Abstract
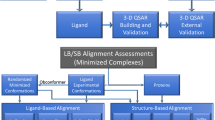
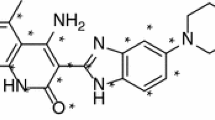
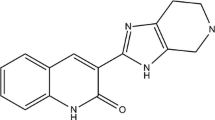
The vascular endothelial growth factor (VEGF) is the main target of tumor treatment. VEGFR-2 is the main functional receptor of VEGF, which is involved in the regulation of angiogenesis. Based on hologram quantitative structure activity relationships (HQSAR) and topomer comparative molecular field analysis (topomer CoMFA), the contribution of 6-amide-2-aryl benzoxazole/benzimidazole derivatives (VEGFR-2 kinase inhibitors) to these structures was discussed and the corresponding modification strategies were proposed. The most effective HQSAR and topomer CoMFA models are generated by using a training set of 33 compounds. In order to ensure the robustness of the model, the randomization test was used, and 11 compounds were selected as the test set. The results show that the q2 of cross-validation is 0.646/0.659, and the r2 of non-cross-validation is 0.871/0.867, respectively. The data show that both models are reliable. Topomer CoMFA’s steric/electrostatic contour and HQSAR’s atomic contribution map show the structural characteristics controlling its inhibition ability. In addition, molecular docking is also used to study the interaction between these drugs and large proteins, and the ligand pair is connected to the active site of VEGFR-2 kinase, revealing the possible biological active conformation. This study showed that there was a wide interaction between 6-amide-2-aryl benzoxazole/benzimidazole derivatives and Hrg136 and Tyr356 residues of VEGFR-2 kinase active site. Finally, we used ADMET properties and drug-like properties to predict the newly designed molecules, and the results showed that they meet the conditions for becoming drugs and are expected to become potential anti-VEGFR-2 inhibitors. This study can provide a theoretical reference for the synthesis of target products of VEGFR-2 inhibitors.






Similar content being viewed by others
References
Aarjane M, Aouidate A, Slassi S, Amine A (2020) Synthesis, antibacterial evaluation, in silico ADMET and molecular docking studies of new N-acylhydrazone derivatives from acridone. Arab J Chem 13(7):6236–6245
Agahi F, Juan C, Font G, Juan-García A (2020) In silico methods for metabolomic and toxicity prediction of zearalenone, α-zearalenone and β-zearalenone. J Food Chem Tox 146:111818
Ancuceanu R, Tamba B, Stoicescu CS (2020) Use of qsar global models and molecular docking for developing new inhibitors of c-src tyrosine kinase. Internation J Mole Sci 21(1):199–215
Bray F, Mller B (2006) Predicting the future burden of cancer. Nat Rev Cancer 6(1):63–74
Bray F, Jemal A, Grey N, Ferlay J, Forman D (2012) Global cancer transitions according to the Human Development Index (2008–2030): a population-based study. Lancet Oncol 13(8):790–801
Cramer RD (2012) R-group template CoMFA combines benefits of “ad hoc” and topomer alignments using 3D-QSAR for lead optimization. J Comput Aid Mol Des 26(7):805–819
Doddareddy MR, Lee YJ, Yong SC, Choi KI, Koh HY, Pae AN (2004) Hologram quantitative structure activity relationship studies on 5-HT6 antagonists. Bioorgan Med Chem 12(14):3815–3824
Farhood B, Raei B, Malekzadeh R (2019) A review of incidence and mortality of colorectal, lung, liver, thyroid, and bladder cancers in Iran and compared to other countries. Contemp Onc Wsp Onk 23(1):7–15
Ferlay J, Shin H-R, Bray F, Forman D, Parkin DM (2010) Estimates of worldwide burden of cancer in 2008: GLOBOCAN. Int J Cancer 127(12):2893–2917
Ferrara N, Kerbel RS (2005) Angiogenesis as a therapeutic target. Nature 438(7070):967–974
Gersten O, Wilmoth JR (2002) The cancer transition in Japan since 1951. Demogra Res 7(5):271–306
Golbraikh A, Tropsha A (2002) Beware of q2! J Mol Graph Model 20(4):269–276
Jain AN (2007) Surflex Dock 2.1: robust performance from ligand energetic modeling ring flexibility and knowledge-based search. J Comput Aid Mol Des 21(5):281–306
Ajay JN (2003) Surflex:? fully automatic flexible molecular docking using a molecular similarity-based search engine. J Med Chem. 46(4):499–511
Jemal A, Bray F, Center MM, Ferlay JJ, Forman D (2011) Global cancer statistics. J Clin 61(2):69–90
Kallab T, Schfer B, Viveiros A (2020) Penetrance, cancer incidence and survival of hemochromatosis in a long-term follow-up and epidemiological modeling study. Z Gastroenterol 58(05):144–156
Kumar S, Tiwari M (2015) Topomer-CoMFA-based predictive modelling on 2,3-diaryl-substituted-1,3-thiazolidin-4-ones as non-nucleoside reverse transcriptase inhibitors. Med Chem Res 24(1):245–257
Lee HS, Lee NCO, Kouprina N (2016) Effects of anticancer drugs on chromosome instability and new clinical implications for tumor-suppressing therapies. Cancer Resear 76(4):902–911
Lee WS, Yang H, Chong HJ (2020) Combination of anti-angiogenic therapy and immune checkpoint blockade normalizes vascular-immune crosstalk to potentiate cancer immunity. Exper Mol Med 52(9):1475–1485
Leeson PD, Oprea TI (2011) Chapter 2:Drug-Like Physicochemical Properties. Quant Approach, Drug Des Strat, pp 35–59
Li C, Wang P, Zhen Z (2017) 3D-QSAR, Topomer CoMFA, docking analysis, and ADMET prediction of thioether pleuromutilin derivatives as antibacterial agents. Lett Drug Des Discov 14(8):869–879
Li X, Kleinstreuer NC, Fourches D (2020) Hierarchical quantitative structure-activity relationship modeling approach for integrating binary, multiclass, and regression models of acute oral systemic toxicity. Chem Resear Toxic 33(2):353–366
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings1. Advan Drug Del Rev 64:4–17
Papetti M, Herman IM (2002) Mechanisms of normal and tumor-derived angiogenesis. Am J Physiol-Cell Physiol 282(5):C947–C970
Purcell WP, Singer JA (1967) A brief review and table of semiempirical parameters used in the Hueckel molecular orbital method. J Chem Eng Data 12(2):235–246
Roskoski R (2007) Vascular endothelial growth factor (VEGF) signaling in tumor progression. Crit Rev Oncol Hemat 62(3):179–213
Roy K, Das RN, Ambure P, Aher RB (2016) Be aware of error measures. Further studies on validation of predictive QSAR models. Chemometr Intell Lab 152:18–33
Seal A, Aykkal R, Babu RO, Ghosh MG (2011) Docking study of HIV-1 reverse transcriptase with phytochemicals. Bioinformation 5(10):430–439
Shibuya M (2011) Vascular endothelial growth factor (VEGF) and its receptor (VEGFR) signaling in angiogenesis: a crucial target for anti- and pro-angiogenic therapies. Genes Cancer 2(12):1097–1105
Tong JB, Bai M, Zhao X (2016) 3D-QSAR and docking studies of HIV-1 protease inhibitors using R-group search and Surflex-dock. Med Chem Res 25(11):2619–2630
Tong JB, Wang Y, Lei S (2019) Comprehensive 3D-QSAR and binding mode of dapy inhibitors using r-group search and molecular docking. J Struct Chem 38(1):25–36
Ugarkar AG, Ambre PK, Coutinho EC, Nandan S, Pissurlenkar RRS (2014) Extracting structural requirements for activity of GPR119 agonists: a hologram quantitative structure activity relationship (HQSAR) study. Can J Chem 92(7):670–676
Waller CL (2004) A comparative qsar study using CoMFA, HQSAR, and FRED/SKEYS paradigms for estrogen receptor binding affinities of structurally diverse compounds. J Chem Inf Model 44(2):758–765
Wang Y, Yu S, Ruan X, Yuan J, Shi J (2015) HQSAR and topomer CoMFA for predicting melanocortin-4 receptor binding affinities of trans-4-(4-chlorophenyl) pyrrolidine-3-carboxamides. Chemometr Intell Lab 146:34–41
Wasko MJ, Pellegrene KA, Madura JD (2015) A role for fragment-based drug design in developing novel lead compounds for central nervous system targets. Front Neur 6:197
Tong W, David Lowis R, Perkins R, Chen Yu, William J (1998) Evaluation of quantitative structure−activity relationship methods for large-scale prediction of chemicals binding to the estrogen receptor. J Chem Inf Model 38(4):669–677
Yadav DK, Khan F, Negi AS (2012) Pharmacophore modeling, molecular docking, QSAR, and in silico ADMET studies of gallic acid derivatives for immunomodulatory activity. J Mol Mod 18(6):2513–2525
Yang HB, Luo CF, Sun LX, Li J, Cai YC, Wang Z, Li WH, Liu GX, Tang Y (2019) admetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics 35(6):1067–1069
Yuan X, Yang Q, Liu T, Li K, Liu Y, Zhu C, Zhang Z, Li L, Zhang C, Xie M, Lin J, Zhang J, Jin Y (2019) Design, synthesis and in vitro evaluation of 6-amide-2-aryl benzoxazole/benzimidazole derivatives against tumor cells by inhibiting VEGFR-2 kinase. Eur J Med Chem 179:147–165
Zhang Z, Song J, Xiang Y (2014) Topomer CoMFA and virtual screening studies of azaindole class renin inhibitors. Comb Chem High Throughtput Screen 17(5):458–472
Zhong Z, Yu J, Virshup DM (2020) Wnts and the hallmarks of cancer. Cancer Metasta Rev 39:625–645
Acknowledgements
This work was supported by the National Natural Science Funds of China (21475081), the Natural Science Foundation of Shaanxi Province of China (2019JM-237), and the Graduate Innovation Fund of Shaanxi University of Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tong, JB., Feng, Y., Luo, D. et al. 6-amide-2-aryl benzoxazole/benzimidazole derivatives as VEFGR-2 inhibitors in two-and three-dimensional QSAR studies: topomer CoMFA and HQSAR. Chem. Pap. 75, 3551–3562 (2021). https://doi.org/10.1007/s11696-021-01588-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-021-01588-w