Summary
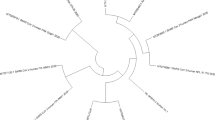
Severe acute respiratory syndrome Coronavirus 2 (SARS-CoV-2) with unknown origin spread rapidly to 222 countries, areas or territories. To investigate the genomic evolution and variation in the early phase of COVID-19 pandemic in Guangdong, 60 specimens of SARS-CoV-2 were used to perform whole genome sequencing, and genomics, amino acid variation and Spike protein structure modeling analyses. Phylogenetic analysis suggested that the early variation in the SARS-CoV-2 genome was still intra-species, with no evolution to other coronaviruses. There were one to seven nucleotide variations (SNVs) in each genome and all SNVs were distributed in various fragments of the genome. The Spike protein bound with human receptor, an amino acid salt bridge and a potential furin cleavage site were found in the SARS-CoV-2 using molecular modeling. Our study clarified the characteristics of SARS-CoV-2 genomic evolution, variation and Spike protein structure in the early phase of local cases in Guangdong, which provided reference for generating prevention and control strategies and tracing the source of new outbreaks.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Su S, Wong G, Shi W, et al. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol, 2016,24(6):490–502
Ksiazek T G, Erdman D, Goldsmith C S, et al. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med, 2003,348(20):1953–1966
Kuiken T, Fouchier R A, Schutten M, et al. Newly discovered coronavirus as the primary cause of severe acute respiratory syndrome. Lancet, 2003,362(9380): 263–270
Zaki AM, van Boheemen S, Bestebroer TM, et al. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med, 2012,367(19):1814–1820
WHO. Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July 2003. Dec 31, 2003. https://www.who.int/csr/sars/country/table2004_04_21/en/ (accessed Jan 19, 2020).
WHO. Middle East respiratory syndrome coronavirus (MERS-CoV). November, 2019. http://www.who.int/emergencies/mers-cov/en/ (accessed Jan 19, 2020).
Lu R, Zhao X, Li J, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet, 2020, 395(10224):565–574
Bankevich A, Nurk S, Antipov D, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol, 2012,19(5):455–477
Elbe S, Buckland-Merrett G. Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob Chall, 2017,1(1):33–46
Kumar S, Stecher G, Li M, et al. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol, 2018,35(6):1547–1549
Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res, 1988,16(22):10881–10890
Biasini M, Bienert S, Waterhouse A, et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res, 2014,42(Web Server issue):W252–W258
Song W, Gui M, Wang X, et al. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog, 2018,14(8):e1007236
Zhu N, Zhang D, Wang W, et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med, 2020,382(8):727–733
Wrapp D, Wang N, Corbett K S, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science, 2020,367(6483):1260–1263
Alam N, Goldstein O, Xia B, et al. High-resolution global peptide-protein docking using fragments-based PIPER-FlexPepDock. PLoS Comput Biol, 2017,13(12): e1005905
Song W, Gui M, Wang X, et al. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog, 2018,14(8):e1007236
Pillay T S. Gene of the month: the 2019-nCoV/SARS-CoV-2 novel coronavirus spike protein. J Clin Pathol, 2020,73(7):366–369
Follis KE, York J, Nunberg JH. Furin cleavage of the SARS coronavirus spike glycoprotein enhances cell-cell fusion but does not affect virion entry. Virology, 2006,350(2):358–369
Belouzard S, Chu VC, Whittaker GR. Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites. Proc Natl Acad Sci U S A, 2009,106(14):5871–5876
Author information
Authors and Affiliations
Corresponding author
Additional information
Conflict of Interest Statement
The authors declare that there is no conflict of interest with any financial organization or corporation or individual that can inappropriately influence this work.
Rights and permissions
About this article
Cite this article
Li, Bs., Li, Zc., Hu, Y. et al. Genomic Evolution and Variation of SARS-CoV-2 in the Early Phase of COVID-19 Pandemic in Guangdong Province, China. CURR MED SCI 41, 228–235 (2021). https://doi.org/10.1007/s11596-021-2340-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-021-2340-3