Abstract
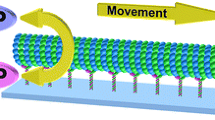
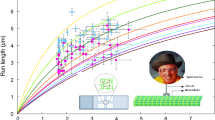
A microtubule gliding assay is a biological experiment observing the dynamics of microtubules driven by motor proteins fixed on a glass surface. When appropriate microtubule interactions are set up on gliding assay experiments, microtubules often organize and create higher-level dynamics such as ring and bundle structures. In order to reproduce such higher-level dynamics on computers, we have been focusing on making a real-time 3D microtubule simulation. This real-time 3D microtubule simulation enables us to gain more knowledge on microtubule dynamics and their swarm movements by means of adjusting simulation parameters in a real-time fashion. One of the technical challenges when creating a real-time 3D simulation is balancing the 3D rendering and the computing performance. Graphics processor unit (GPU) programming plays an essential role in balancing the millions of tasks, and makes this real-time 3D simulation possible. By the use of general-purpose computing on graphics processing units (GPGPU) programming we are able to run the simulation in a massively parallel fashion, even when dealing with more complex interactions between microtubules such as overriding and snuggling. Due to performance being an important factor, a performance model has also been constructed from the analysis of the microtubule simulation and it is consistent with the performance measurements on different GPGPU architectures with regards to the number of cores and clock cycles.
Similar content being viewed by others
References
S. Murata, A. Konagaya, S. Kobayashi, H. Saito, M. Hagiya. Molecular robotics: A new paradigm for artifacts. New Generation Computing, vol. 31, no. 1, pp. 27–45, 2013.
J. Howard, A. J. Hudspeth, R. D. Vale. Movement of microtubules by single kinesin molecules. Nature, vol. 342, no. 6246, pp. 154–158, 1989.
K. J. Böhm, R. Stracke, P. Mühlig, E. Unger. Motor protein-driven unidirectional transport of micrometer-sized cargoes across isopolar microtubule arrays. Nanotechnology, vol. 12, no. 3, pp. 238–244, 2001.
T. Fischer, A. Agarwal, H. Hess. A smart dust biosensor powered by kinesin motors. Nature Nanotechnology, vol.4, no. 3, pp. 162–166, 2009.
A. M. R. Kabir, S. Wada, D. Inoue, Y. Tamura, T. Kajihara, H. Mayama, K. Sada, A. Kakugo, J. P. Gong. Formation of ring-shaped assembly of microtubules with a narrow size distribution at an air–buffer interface. Soft Matter, vol. 8, no. 42, pp. 10863–10867, 2012.
D. Inoue, A. M.R. Kabir, H. Mayama, J. P.Gong, K. Sada, A. Kakugo. Growth of ring-shaped microtubule assemblies through stepwise active self-organisation. Soft Matter, vol. 9, no. 29, pp. 7061–7068, 2013.
A. M. R. Kabir, D. Inoue, A. Kakugo, A. Kamei, J. P. Gong. Prolongation of the active lifetime of a biomolecular motor for in vitro motility assay by using an inert atmosphere. Langmuir, vol. 27, no. 22, pp. 13659–13668, 2011.
A. M. R. Kabir, D. Inoue, A. Kakugo, K. Sada, J. P. Gong. Active self-organization of microtubules in an inert chamber system. Polymer Journal, vol. 44, no. 6, pp. 607–611, 2012.
P. Kraikivski, R. Lipowsky, J. Kierfeld. Enhanced ordering of interacting filaments by molecular motors. Physical Review Letters, vol. 96, no. 25, pp. 258103, 2006.
K. Y. Kong, A. I. Marcus, P. Giannakakou, C. Alberti, M. D. Wang. A two dimensional simulation of microtubule dynamics. In Proceedings of International Conference on Information Technology and Applications in Biomedicine, IEEE, Shenzhen, China, pp. 461–462, 2008.
A. Sherrod, W. Jones. Beginning DirectX 11 Game Programming, Boston, USA: Course Technology PTR, 2011.
F. D. Luna. Introduction to 3D Game Programming: With DirectX 11, Dulles, USA: Mercury Learning & Information, 2012.
Author information
Authors and Affiliations
Corresponding author
Additional information
Recommended by Guest Editor Song Yang
This work was supported by a Grant-in-Aid for Scientific Research on Innovation Areas “Molecular Robotics” (No. 24104004) of the Ministry of Education, Culture, Sports, Science, and Technology, Japan.
Gregory Gutmann received the B. Sc. degree with a double major in computer science and East Asian studies from John Carroll University, USA in 2013. In the summers of 2012 and 2013, he was an intern at National Aeronautics and Space Administration (NASA) in The NASA Center for Climate Simulation and The Earth Sciences Division Departments. He received the M.Eng. degree in the Department of Computational Intelligence and Systems Science from Tokyo Institute of Technology, Japan in 2015. Currently, he is a Ph.D. degree candidate at Tokyo Institute of Technology, Japan.
His research interest covers GPGPU computing, high performance computing, 3D visualization and various science fields. His current research is focused on molecular robotics.
ORCID iD: 0000-0001-9157-6344
Daisuke Inoue received the B. Sc. degree with Division of Marine Life Science, Faculty and Graduate School of Fisheries Sciences, Hokkaido University, Japan in 2010, and received his master of Division of Biological Sciences, Graduate School of Life Science, Hokkaido University, Japan in 2012. Currently, he obtained the Ph.D. degree at Graduate School of Chemical Sciences and Engineering, Hokkaido University, Japan in 2015. He received a JSPS Research Fellowship for Young Scientists in 2014. From 2015, he serves as a postdoctoral researcher in Faculty of Science, Hokkaido University, Japan. Currently, he is studying pattern formation of microtubules in dynamic systems and their responses to change in their surrounding environment.
His research interests include self-organization of active matters, swarm robotics and symmetry breaking in the bio-system.
ORCID iD: 0000-0002-6299-1036
Akira Kakugo is an associate professor at the Hokkaido University, Japan. He obtained the Ph. D. degree of Science from the Hokkaido University, Japan in 2003. He has targeted bio-molecular motor such as actin/myosin and microtubule/kinesin as a power system of the molecular robots and developed prototype models.
His research interests include the designing and engineering of molecular robot that is a set of molecular devices such as sensors, logic gates, and actuators integrated into a consistent system.
ORCID iD: 0000-0002-1591-867X
Akihiko Konagaya received the B. Sc., M. Sc. and Ph.D. degrees of engineering from Tokyo Institute of Technology, Japan in 1978, 1980 and 1995, respectively. He joined NEC Corporation in 1980. Then, he became a professor of Japan Advanced Institute of Science and Technology (JAIST) and a project director of RIKEN Genomic Science Center (GSC) in 1997 and 2003, respectively. Since 2009, he has been a professor of the Department of Computational Intelligence and Systems Science in Tokyo Institute of Technology. He is also a PI of the Amoeba Team in the Grant-in-Aid for Scientific Research on Innovative Areas “Molecular Robotics” of the Ministry of Education, Culture, Sports, Science, and Technology, Japan.
ORCID iD: 0000-0001-6846-7449
Rights and permissions
About this article
Cite this article
Gutmann, G., Inoue, D., Kakugo, A. et al. Real-time 3D microtubule gliding simulation accelerated by GPU computing. Int. J. Autom. Comput. 13, 108–116 (2016). https://doi.org/10.1007/s11633-015-0947-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11633-015-0947-1