Abstract
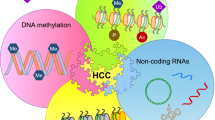
Hepatocellular carcinoma (HCC) development is characterized by the presence of epigenetic alterations, including promoter DNA hypermethylation and post-translational modifications of histone, which profoundly affect expression of a wide repertoire of genes critical for cancer development. Emerging data suggest that deregulation of polycomb group (PcG) proteins, which are key chromatin modifiers repressing gene transcription during developmental stage, plays a causative role in oncogenesis. PcG proteins assemble into polycomb repressive complex 1 (PRC1) and polycomb repressive complex 2 (PRC2) to impose the histone H3 lysine 27 trimethylation (H3K27me3) modification for repression. In this review, we will first recapitulate the mechanisms of two key epigenetic pathways: DNA methylation and histone modifications. Specifically, we will focus our discussion on the molecular roles of PcG proteins. Next, we will highlight recent findings on PcG proteins, their clinicopathological implication and their downstream molecular consequence in hepatocarcinogenesis. Last but not least, we will consider the therapeutic potential of targeting enhancer of zeste homolog 2 (EZH2) as a possible treatment for HCC. Improving our understanding on the roles of PcG proteins in hepatocarcinogenesis can benefit the development of epigenetic-based therapy.
Similar content being viewed by others
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin 2011;61(2):69–90
Ferlay Ja. GLOBOCAN 2008, Cancer Incidence and Mortality Worldwide: IARC CancerBase. 2008; Available from: http://globocan.iarc.fr/(Accessed on October 1, 2012)
Tang ZY. Hepatocellular carcinoma-cause, treatment and metastasis. World J Gastroenterol 2001;7(4):445–454
Jones PA, Baylin SB. The epigenomics of cancer. Cell 2007;128(4):683–692
Jones PA, Laird PW. Cancer epigenetics comes of age. Nat Genet 1999;21(2):163–167
Riggs AD. X inactivation, differentiation, and DNA methylation. Cytogenet Cell Genet 1975;14(1):9–25
Holliday R, Pugh JE. DNA modification mechanisms and gene activity during development. Science 1975;187(4173):226–232
Kondoh N, Wakatsuki T, Hada A, Shuda M, Tanaka K, Arai M, Yamamoto M. Genetic and epigenetic events in human hepatocarcinogenesis. Int J Oncol 2001;18(6):1271–1278
Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002;3(6):415–428
Turker MS. The establishment and maintenance of DNA methylation patterns in mouse somatic cells. Semin Cancer Biol 1999;9(5):329–337
Jones PL, Veenstra GJ, Wade PA, Vermaak D, Kass SU, Landsberger N, Strouboulis J, Wolffe AP. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat Genet 1998;19(2):187–191
Wong CM, Ng YL, Lee JM, Wong CC, Cheung OF, Chan CY, Tung EK, Ching YP, Ng IO. Tissue factor pathway inhibitor-2 as a frequently silenced tumor suppressor gene in hepatocellular carcinoma. Hepatology 2007;45(5):1129–1138
Tung EK, Wong CM, Yau TO, Lee JM, Ching YP, Ng IO. HAI-2 is epigenetically downregulated in human hepatocellular carcinoma, and its Kunitz domain type 1 is critical for anti-invasive functions. Int J Cancer 2009;124(8):1811–1819
Gama-Sosa MA, Slagel VA, Trewyn RW, Oxenhandler R, Kuo KC, Gehrke CW, Ehrlich M. The 5-methylcytosine content of DNA from human tumors. Nucleic Acids Res 1983;11(19):6883–6894
Lister R, Ecker JR. Finding the fifth base: genome-wide sequencing of cytosine methylation. Genome Res 2009;19(6):959–966
Kanai Y. Genome-wide DNA methylation profiles in precancerous conditions and cancers. Cancer Sci 2010;101(1):36–45
Ehrlich M. DNA methylation in cancer: too much, but also too little. Oncogene 2002;21(35):5400–5413
Saito Y, Kanai Y, Sakamoto M, Saito H, Ishii H, Hirohashi S. Expression of mRNA for DNA methyltransferases and methyl-CpG-binding proteins and DNA methylation status on CpG islands and pericentromeric satellite regions during human hepatocarcinogenesis. Hepatology 2001;33(3):561–568
Kornberg RD, Lorch Y. Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell 1999;98(3):285–294
Böhm L, Crane-Robinson C. Proteases as structural probes for chromatin: the domain structure of histones. Biosci Rep 1984;4(5):365–386
Strahl BD, Allis CD. The language of covalent histone modifications. Nature 2000;403(6765):41–45
Jenuwein T, Allis CD. Translating the histone code. Science 2001;293(5532):1074–1080
Lewis EB. A gene complex controlling segmentation in Drosophila. Nature 1978;276(5688):565–570
Levine SS, King IF, Kingston RE. Division of labor in polycomb group repression. Trends Biochem Sci 2004;29(9):478–485
Otte AP, Kwaks TH. Gene repression by Polycomb group protein complexes: a distinct complex for every occasion? Curr Opin Genet Dev 2003;13(5):448–454
Cao R, Wang L, Wang H, Xia L, Erdjument-Bromage H, Tempst P, Jones RS, Zhang Y. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002;298(5595):1039–1043
Sing A, Pannell D, Karaiskakis A, Sturgeon K, Djabali M, Ellis J, Lipshitz HD, Cordes SP. A vertebrate Polycomb response element governs segmentation of the posterior hindbrain. Cell 2009;138(5):885–897
Plath K, Fang J, Mlynarczyk-Evans SK, Cao R, Worringer KA, Wang H, de la Cruz CC, Otte AP, Panning B, Zhang Y. Role of histone H3 lysine 27 methylation in X inactivation. Science 2003;300(5616):131–135
Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell 2008;32(2):232–246
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007;129(7):1311–1323
Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM, Chevalier B, Johnstone SE, Cole MF, Isono K, Koseki H, Fuchikami T, Abe K, Murray HL, Zucker JP, Yuan B, Bell GW, Herbolsheimer E, Hannett NM, Sun K, Odom DT, Otte AP, Volkert TL, Bartel DP, Melton DA, Gifford DK, Jaenisch R, Young RA. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 2006;125(2):301–313
Bracken AP, Dietrich N, Pasini D, Hansen KH, Helin K. Genomewide mapping of Polycomb target genes unravels their roles in cell fate transitions. Genes Dev 2006;20(9):1123–1136
Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI, Levine SS, Wernig M, Tajonar A, Ray MK, Bell GW, Otte AP, Vidal M, Gifford DK, Young RA, Jaenisch R. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 2006;441(7091):349–353
Juan AH, Kumar RM, Marx JG, Young RA, Sartorelli V. Mir-214-dependent regulation of the polycomb protein Ezh2 in skeletal muscle and embryonic stem cells. Mol Cell 2009;36(1):61–74
Montgomery ND, Yee D, Chen A, Kalantry S, Chamberlain SJ, Otte AP, Magnuson T. The murine polycomb group protein Eed is required for global histone H3 lysine-27 methylation. Curr Biol 2005 24;15(10): 942–947
Pasini D, Bracken AP, Jensen MR, Lazzerini Denchi E, Helin K. Suz12 is essential for mouse development and for EZH2 histone methyltransferase activity. EMBO J 2004;23(20):4061–4071
Cao R, Zhang Y. SUZ12 is required for both the histone methyltransferase activity and the silencing function of the EEDEZH2 complex. Mol Cell 2004;15(1):57–67
Cai MY, Tong ZT, Zheng F, Liao YJ, Wang Y, Rao HL, Chen YC, Wu QL, Liu YH, Guan XY, Lin MC, Zeng YX, Kung HF, Xie D. EZH2 protein: a promising immunomarker for the detection of hepatocellular carcinomas in liver needle biopsies. Gut 2011;60(7):967–976
Sasaki M, Ikeda H, Itatsu K, Yamaguchi J, Sawada S, Minato H, Ohta T, Nakanuma Y. The overexpression of polycomb group proteins Bmi1 and EZH2 is associated with the progression and aggressive biological behavior of hepatocellular carcinoma. Lab Invest 2008;88(8):873–882
Sudo T, Utsunomiya T, Mimori K, Nagahara H, Ogawa K, Inoue H, Wakiyama S, Fujita H, Shirouzu K, Mori M. Clinicopathological significance of EZH2 mRNA expression in patients with hepatocellular carcinoma. Br J Cancer 2005;92(9):1754–1758
Au SL, Wong CC, Lee JM, Fan DN, Tsang FH, Ng IO, Wong CM. Enhancer of zeste homolog 2 epigenetically silences multiple tumor suppressor microRNAs to promote liver cancer metastasis. Hepatology 2012;56(2):622–631
Cai MY, Hou JH, Rao HL, Luo RZ, Li M, Pei XQ, Lin MC, Guan XY, Kung HF, Zeng YX, Xie D. High expression of H3K27me3 in human hepatocellular carcinomas correlates closely with vascular invasion and predicts worse prognosis in patients. Mol Med 2011;17(1–2):12–20
Takigawa Y, Brown AM. Wnt signaling in liver cancer. Curr Drug Targets 2008;9(11):1013–1024
Cheng AS, Lau SS, Chen Y, Kondo Y, Li MS, Feng H, Ching AK, Cheung KF, Wong HK, Tong JH, Jin H, Choy KW, Yu J, To KF, Wong N, Huang TH, Sung JJ. EZH2-mediated concordant repression of Wnt antagonists promotes β-catenin-dependent hepatocarcinogenesis. Cancer Res 2011;71(11):4028–4039
Gramantieri L, Fornari F, Callegari E, Sabbioni S, Lanza G, Croce CM, Bolondi L, Negrini M. MicroRNA involvement in hepatocellular carcinoma. J Cell Mol Med 2008;12(6A):2189–2204
Ji J, Wang XW. New kids on the block: diagnostic and prognostic microRNAs in hepatocellular carcinoma. Cancer Biol Ther 2009;8(18):1686–1693
Huntzinger E, Izaurralde E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat Rev Genet 2011;12(2):99–110
Zheng F, Liao YJ, Cai MY, Liu YH, Liu TH, Chen SP, Bian XW, Guan XY, Lin MC, Zeng YX, Kung HF, Xie D. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut 2012;61(2):278–289
Varambally S, Cao Q, Mani RS, Shankar S, Wang X, Ateeq B, Laxman B, Cao X, Jing X, Ramnarayanan K, Brenner JC, Yu J, Kim JH, Han B, Tan P, Kumar-Sinha C, Lonigro RJ, Palanisamy N, Maher CA, Chinnaiyan AM. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 2008;322(5908):1695–1699
Sander S, Bullinger L, Klapproth K, Fiedler K, Kestler HA, Barth TF, Möller P, Stilgenbauer S, Pollack JR, Wirth T. MYC stimulates EZH2 expression by repression of its negative regulator miR-26a. Blood 2008;112(10):4202–4212
Cao Q, Mani RS, Ateeq B, Dhanasekaran SM, Asangani IA, Prensner JR, Kim JH, Brenner JC, Jing X, Cao X, Wang R, Li Y, Dahiya A, Wang L, Pandhi M, Lonigro RJ, Wu YM, Tomlins SA, Palanisamy N, Qin Z, Yu J, Maher CA, Varambally S, Chinnaiyan AM. Coordinated regulation of polycomb group complexes through microRNAs in cancer. Cancer Cell 2011;20(2):187–199
Squazzo SL, O’Geen H, Komashko VM, Krig SR, Jin VX, Jang SW, Margueron R, Reinberg D, Green R, Farnham PJ. Suz12 binds to silenced regions of the genome in a cell-type-specific manner. Genome Res 2006;16(7):890–900
Kirmizis A, Bartley SM, Farnham PJ. Identification of the polycomb group protein SU(Z)12 as a potential molecular target for human cancer therapy. Mol Cancer Ther 2003;2(1):113–121
Wang WH, Studach LL, Andrisani OM. Proteins ZNF198 and SUZ12 are down-regulated in hepatitis B virus (HBV) X proteinmediated hepatocyte transformation and in HBV replication. Hepatology 2011;53(4):1137–1147
Studach LL, Menne S, Cairo S, Buendia MA, Hullinger RL, Lefrançois L, Merle P, Andrisani OM. A subset of Suz12/PRC2 target genes is activated during HBV replication and liver carcinogenesis associated with hepatitis B virus X protein. Hepatology 2012;56(4):1240–1251
Gao Z, Zhang J, Bonasio R, Strino F, Sawai A, Parisi F, Kluger Y, Reinberg D. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol Cell 2012;45(3):344–356
Morey L, Pascual G, Cozzuto L, Roma G, Wutz A, Benitah SA, Di Croce L. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell Stem Cell 2012;10(1):47–62
van der Lugt NM, Alkema M, Berns A, Deschamps J. The Polycomb-group homolog Bmi-1 is a regulator of murine Hox gene expression. Mech Dev 1996;58(1–2):153–164
Akasaka T, Kanno M, Balling R, Mieza MA, Taniguchi M, Koseki H. A role for mel-18, a polycomb group-related vertebrate gene, during theanteroposterior specification of the axial skeleton. Development 1996;122(5):1513–1522
Effendi K, Mori T, Komuta M, Masugi Y, Du W, Sakamoto M. Bmi-1 gene is upregulated in early-stage hepatocellular carcinoma and correlates with ATP-binding cassette transporter B1 expression. Cancer Sci 2010;101(3):666–672
Wang H, Pan K, Zhang HK, Weng DS, Zhou J, Li JJ, Huang W, Song HF, Chen MS, Xia JC. Increased polycomb-group oncogene Bmi-1 expression correlates with poor prognosis in hepatocellular carcinoma. J Cancer Res Clin Oncol 2008;134(5):535–541
Schuringa JJ, Vellenga E. Role of the polycomb group gene BMI1 in normal and leukemic hematopoietic stem and progenitor cells. Curr Opin Hematol 2010;17(4):294–299
Xu CR, Lee S, Ho C, Bommi P, Huang SA, Cheung ST, Dimri GP, Chen X. Bmi1 functions as an oncogene independent of Ink4A/Arf repression in hepatic carcinogenesis. Mol Cancer Res 2009;7(12):1937–1945
Chiba T, Zheng YW, Kita K, Yokosuka O, Saisho H, Onodera M, Miyoshi H, Nakano M, Zen Y, Nakanuma Y, Nakauchi H, Iwama A, Taniguchi H. Enhanced self-renewal capability in hepatic stem/ progenitor cells drives cancer initiation. Gastroenterology 2007;133(3):937–950
Chiba T, Miyagi S, Saraya A, Aoki R, Seki A, Morita Y, Yonemitsu Y, Yokosuka O, Taniguchi H, Nakauchi H, Iwama A. The polycomb gene product BMI1 contributes to the maintenance of tumorinitiating side population cells in hepatocellular carcinoma. Cancer Res 2008;68(19):7742–7749
O’Loghlen A, Muñoz-Cabello AM, Gaspar-Maia A, Wu HA, Banito A, Kunowska N, Racek T, Pemberton HN, Beolchi P, Lavial F, Masui O, Vermeulen M, Carroll T, Graumann J, Heard E, Dillon N, Azuara V, Snijders AP, Peters G, Bernstein E, Gil J. MicroRNA regulation of Cbx7 mediates a switch of Polycomb orthologs during ESC differentiation. Cell Stem Cell 2012;10(1):33–46
Karamitopoulou E, Pallante P, Zlobec I, Tornillo L, Carafa V, Schaffner T, Borner M, Diamantis I, Esposito F, Brunner T, Zimmermann A, Federico A, Terracciano L, Fusco A. Loss of the CBX7 protein expression correlates with a more aggressive phenotype in pancreatic cancer. Eur J Cancer 2010;46(8):1438–1444
Pallante P, Terracciano L, Carafa V, Schneider S, Zlobec I, Lugli A, Bianco M, Ferraro A, Sacchetti S, Troncone G, Fusco A, Tornillo L. The loss of the CBX7 gene expression represents an adverse prognostic marker for survival of colon carcinoma patients. Eur J Cancer 2010;46(12):2304–2313
Forzati F, Federico A, Pallante P, Abbate A, Esposito F, Malapelle U, Sepe R, Palma G, Troncone G, Scarfò M, Arra C, Fedele M, Fusco A. CBX7 is a tumor suppressor in mice and humans. J Clin Invest 2012;122(2):612–623
Brown JL, Mucci D, Whiteley M, Dirksen ML, Kassis JA. The Drosophila Polycomb group gene pleiohomeotic encodes a DNA binding protein with homology to the transcription factor YY1. Mol Cell 1998;1(7):1057–1064
Wilkinson FH, Park K, Atchison ML. Polycomb recruitment to DNA in vivo by the YY1 REPO domain. Proc Natl Acad Sci USA 2006;103(51):19296–19301
Notarbartolo M, Giannitrapani L, Vivona N, Poma P, Labbozzetta M, Florena A M, Porcasi R, Rosario Muggeo V M, Sandonato L, Cervello M, Montalto G, D’Alessandro N. Frequent alteration of the Yin Yang 1/Raf-1 kinase inhibitory protein ratio in hepatocellular carcinoma. OMICS: A Journal of Integrative Biology 2011;15(5):267–272
Zhang S, Jiang T, Feng L, Sun J, Lu H, Wang Q, Pan M, Huang D, Wang X, Wang L, Jin H. Yin Yang-1 suppresses differentiation of hepatocellular carcinoma cells through the downregulation of CCAAT/enhancer-binding protein alpha. J Mol Med (Berl) 2012;90(9):1069–1077
Zhang L, Cai X, Chen K, Wang Z, Wang L, Ren M, Huang A, Tang H. Hepatitis B virus protein up-regulated HLJ1 expression via the transcription factor YY1 in human hepatocarcinoma cells. Virus Res 2011;157(1):76–81
García E, Marcos-Gutiérrez C, del Mar Lorente M, Moreno JC, Vidal M. RYBP, a new repressor protein that interacts with components of the mammalian polycomb complex, and with the transcription factor YY1. EMBO J 1999;18(12):3404–3418
Tavares L, Dimitrova E, Oxley D, Webster J, Poot R, Demmers J, Bezstarosti K, Taylor S, Ura H, Koide H, Wutz A, Vidal M, Elderkin S, Brockdorff N. RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell 2012;148(4):664–678
Chen D, Zhang J, Li M, Rayburn ER, Wang H, Zhang R. RYBP stabilizes p53 by modulating MDM2. EMBO Rep 2009;10(2):166–172
Baylin SB, Jones PA. A decade of exploring the cancer epigenomebiological and translational implications. Nat Rev Cancer 2011;11(10):726–734
Chiang PK, Cantoni GL. Perturbation of biochemical transmethylations by 3-deazaadenosine in vivo. Biochem Pharmacol 1979;28(12):1897–1902
Tan J, Yang X, Zhuang L, Jiang X, Chen W, Lee PL, Karuturi RK, Tan PB, Liu ET, Yu Q. Pharmacologic disruption of Polycombrepressive complex 2-mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev 2007;21(9):1050–1063
Miranda TB, Cortez CC, Yoo CB, Liang G, Abe M, Kelly TK, Marquez VE, Jones PA. DZNep is a global histone methylation inhibitor that reactivates developmental genes not silenced by DNA methylation. Mol Cancer Ther 2009;8(6):1579–1588
Fiskus W, Wang Y, Sreekumar A, Buckley KM, Shi H, Jillella A, Ustun C, Rao R, Fernandez P, Chen J, Balusu R, Koul S, Atadja P, Marquez VE, Bhalla KN. Combined epigenetic therapy with the histone methyltransferase EZH2 inhibitor 3-deazaneplanocin A and the histone deacetylase inhibitor panobinostat against human AML cells. Blood 2009;114(13):2733–2743
Cheng LL, Itahana Y, Lei ZD, Chia NY, Wu Y, Yu Y, Zhang SL, Thike AA, Pandey A, Rozen S, Voorhoeve PM, Yu Q, Tan PH, Bay BH, Itahana K, Tan P. TP53 genomic status regulates sensitivity of gastric cancer cells to the histone methylation inhibitor 3-deazaneplanocin A (DZNep). Clin Cancer Res 2012 1;18(15):4201–4212
Chiba T, Suzuki E, Negishi M, Saraya A, Miyagi S, Konuma T, Tanaka S, Tada M, Kanai F, Imazeki F, Iwama A, Yokosuka O. 3-Deazaneplanocin A is a promising therapeutic agent for the eradication of tumor-initiating hepatocellular carcinoma cells. Int J Cancer 2012;130(11):2557–2567
Chen Y, Lin MC, Yao H, Wang H, Zhang AQ, Yu J, Hui CK, Lau GK, He ML, Sung J, Kung HF. Lentivirus-mediated RNA interference targeting enhancer of zeste homolog 2 inhibits hepatocellular carcinoma growth through down-regulation of stathmin. Hepatology 2007;46(1):200–208
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Au, S.LK., Ng, I.OL. & Wong, CM. Epigenetic dysregulation in hepatocellular carcinoma: focus on polycomb group proteins. Front. Med. 7, 231–241 (2013). https://doi.org/10.1007/s11684-013-0253-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11684-013-0253-7