Abstract
Purpose
To predict solid and micropapillary components in lung invasive adenocarcinoma using radiomic analyses based on high-spatial-resolution CT (HSR-CT).
Materials and methods
For this retrospective study, 64 patients with lung invasive adenocarcinoma were enrolled. All patients were scanned by HSR-CT with 1024 matrix. A pathologist evaluated subtypes (lepidic, acinar, solid, micropapillary, or others). Total 61 radiomic features in the CT images were calculated using our modified texture analysis software, then filtered and minimized by least absolute shrinkage and selection operator (LASSO) regression to select optimal radiomic features for predicting solid and micropapillary components in lung invasive adenocarcinoma. Final data were obtained by repeating tenfold cross-validation 10 times. Two independent radiologists visually predicted solid or micropapillary components on each image of the 64 nodules with and without using the radiomics results. The quantitative values were analyzed with logistic regression models. The receiver operating characteristic curves were generated to predict of solid and micropapillary components. P values < 0.05 were considered significant.
Results
Two features (Coefficient Variation and Entropy) were independent indicators associated with solid and micropapillary components (odds ratio, 30.5 and 11.4; 95% confidence interval, 5.1–180.5 and 1.9–66.6; and P = 0.0002 and 0.0071, respectively). The area under the curve for predicting solid and micropapillary components was 0.902 (95% confidence interval, 0.802 to 0.962). The radiomics results significantly improved the accuracy and specificity of the prediction of the two radiologists.
Conclusion
Two texture features (Coefficient Variation and Entropy) were significant indicators to predict solid and micropapillary components in lung invasive adenocarcinoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the leading cause of cancer death worldwide [1]. Nearly 70–90% of all surgically resected lung cancers are invasive adenocarcinomas. The morphologic and pathologic manifestations of invasive adenocarcinoma (IVA) have been well characterized according to a multidisciplinary classification proposed by the International Association for the Study of Lung Cancer, the American Thoracic Society, and the European Respiratory Society (IASLC/ATS/ERS) [2]. IVA is divided into five subtypes: lepidic, acinar, papillary, solid, and micropapillary patterns. Most IVAs appear as a sequential tissue transition between two or more histologic patterns [2]. Micropapillary and solid pattern predominant tumors are known to have worse survival prognosis and higher recurrence rates. Furthermore, they are considered to have a worse prognosis even when their pattern is not predominant [3,4,5]. In particular, micropapillary pattern is associated with an increased risk of local recurrence in patients treated with limited resection [6]. Therefore, it is important to predict micropapillary and solid components when deciding on the surgical approach.
Preoperative diagnostic methods include percutaneous needle biopsy or bronchoscopy, but these are invasive, especially in patients with underlying diseases such as emphysema, and are sometimes difficult in patients with poor performance status. There remain problems with the accuracy and reliability of biopsy, as only a portion of the tissue is sampled. CT is a relatively simple and minimally invasive examination at the earliest stage for treating lung cancer than biopsy. Some samples do not provide a complete picture of the tumor, and the nature of the tumor may differ depending on where the sample is taken.
Imaging, on the other hand, is superior in that it provides an entire image of the tumor.
Some studies have investigated the correlation between CT morphologic and histologic features [7]. High-spatial-resolution (HSR) CT scanners, which have a spatial resolution of up to 150 mm, have been available in the clinical setting since 2017.
Recently, HSR-CT has been reported to improve diagnostic performance by subjective assessment of pathological invasiveness in adenocarcinoma due to its improved spatial resolution [8]. However, it is still difficult to distinguish subtypes of lung adenocarcinoma by morphological features. Radiomics features extracted from image data have been used in several recent studies [9].
The radiomics approach has also been used to distinguish subtypes of lung adenocarcinoma [10]. We hypothesized that the radiomics approach with improved spatial resolution of HSR-CT data could predict solid and micropapillary components in lung invasive adenocarcinoma.
The purpose of our study is to predict solid and micropapillary components in lung invasive adenocarcinoma using HSR-CT-based radiomic analysis.
Materials and methods
Study participants
This retrospective study was approved by our institution's internal ethics review board. Informed consent was waived for review of patient records and images. We reviewed patients who underwent surgery at a single institution between January 2018 and December 2019, and found 248 patients who underwent preoperative CT. Of these, a total of 157 patients with 159 nodules were included.
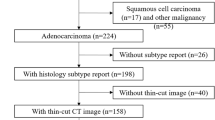
The inclusion criteria were as follows (Fig. 1): (I) histologically diagnosed adenocarcinoma, (II) CT was performed with a 1024 matrix and 0.25-mm thickness, (III) clinical stage I or II lung cancer, (IV) no previous treatment, (V) age 20 years or older.
Of the 159 nodules, 115 were histologically diagnosed with adenocarcinoma. Of the 115 nodules, four with atypical adenomatous hyperplasia and four with mucinous adenocarcinoma were excluded because they differed from non-mucinous adenocarcinoma in genetic factors and cancer genesis. 21 patients (21 nodules) were excluded: patients with nodules too small to evaluate, loss of raw CT data and inability to calculate radiomic features, and those who underwent CT more than 3 months before surgery.
A total of 64 nodules with minimally invasive adenocarcinoma or invasive adenocarcinoma were finally included in our study.
Image acquisition
All CT images were acquired on an HSR-CT scanner (Aquilion Precision: Canon Medical Systems) with 160 detector rows and 1792 detector channels. The CT parameters were as follows: tube voltage, 120 kVp; tube current, auto-exposure control; focus size, 0.6 × 0.6 mm; gantry rotation time, 0.5 s in spiral mode. Clinical images were acquired during breath-hold with full inspiration. All CT images were reconstructed with the following settings: matrix size, 1024 × 1024; slice thickness, 0.25 mm; slice interval, 0.25 mm; field of view, 34.5 cm; FC 51 (a lung algorithm) with adaptive iterative dose reduction. Volume CT dose index was 13.65 mGy ± 2.44.
Histopathologic data
All pathologic specimens were stained with hematoxylin–eosin and evaluated by pathologists at our institution according to the multidisciplinary adenocarcinoma criteria [2]. The extent of all five growth patterns (lepidic, acinar, papillary, micropapillary, and solid) was recorded by the percentage.
Image analysis
Commercially available software (WatchinGGO; LISIT, Co., Ltd., Tokyo, Japan) was used to segment pulmonary nodules. This software was modified for texture analysis. It can calculate radiomics features of selected two-dimensional (2D) CT images with a 1024 matrix. A total of 61 radiomic features were calculated in the segmented CT images using the software. The 61 radiomics features are listed in Online Resource 1.
Semi-automatic segmentation was performed in the maximum cross-sectional image of each tumor. The maximum cross-sectional image of each tumor was selected by a chest radiologist (MY). In the case of part-solid GGN lesions, the cross-sectional image with the maximum size of the solid component was selected. For each tumor, two independent radiologists (MT and YS) delineated regions of interest on the axial images (Fig. 2). Then, 61 radiomics features were calculated for each segmentation using our software. The average value of the radiomics features by two independent radiologists was statistically analyzed.
Subjective image analysis
Two image review sessions were performed. In the first session, two chest radiologists (R1 (NK) and R2 (YY)) independently classified the 64 nodules into two groups (score = 1: nodules with solid and/or micropapillary components, and score = 0: those without solid or micropapillary components). They were given the following reference information beforehand; micropapillary and solid patterns are more likely to be: (a) larger (total size≧2.5 cm), and/or (b) solid component predominant, and/or (c) spiculation or lobulation, whereas lepidic pattern is likely to contain a greater proportion of ground-glass opacities and/or an air bronchogram [7]. In the second session, performed 1 month after the first session, the same 64 nodules were reclassified into the two above-mentioned groups on the basis of the radiomics results which were significant indicators to predict the score = 1. At this time, they were also presented with the AUC value.
Statistical analysis
Statistical analyses were performed using commercially available software (MedCalc® Statistical Software version 20.216, Ostend, Belgium). Python 3.8.6 and scikit-learn 0.24.1 were used for the radiomics analysis.
The 61 radiomic features were filtered by least absolute shrinkage and selection operator (LASSO) regression with tenfold cross-validation 10 times. We determined feature importance by counting the number of times the feature had non-zero regression coefficients through repeated cross-validation. We created the radiomics features using a linear combination of the selected features. A radiomics score was calculated for each patient using a linear combination of the selected features, weighted according to their coefficients. For each higher relevant feature, the cutoff value that yielded the largest difference in the number of patients with and without solid and micropapillary components was determined using the receiver operating characteristic (ROC) method. Optimal cutoff values were determined for each variable separately using the Youden index (the maximum value of sensitivity and specificity). Associations between solid and micropapillary components and each binary group (score = 0 or score = 1) designated by the cutoff value for the ten radiomics features were evaluated by univariate logistic regression analysis. Significant features identified by univariate analysis were included in multiple logistic regression (stepwise method; P value of 0.05 or less was used for entry into the model and P value greater than 0.1 was selected for removal).
Diagnostic performance for predicting solid and micropapillary components was analyzed using ROC curves: sensitivity, specificity, and area under the curve (AUC). We used the “Comparison of ROC curves” of the MedCalc software to test the difference in the AUCs between predictive performance with and without radiomics features. Comparisons of accuracy, sensitivity, and specificity were performed using McNemar’s test. P value < 0.05 was considered significant.
Results
Clinical and pathological characteristics
Table1 shows the clinical and pathologic characteristics. Of the 64 patients, 16 patients had solid or micropapillary components and 48 patients had no solid or micropapillary components.
Prediction of solid and micropapillary components using radiomic analysis
We extracted ten optimal radiomic features for predicting solid and micropapillary components: LRHGE, LZHGE, Variance, GLN, Mean, GLCM: Correlation, Coefficient Variation, Entropy, GLV and Mean intensity (Fig. 3). The formula used to calculate the ten radiomic features is shown in Online Resource 2.
Importance of radiomic features for predicting solid and micropapillary components. We defined a radiomic feature as important if it had a non-zero LASSO regression coefficient over the repeated cross-validation. The maximum number of repetitions was 100. LRHGE large run high gray-level emphasis, LZHGE large zone high gray-level emphasis, GLN gray level non-uniformity, GLCM gray level co-occurrence matrix, GLV gray level variance, RLV run length variance, SZHGE small zone low gray-level emphasis, LRE long run emphasis, NGTDM neighborhood gray-tone-difference matrix, SRE short run emphasis, LZE large zone emphasis, HGRE high gray-level run emphasis, ZP zone percentage, HGZE high gray-level zone emphasis, LZLGE large zone low gray-level emphasis, RLN run-length non-uniformity, ZLV zone level variance, RP run percentage, SRHGE short run high gray-level emphasis, LRLGE long run low gray-level emphasis, SRLGE short run low gray-level emphasis, LGRE low gray-level run emphasis, SZE small zone emphasis, ZSN zone size non-uniformity, LGZE low gray-level zone emphasis, SZLGE small zone low gray-level emphasis, GLCM gray-level co-occurrence matrix, GLN gray-level non-uniformity, LRHGE large run high gray-level emphasis, GLV gray level variance, LZHGE large zone high gray-level emphasis
We determined cutoff values for each of the ten features using ROC analysis (Table 2). Univariable logistic regression analyses of these features revealed that all ten radiomic features were statistically significant for predicting micropapillary and solid patterns. Multiple logistic regression analysis revealed that two features (Coefficient of Variation and Entropy) were independent indicators associated with solid and micropapillary components (odds ratio, 30.5 and 11.4; 95% confidence interval, 5.1–180.5 and 1.9–66.6; and P = 0.0002 and 0.0071, respectively). The area under the curve for predicting solid and micropapillary components was 0.902 (95% confidence interval, 0.802 to 0.962).
Prediction of solid and micropapillary component by radiologists with and without radiomics results
The prediction by the two chest radiologists without the radiomic results were as follows: accuracy, sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV): R1 and R2, 54.7 and 54.7%, 100 and 93.7%, 39.6 and 41.7%, 35.6 and 34.9%, 100 and 95.2%, respectively). The radiomics results improved the accuracy and specificity significantly, while it worsened the sensitivity of R2, not significantly (Table 3). The AUC values for prediction with radiomics results were higher than without radiomics, but not significant: 0.698 (95% CI, 0.580–0.806) vs. 0.802 (0.684–0.891), P = 0.14 and 0.677 (0.549–0.789) vs. 0.823 (0.707–0.907), P = 0.07.
Discussion
Our study showed that radiomic analysis of HSR-CT data with 1024 matrix could achieve relatively accurate prediction performance of micropapillary and solid patterns. (AUC value of 0.90; 95% CI: 0.80–0.96). Considering that micropapillary and solid pattern predominant tumors had worse survival prognosis and higher recurrence rates, the good performance of our radiomics model could be an effective tool for prognosis prediction and clinical treatment planning decision, especially in cases where surgery or biopsy is not feasible. In particular, it may be more useful when a tumor has only a small portion of the solid and micropapillary pattern, which are difficult to detect by biopsy. Tissue biopsy is essential to examine the histopathology of a tumor. However, the spatially and temporally heterogeneous properties of tumors remain a problem for the accuracy and reliability of biopsy using only a portion of the tissue. If the CT-based radiomics approach, which is a simple and non-invasive examination, can predict the pathological diagnosis, the clinical benefit to patients is also considered to be significant.
Some previous studies have attempted to predict subtypes of lung adenocarcinoma. Some studies have reported radiologic features of each subtype, but they are not specific [7]. Recently, a radiomic approach has been used to predict micropapillary and solid patterns. Bingxi demonstrated that the radiomic method achieved good prediction performance of micropapillary and solid patterns (AUC value of 0.75; 95% CI: 0.65–0.85) [11]. Yunyu X et al. [12] reported that the combination of radiomics and clinical features had a statistically better prediction performance of micropapillary pattern (AUC = 0.739; 95% CI: 0.594–0.885) than radiomics analysis alone (AUC = 0.722; 95% CI: 0.574–0.870). A previous study showed that the radiomic approach achieved a higher PPV than radiologists for solid or micropapillary-predominant adenocarcinoma [13]. We performed radiomics analysis using HSR-CT data and achieved better predictive performance than conventional CT data. HSR-CT can provide higher image quality than conventional CT by improving spatial resolution (0.15 mm in plane, 0.20 mm through plane) and reducing artifacts due to undershoot [14]. A large matrix size maintained spatial resolution and improved image quality and assessment of lung disease [15]. The 1024 matrix size can provide detailed texture features compared to conventional CT. Tao et al. reported that the radiomics features with HRCT data of small pixel size achieved higher AUC in predicting the invasiveness of GGNs than larger pixel size [16]. The accuracy of radiomics features with HSR-CT data may improve the accuracy of predicting solid and micropapillary components.
The radiomics results significantly improved the specificity and, not significantly, the readers' AUC value. Visual assessment seems to have a tendency to predict that nodules have micropapillary or solid component, because many lesions had findings suggestive of micropapillary or solid components, such as solid portion or spiculation. However, nodules consisting mainly of solid components can also be those without micropapillary or solid portions. Therefore, there are many false positives in visual assessment, and radiomics results can significantly reduce them (Fig. 4).
There is a part-solid nodule in the left lower lobe. It is a lesion with a large proportion of solid component and it seems to be suggestive of a solid or micropapillary component. However, it has some degree of GGO and it is difficult to predict visually with certainty. Pathologically, it was a nodule without solid or micropapillary component. The radiologists predicted that it had a solid or micropapillary component. By referring to the radiomics results, they were able to predict correctly. The radiomics result entropy 3.36, coefficient of variance: − 0.817
On the other hand, it may be a problem that the criteria for whether to use the radiomics results vary from one radiologist to another (Fig. 5). For example, it may be difficult to use the radiomics results if they do not agree with the visual assessment, even though the performance of the radiomics method is good (Fig. 6). That’s because the basis for radiomics predictions is unclear. It is likely to be difficult to prove, and it may be necessary to set standards for whether to use radiomics results.
The three nodules with micropapillary and solid component: The radiomics method of this study predicted that they did not have micropapillary and solid component. The radiologists correctly predicted them without the radiomics results. After referring to the radiomics results, one did not change her prediction: it was correct, and the other did change it: it was incorrect. Thus, the decision to use radiomics results sometimes differs between radiologists. a A part-solid nodule of the left upper lobe with a small percentage of ground-glass opacity and spiculation. (The radiomics results: entropy 3.25, coefficient of variance − 1.34). b A solid nodule of the left upper lobe with almost no spiculation. (The radiomics results: entropy 3.19, coefficient of variance − 1.55). c A part-solid nodule of the right lower lobe, composed mainly of solid components and with spiculation. (The radiomics results: entropy 3.36, coefficient of variance − 0.877)
a There is a solid nodule in the right middle lobe. It has solid component. (The radiomics results: entropy 3.43, coefficient of variance − 1.93) The radiologists correctly predicted with and without the radiomics results. bThere is a solid nodule in the left lower lobe. It is a nodule without solid or micropapillary component. (The radiomics results: entropy 3.13, coefficient of variance − 0.918) The radiologists predicted wrongly without the radiomics results. After referring to the radiomics results, one did not change her prediction: it was incorrect, and the other did change it: it was correct. The two nodules are both visually solid and there appears to be little difference in appearance between them. However, a is a nodule with solid components, whereas b is without solid or micropapillary components. Although the radiomics method predicted correctly, it may be difficult to decide whether to accept the results because both are visually suspected to be nodules with solid or micropapillary components
Lung adenocarcinomas are usually heterogeneous with multiple subtypes. Thus, extracted radiomics values do not necessarily reflect the characteristics of each subtype itself, but those of the entire lesion with multiple subtypes. To reduce the influence of mixed subtypes, Li-Wei C et al. performed radiomics analysis with “near-pure” histopathological subtypes (a predominant subtype > 70% of tumor volume) and achieved high sensitivity and moderate specificity [17].
In this study, higher entropy and coefficient of variation were significant indicators to predict solid and micropapillary components in lung invasive adenocarcinoma. According to some previous studies, higher histogram entropy was associated with worse outcome. This may reflect that micropapillary and solid components have poor prognosis [18, 19]. Entropy and coefficient of variation are first-order statistics that describe the distribution of voxel intensities within the image region. Entropy describes the uncertainty or randomness of the image values. The coefficient of variation is the ratio of the standard deviation to the mean, and the higher the coefficient of variation, the greater the dispersion. This indicates that tumors with solid and micropapillary components have heterogeneity, and even invisible heterogeneity can be identified with the radiomics approach. In this study, detailed radiomics was performed using HSR-CT data with 1024 matrix, but the results were based on 2D analysis. The pathological tumor heterogeneity is complex, and further analysis is needed to develop dedicated software for three-dimensional (3D) texture analysis on HSR-CT with 1024 matrix, and to investigate the relationship between micropapillary and solid component and 3D texture analysis using a large cohort in the future.
Our study has several limitations. First, this study was based on 2D texture as described above. 3D texture analysis is desirable to analyze the complex heterogeneity of tumors. However, we analyzed not a whole lesion of the tumors but cross-sectional CT images where the solid portion is the maximum because currently there is no 3D texture analysis software for data on HSR-CT with large matrix size. Second, this study was a retrospective study at a single institution, and the number of cases with micropapillary was small. Because of the small data in this study, ten-fold cross-validation was performed for the evaluations. Further studies with larger cohorts including micropapillary and solid patterns are needed to confirm our results. Third, we used only a single texture analysis software. Texture features are known to be significantly affected by the CT acquisition protocol and pre-processing methods [20]. The way of segmentation and selection of cross-sections may vary the radiomic features. In this study, there was no influence of the selection of the cross-sections, because the two radiologists segmented the same cross-sectional CT images that were selected in advance.
In conclusion, two texture features (coefficient variation and entropy) from HSR-CT with 1024 matrix were significant indicators to predict solid and micropapillary components in lung invasive adenocarcinoma. This may be useful to predict the prognosis of each patient and determine a more appropriate treatment plan. However, since different radiologists have different criteria for using radiomics results, it may be necessary to build an explanatory radiomics prediction model and establish criteria for our future use of radiomics results.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries CA Cancer. J Clin. 2021;71(3):209–49.
William DT, Elisabeth B, Noguchi M, Andrew GN, Kim RG, Yatabe Y, et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol. 2011;6(2):244–85.
Cha MJ, Lee HY, Lee KS, Jeong JY, Han J, Shim YM, et al. Micropapillary and solid subtypes of invasive lung adenocarcinoma: clinical predictors of histopathology and outcome. J Thorac Cardiovasc Surg. 2014;147:921–8.
Zhao ZR, To KF, Mok TK, Ng CS. Is there significance in identification of non-predominant micropapillary or solid components in early-stage lung adenocarcinoma? Interact Cardiovasc Thorac Surg. 2017;24(1):121–5.
Lee G, Lee HY, Jeong JY, Han J, Cha MJ, Lee KS, et al. Clinical impact of minimal micropapillary pattern in invasive lung adenocarcinoma: prognostic significance and survival outcomes. Am J Surg Pathol. 2015;39:660–6.
Nitadori J, Bograd AJ, Kadota K, Sima CS, Rizk NP, Morales EA, et al. Impact of micropapillary histologic subtype in selecting limited resection vs lobectomy for lung adenocarcinoma of 2cm or smaller. J Natl Cancer Inst. 2013;105:1212–20.
Mathieu L, Michael P, Thomas M, Philipp AS, Albrecht S, Hans-Ulrich K, et al. Correlation of radio- and histomorphological pattern of pulmonary adenocarcinoma. Eur Respir J. 2013;41(4):943–51.
Yanagawa M, Tsubamoto M, Satoh Y, Hata A, Miyata T, Yoshida Y, et al. Lung adenocarcinoma at CT with 0.25-mm section thickness and a 2048 matrix: high-spatial-resolution imaging for predicting invasiveness. Radiology. 2020;297(2):462–71.
Ryan W, Anand D. Radiomics of pulmonary nodules and lung cancer. Transl Lung Cancer Res. 2017;6:86–91.
Song SH, Park H, Lee G, Lee HY, Sohn I, Kim HS, et al. Imaging phenotyping using radiomics to predict micropapillary pattern within lung adenocarcinoma. J Thorac Oncol. 2017;12(4):624–32.
He B, Yongxiang S, Lili W, et al. A machine learning-based prediction of the micropapillary/solid growth pattern in invasive lung adenocarcinoma with radiomics. Transl Lung Cancer Res. 2021;10(2):955–64.
Xu Y, Ji W, Hou L, Lin S, Shi Y, Zhou C, et al. Enhanced CT-based radiomics to predict micropapillary pattern within lung invasive adenocarcinoma. Front Oncol. 2021;11:704994.
Park S, Lee SM, Noh HN, Hwang HJ, Kim S, Do KH, et al. Differentiation of predominant subtypes of lung adenocarcinoma using a quantitative radiomics approach on CT. Eur Radiol. 2020;30(9):4883–92.
Yanagawa M, Hata A, Honda O, Kikuchi N, Miyata T, Uranishi A, et al. Subjective and objective comparisons of image quality between ultra-high-resolution CT and conventional area detector CT in phantoms and cadaveric human lungs. Eur Radiol. 2018;28:5060–8.
Hata A, Yanagawa M, Honda O, Kikuchi N, Miyata T, Tsukagoshi S, et al. Effect of matrix size on the image quality of ultra-high-resolution CT of the Lung: comparison of 512 × 512, 1024 × 1024, and 2048 × 2048. Acad Radiol. 2018;25(7):869–76.
Tao G, Yin L, Shi D, Ye J, Lu Z, Zhou Z, et al. Dependence of radiomic features on pixel size affects the diagnostic performance of radiomic signature for the invasiveness of pulmonary ground-glass nodule. Br J Radiol. 2021;94(1118):20200089.
Chen LW, Yang SM, Wang HJ, Chen YC, Lin MW, Hsieh MS, et al. Prediction of micropapillary and solid pattern in lung adenocarcinoma using radiomic values extracted from near-pure histopathological subtypes. Eur Radiol. 2021;31:5127–38.
Bak SH, Park H, Sohn I, Lee SH, Ahn MJ, Lee HY. Prognostic impact of longitudinal monitoring of radiomic features in patients with advanced non-small cell lung cancer. Sci Rep. 2019;9:8730.
Tu SJ, Wang CW, Pan KT, Wu YC, Wu CT. Localized thin-section CT with radiomics feature extraction and machine learning to classify early-detected pulmonary nodules from lung cancer screening. Phys Med Biol. 2018;63(6):065005.
Au RC, Tan WC, Bourbeau J, Hogg JC, Kirby M, et al. Impact of image pre-processing methods on computed tomography radiomics features in chronic obstructive pulmonary disease. Phys Med Biol. 2021;66:245015.
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This retrospective study was approved by our institution's internal ethics review board (No.16321-5).
Informed consent
Informed consent was waived for review of patient records and images.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Ninomiya, K., Yanagawa, M., Tsubamoto, M. et al. Prediction of solid and micropapillary components in lung invasive adenocarcinoma: radiomics analysis from high-spatial-resolution CT data with 1024 matrix. Jpn J Radiol (2024). https://doi.org/10.1007/s11604-024-01534-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11604-024-01534-2