Abstract
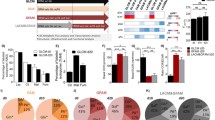
Knockout of multifunction gene cysteine- and glycine-rich protein 3 (CSRP3) in cardiomyocytes (CMs) of mice leads to heart dilation, severely affecting its functions. In humans, CSRP3 mutations are associated with hypertrophic (HCM) and dilated cardiomyopathy (DCM). The absence of the CSRP3 expression produces unknown effects on in vitro neonatal CMs’ metabolism. The metabolome changes in culture media conditioned by CSRP3 knockout (KO-CSRP3), and wild type (WT) neonatal cardiomyocytes were investigated under untreated or after metabolic challenging conditions produced by isoproterenol (ISO) stimulation, by in vitro high-resolution proton magnetic resonance spectroscopy (1H-MRS)-based metabolomics. Metabolic differences between neonatal KO-CSRP3 and WT rats’ CMs were identified. After 72 h of culture, ISO administration was associated with increased CMs’ energy requirements and increased levels of threonine, alanine, and 3-hydroxybutyrate in both neonatal KO-CSRP3 and WT CMs conditioned media. When compared with KO-CSRP3, culture media derived from WT cells presented higher lactate concentrations either under basal or ISO-stimulated conditions. The higher activity of ketogenic biochemical pathways met the elevated energy requirements of the contractile cells. Both cells are considered phenotypically indistinguishable in the neonatal period of animal lives, but the observed metabolic stress responses of KO-CSRP3 and WT CMs to ISO were different. KO-CSRP3 CMs produced less lactate than WT CMs in both basal and stimulated conditions. Mainly, ISO-stimulated conditions produced evidence for lactate overload within KO-CSRP3 CMs, while WT CMs succeeded to manage the metabolic stress. Thus, 1H-MRS-based metabolomics was suitable to identify early inefficient energetic metabolism in neonatal KO-CSRP3 CMs. These results may reflect an apparent lower lactate transport and consumption, in association with protein catabolism.




Similar content being viewed by others
References
Aliu E, Kanungo S, Arnold GL (2018) Amino acid disorders. Ann Transl Med 6(24):471–471. https://doi.org/10.21037/atm.2018.12.12
Arber S, Hunter JJ, Ross J, Hongo M, Sansig G, Borg J et al (1997) MLP-deficient mice exhibit a disruption of cardiac cytoarchitectural organization, dilated cardiomyopathy, and heart failure. Cell 88(3):393–403. Retrieved from https://doi.org/10.1186/2193-1801-3-470
Bacchi PS, Bloise AC, Bustos SO, Zimmermann L, Chammas R, Rabbani SR (2014) Metabolism under hypoxia in Tm1 murine melanoma cells is affected by the presence of galectin-3, a metabolomics approach. SpringerPlus 3:470
Boateng SY, Senyo SE, Qi L, Goldspink PH, Russell B (2009) Myocyte remodeling in response to hypertrophic stimuli requires nucleocytoplasmic shuttling of muscle LIM protein. J Mol Cell Cardiol 47(4):426–435. https://doi.org/10.1016/j.yjmcc.2009.04.006
Campos LCG, Ribeiro-Silva JC, Menegon AS, Barauna VG, Miyakawa AA, Krieger JE (2018) Cyclic stretch-induced Crp3 sensitizes vascular smooth muscle cells to apoptosis during vein arterialization remodeling. Clin Sci 132(4):449–459. https://doi.org/10.1042/cs20171601
Chaudhari U, Ellis JK, Wagh V, Nemade H, Hescheler J, Keun HC, Sachinidis A (2017) Metabolite signatures of doxorubicin induced toxicity in human induced pluripotent stem cell-derived cardiomyocytes. Amino Acids 49(12):1955–1963. https://doi.org/10.1007/s00726-017-2419-0
Ehsan M, Kelly M, Hooper C, Yavari A, Beglov J, Bellahcene M et al (2018) Mutant muscle LIM protein C58G causes cardiomyopathy through protein depletion. J Mol Cell Cardiol 121:287–296. https://doi.org/10.1016/j.yjmcc.2018.07.248
Ellinger JJ, Chylla RA, Ulrich EL, Markley JL (2013) Databases and software for NMR-based metabolomics. Curr Metabolomics 1(1):28–40. https://doi.org/10.2174/2213235x11301010028
Emwas AHM (2015) The strengths and weaknesses of NMR spectroscopy and mass spectrometry with particular focus on metabolomics research. Methods Mol Biol 1277:161–193. https://doi.org/10.1007/978-1-4939-2377-9_13
Finckenberg P, Mervaala E (2010) Novel regulators and drug targets of cardiac hypertrophy. J Hypertens 28(Suppl 1):S33–S38. https://doi.org/10.1097/01.hjh.0000388492.73954.0b
Geier C, Perrot A, Özcelik C, Binner P, Counsell D, Hoffmann K et al (2003) Mutations in the human muscle LIM protein gene in families with hypertrophic cardiomyopathy. Circulation 107(10):1390–1395. https://doi.org/10.1161/01.CIR.0000056522.82563.5F
Griffin JL, Shockcor JP (2004) Metabolic profiles of cancer cells. Nat Rev Cancer 4(7):551
Heckler CE (2005) Applied multivariate statistical analysis: applied multivariate statistical analysis. Technometrics 47(4):517–517. https://doi.org/10.1198/tech.2005.s319
Hernandez-Carretero A, Weber N, LaBarge SA, Peterka V, Doan NYT, Schenk S, Osborn O (2018) Cysteine- and glycine-rich protein 3 regulates glucose homeostasis in skeletal muscle. Am J Physiol-Endocrinol Metab 315(2):E267–E278. https://doi.org/10.1152/ajpendo.00435.2017
Jensen L, Neri E, Bassaneze V, De Almeida Oliveira NC, Dariolli R, Turaça LT, … Krieger JE (2018) Integrated molecular, biochemical, and physiological assessment unravels key extraction method mediated influences on rat neonatal cardiomyocytes. J Cell Physiol 233(7):5420–5430. https://doi.org/10.1002/jcp.26380
Kaddurah-Daouk R, Kristal BS, Weinshilboum RM (2008) Metabolomics: a global biochemical approach to drug response and disease. Annu Rev Pharmacol Toxicol 48:653–683. https://doi.org/10.1146/annurev.pharmtox.48.113006.094715
Klos M, Morgenstern S, Hicks K, Suresh S, Devaney EJ (2019) The effects of the ketone body β-hydroxybutyrate on isolated rat ventricular myocyte excitation-contraction coupling. Arch Biochem Biophys 662:143–150. https://doi.org/10.1016/J.ABB.2018.11.027
Kostidis S (2018) Quantitative analysis of central energy metabolism in cell culture samples. Methods Mol Biol 1730:329–342. https://doi.org/10.1007/978-1-4939-7592-1_25
Lange S, Gehmlich K, Lun AS, Blondelle J, Hooper C, Dalton ND et al (2016) MLP and CARP are linked to chronic PKCα signaling in dilated cardiomyopathy. Nat Commun 7(1):12120. https://doi.org/10.1038/ncomms12120
Li R, Yan G, Zhang Q, Jiang Y, Sun H, Hu Y et al (2013) miR-145 inhibits isoproterenol-induced cardiomyocyte hypertrophy by targeting the expression and localization of GATA6. FEBS Lett 587:1754–1761
Li X, Lu WJ, Li Y, Wu F, Bai R, Ma S et al (2019) MLP-deficient human pluripotent stem cell derived cardiomyocytes develop hypertrophic cardiomyopathy and heart failure phenotypes due to abnormal calcium handling. Cell Death Dis 10(8). https://doi.org/10.1038/s41419-019-1826-4
Lindon JC, Holmes E, Nicholson JK (2006) Metabonomics techniques and applications to pharmaceutical research & development. Pharm Res 23(6):1075–1088. https://doi.org/10.1007/s11095-006-0025-z
Lindon JC, Nicholson JK, Holmes E (2007) The Handbook of Metabonomics and Metabolomics. In: The Handbook of Metabonomics and Metabolomics. https://doi.org/10.1016/B978-0-444-52841-4.X5000-0
Madhu B, Dadulescu M, Griffiths J (2015) Artifacts in 1H NMR-based metabolomic studies on cell cultures. MAGMA 28(2):161–171. https://doi.org/10.1007/s10334-014-0458-z
Madsen R, Lundstedt T, Trygg J (2010) Chemometrics in metabolomics - a review in human disease diagnosis. Anal Chim Acta 659:23–33
Mandenius C-F, Steel D, Noor F, Meyer T, Heinzle E, Asp J et al (2011) Cardiotoxicity testing using pluripotent stem cell-derived human cardiomyocytes and state-of-the-art bioanalytics: a review. J Appl Toxicol 31(3):191–205. https://doi.org/10.1002/jat.1663
Martínez MS, García A, Luzardo E, Chávez-Castillo M, Olivar LC, Salazar J et al (2017) Energetic metabolism in cardiomyocytes: molecular basis of heart ischemia and arrhythmogenesis. Vessel Plus 1(12). https://doi.org/10.20517/2574-1209.2017.34
Martins-Bach AB, Bloise AC, Vainzof M, Rabbani SR (2012) Metabolic profile of dystrophic mdx mouse muscles analyzed with in vitro magnetic resonance spectroscopy (MRS). Magn Reson Imaging 30:1167–1176
Massad E, de Menezes RX, Silveira PSP, Ortega NRS (2004) Métodos Quantitativos em Medicina. Manole
Mohmad-Saberi SE, Hashim YZH-Y, Mel M, Amid A, Ahmad-Raus R, Packeer-Mohamed V (2013) Metabolomics profiling of extracellular metabolites in CHO-K1 cells cultured in different types of growth media. Cytotechnology 65(4):577–586. https://doi.org/10.1007/s10616-012-9508-4
Nalbandian M, Takeda M (2016) Lactate as a signaling molecule that regulates exercise-induced adaptations. Biology 5(4):38. https://doi.org/10.3390/biology5040038
Nicholson JK (2010) 2020 visions. Nature 463(7277):26–32. https://doi.org/10.1038/463026a
Pereira T, Ivanova G, Caseiro AR, Barbosa P, Bártolo PJ, Santos JD et al (2014) MSCs conditioned media and umbilical cord blood plasma metabolomics and composition. PLoS ONE 9(11):e113769. https://doi.org/10.1371/journal.pone.0113769
Saborano R, Eraslan Z, Roberts J, Khanim FL, Lalor PF, Reed MAC, Günther UL (2019) A framework for tracer-based metabolism in mammalian cells by NMR. Sci Rep 9(1). https://doi.org/10.1038/s41598-018-37525-3
Triba MN, Starzec A, Bouchemal N, Guenin E, Perret GY, Le Moyec L (2010) Metabolomic profiling with NMR discriminates between biphosphonate and doxorubicin effects on B16 melanoma cells. NMR Biomed 23:1009–1016. https://doi.org/10.1002/nbm.1516
Wishart DS, Jewison T, Guo AC, Wilson M, Knox C (2013) HMDB 3.0 - The human metabolome database in 2013. Nucleic Acids Res 41:801–807
Worley B, Powers R (2013) Multivariate analysis in metabolomics. Curr Metabolomics 1(1):92–107. https://doi.org/10.2174/221323X11301010092
Xia J, Wishart DS (2016) Using MetaboAnalyst 3.0 for comprehensive metabolomics data analysis. Curr Protoc Bioinformatics 55:14.10.1–14.10.91
Zhang Y, Sekar RB, McCulloch AD, Tung L (2008) Cell cultures as models of cardiac mechanoelectrical feedback. Prog Biophys Mol 97:367–382
Acknowledgments
The authors would like to acknowledge the São Paulo Research Foundation (FAPESP) (grants 2011/19678-1; 2013/17368-0; 2014/22102-2; 2014/21646-9; 2018/20910-5) and Medical Sciences Graduate Program-CAPES/PROEX for financial support; LNBio-CNPEM (20992) for providing spectrometer and facilities; Laboratory of Genetics and Molecular Cardiology, Heart Institute (InCor) - University of Sao Paulo School of Medicine for providing cells and scientific assistance; and National Institute of Science and Technology Complex Fluids (INCT-FCX) for daily financial aid.
Author information
Authors and Affiliations
Contributions
ACB and VB equally contributed to cell experiments. ACB contributed with sample preparation, NMR experiments, data processing, statistical analysis, and biological interpretations and drafted the manuscript. LFG critically revised the manuscript. JAS, IVB, VB, and AMA contributed to the design of the research and cell experimental setup. All authors agree to be fully accountable for ensuring the integrity and accuracy of the work and read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethics approval
Approval was obtained from the institutional review board from the University of Sao Paulo, School of Medicine, Brazil (#340/12).
Additional information
Editor: Tetsuji Okamoto
Electronic supplementary material
ESM 1
(DOCX 861 kb)
Rights and permissions
About this article
Cite this article
Bloise, A.C., dos Santos, J.A., de Brito, I.V. et al. Discriminating aspects of global metabolism of neonatal cardiomyocytes from wild type and KO-CSRP3 rats using proton magnetic resonance spectroscopy of culture media samples. In Vitro Cell.Dev.Biol.-Animal 56, 604–613 (2020). https://doi.org/10.1007/s11626-020-00497-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-020-00497-8