynamics and Markov state model. Biophys Physicobiol 18:305–316
Hicklin DJ, Ellis LM (2005) Role of the vascular endothelial growth factor pathway in tumor growth and angiogenesis. J Clin Oncol 23:1011–1027
Jabeen A, Mohamedali A, Ranganathan S (2019) Protocol for Protein structure modelling. Encyclopedia of bioinformatics and computational biology. Academic Press, Oxford
Jiang Z, You L, Dou W, Sun T, Xu P (2019) Effects of an electric field on the conformational transition of the protein: a molecular dynamics simulation study. Polymers [Online] 11
Jitendra S, Vinay R (2011) Structure based drug designing of a novel antiflaviviral inhibitor for nonstructural 3 protein. Bioinformation 6:57–60
Kumar R, Sharma A (2023) Chapter 15—Computational strategies and tools for protein tertiary structure prediction. Basic biotechniques for bioprocess and bioentrepreneurship. Academic Press, Cambridge
Küpeli Akkol E, Genç Y, Karpuz B, Sobarzo-Sánchez E, Capasso R (2020) Coumarins and coumarin-related compounds in pharmacotherapy of cancer. Cancers [Online] 12
Lugano R, Ramachandran M, Dimberg A (2020) Tumor angiogenesis: causes, consequences, challenges and opportunities. Cell Mol Life Sci 77:1745–1770
Ma X-L, Chen C, Yang J (2005) Predictive model of blood-brain barrier penetration of organic compounds. Acta Pharmacol Sin 26:500–512
Maria João M, Lourdes S, Eugenio U, Orlando AA, Enrique M, Estela Guardado Y (2015) Coumarins—an important class of phytochemicals. Phytochemicals. IntechOpen
Meng XY, Zhang HX, Mezei M, Cui M (2011) Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des 7:146–157
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
NaderiAlizadeh M, Rashidi M, Muhammadnejad A, MoeiniZanjani T, Ziai SA (2018) Antitumor effects of umbelliprenin in a mouse model of colorectal cancer. Iran J Pharm Res 17:976–985
Nilsson M, Heymach JV (2006) Vascular endothelial growth factor (VEGF) pathway. J Thorac Oncol 1:768–770
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open babel: an open chemical toolbox. J Cheminformatics 3:33
Önder A (2020) Chapter 3—Anticancer activity of natural coumarins for biological targets. In: Atta Ur R (ed) Studies in natural products chemistry. Elsevier, Amsterdam
Peng F-W, Liu D-K, Zhang Q-W, Xu Y-G, Shi L (2017) VEGFR-2 inhibitors and the therapeutic applications thereof: a patent review (2012–2016). Expert Opin Ther Pat 27:987–1004
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—A visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Pinedo HM, Slamon DJ (2000) INTRODUCTION: Translational Research: The Role of VEGF in Tumor Angiogenesis. Oncologist 5:1–2
S V, Kajal K, Mondal S, Wahan SK, Das Kurmi B, Das Gupta G, Patel P (2023) Novel VEGFR-2 kinase inhibitors as anticancer agents: a review focusing on SAR and molecular docking studies (2016–2021). Chem Biodivers 20:e202200847
Sharifi-Rad J, Cruz-Martins N, López-Jornet P, Lopez EP-F, Harun N, Yeskaliyeva B, Beyatli A, Sytar O, Shaheen S, Sharopov F, Taheri Y, Docea AO, Calina D, Cho WC (2021) Natural coumarins: exploring the pharmacological complexity and underlying molecular mechanisms. Oxid Med Cell Longev 2021:6492346
Singh R, Pokle AV, Ghosh P, Ganeshpurkar A, Swetha R, Singh SK, Kumar A (2022) Pharmacophore-based virtual screening, molecular docking and molecular dynamics simulations study for the identification of LIM kinase-1 inhibitors. J Biomol Struct Dyn 41:1–15
Skariyachan S, Garka S (2018) Chapter 1—Exploring the binding potential of carbon nanotubes and fullerene towards major drug targets of multidrug resistant bacterial pathogens and their utility as novel therapeutic agents. In: Grumezescu AM (ed) Fullerens graphenes and nanotubes. William Andrew Publishing
Sneha P, George Priya Doss C (2016) Chapter seven—Molecular dynamics: new frontier in personalized medicine. In: Donev R (ed) Advances in protein chemistry and structural biology. Academic Press
Sobolev OV, Afonine PV, Moriarty NW, Hekkelman ML, Joosten RP, Perrakis A, Adams PD (2020) A global Ramachandran score identifies protein structures with unlikely stereochemistry. Structure 28:1249-1258.e2
Sorokina M, Merseburger P, Rajan K, Yirik MA, Steinbeck C (2021) COCONUT online: collection of open natural products database. J Cheminformatics 13:1–13
Subramaniam S, Mehrotra M, Gupta D (2008) Virtual high throughput screening (vHTS)—a perspective. Bioinformation 3:14–17
Sunseri J, Koes DR (2016) Pharmit: interactive exploration of chemical space. Nucl Acids Res 44:W442–W448
Swetha R, Sharma A, Singh R, Ganeshpurkar A, Kumar D, Kumar A, Singh SK (2022) Combined ligand-based and structure-based design of PDE 9A inhibitors against Alzheimer’s disease. Mol Divers 26:2877–2892
Tripathi N, Goel B, Bhardwaj N, Sahu B, Kumar H, Jain SK (2022) Virtual screening and molecular simulation study of natural products database for lead identification of novel coronavirus main protease inhibitors. J Biomol Struct Dyn 40:3655–3667
Wang X, Bove AM, Simone G, Ma B (2020) Molecular Bases of VEGFR-2-Mediated Physiological Function and Pathological Role. Front Cell Dev Biol 8:599281
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, De Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucl Acids Res 46:W296–W303
Williams CJ, Headd JJ, Moriarty NW, Prisant MG, Videau LL, Deis LN, Verma V, Keedy DA, Hintze BJ, Chen VB, Jain S, Lewis SM, Arendall WB III, Snoeyink J, Adams PD, Lovell SC, Richardson JS, Richardson DC (2018) MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci 27:293–315
Acknowledgements
The authors extend their gratitude toward Professor David A. Case, Department of Chemistry & Chemical Biology, Rutgers University, New Jersey, USA, for granting a licence for Amber 20. NT and NB are also thankful to IIT (BHU) and Ministry of Education (MoE), New Delhi for providing teaching assistantship.
Funding
This work was partially supported by the SERB-CRG Grant (Grant No. CRG/2022/001637).
Author information
Authors and Affiliations
Contributions
Nancy Tripathi: Conceived, designed, and performed the experiments; Analyzed the data; Writing original draft. Nivedita Bhardwaj: Prepared figures and tables; Paper formatting. Bikarma Singh: Conceived and designed the experiments. Shreyans K. Jain: Conceived and designed the experiments; Supervision; Final paper review and formatting.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tripathi, N., Bhardwaj, N., Singh, B. et al. In-silico identification of Coumarin-based natural compounds as potential VEGFR-2 inhibitors. Chem. Pap. 78, 4347–4360 (2024). https://doi.org/10.1007/s11696-024-03395-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-024-03395-5