Abstract
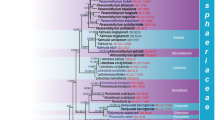
Seven species of Podoscypha (including four new species) and two species of Cymatoderma collected from the Guangxi Zhuang Autonomous Region in Southwest China were studied. Phylogenetic analyses of both the ITS and ITS + nLSU datasets revealed that species of Cymatoderma were included in two different groups and species of Podocypha were nested into a large clade including several groups. By combining morphological characteristics and phylogenetic analyses, four new species, Podoscypha lactea, P. densidisca, P. guangxiensis and P. infundibula, were described, and a key to Podoscypha species from China was also provided.












Similar content being viewed by others
Availability of data and material
The sequence data generated in this study are deposited in NCBI GenBank and the specimens are preserved at the Herbarium of Edible Fungi Institute of Guangxi University (GXU).
References
Bau T, Kang GP, Fan YG, Wang Y, Liang H (2011) Checklist of macrofungi collected from different forests in Changbai Mountain (IV): coniferous and broad-leaved mixed forest. J Fungal Res 1:21–36. https://doi.org/10.13341/j.jfr.2011.01.005
Binder M, Justo A, Riley R, Salamov A, Lopez-Giraldez F, Sjökvist E, Copeland A, Foster B, Sun H, Larsson E, Larsson KH, Townsend J, Grigoriev IV, Hibbett DS (2013) Phylogenetic and phylogenomic overview of the Polyporales. Mycologia 105(6):1350–73. https://doi.org/10.3852/13-003
Boidin J, Lanquetin P (1973) Podoscypha involuta (Klotzsch) Imaz, estuneespèce composite (Basidiomycetes, Podoscyphaceae). Persoonia 7(2):141–150
Dai YC (2010) Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 45(1):131–343
Dai YC (2011) A revised checklist of corticioid and hydnoid fungi in China for 2010. Mycoscience 52:69–79. https://doi.org/10.1007/s10267-010-0068-1
Dai YC, Yang ZL, Cui BK, Wu G, Yuan HS, Zhou LW, He SH, Ge ZW, Wu F, Wei YL, Yuan Y, Si J (2021) Diversity and systematics of the important macrofungi in Chinese forests. Mycosystema 40:770–805
Douanla-Meli C, Langer E (2004) A taxonomic study of the family Podoscyphaceae (Basidiomycetes), new species and new records in Cameroon. Mycotaxon 90:323–335
Drechsler-Santos ER, Gibertoni TB, Cavalcanti MAdeQ, (2007) Podoscypha aculeata, a new record for the neotropics. Mycotaxon 101:69–72
Guo ZT (1987) Stereaceae in China (II). BBR 7(2):53–79
Huang FC, Liu B, Wu H, Qin PH, Li JF (2018) Podoscypha (Polyporales, Basidiomycota) species from Nonggang National Nature Reserve of Guangxi. J Fungal Res 16(04):205–209. https://doi.org/10.13341/j.jfr.2014.1188
Li TH, Dang WQ, Song B, Cao H, Guan SM, Lin WF, Chen XB, Liang SH (2002) Macrofungi from Diaoluoshan nature reserve in Hainan province. Trop Forest 3:37–44
Liu S, Xu TM, Song CG, Zhao CL, Wu DM, Cui BK (2022) Species diversity, molecular phylogeny and ecological habits of Cyanosporus (Polyporales, Basidiomycota) with an emphasis on Chinese collections. MycoKeys 86:19–46. https://doi.org/10.3897/mycokeys.86.78305
Liu S, Chen YY, Sun YF, He XL, Song CG, Si J, Liu DM, Gates G, Cui BK (2023) Systematic classification and phylogenetic relationships of the brown-rot fungi within the Polyporales. Fungal Divers 118:1–94. https://doi.org/10.1007/s13225-022-00511-2
Mao WL, Wu YD, Liu HG, Yuan Y, Dai YC (2023) A contribution to Porogramme (Polyporaceae, Agaricomycetes) and related genera. IMA Fungus 14:5. https://doi.org/10.1186/s43008-023-00110-z
Miettinen O, Larsson E, Sjokvist E, Larsson KH (2012) Comprehensive taxon sampling reveals unaccounted diversity and morphological plasticity in a group of dimitic polypores (Polyporales, Basidiomycota). Cladistics 28:251–270. https://doi.org/10.1111/j.1096-0031.2011.00380.x
Munsell Color (Firm) (2019) Munsell Soil Color Charts: with Genuine Munsell Color Chips. 2009 year Revised, 2019 Production. Produced by Munsell color x-rite, Grand Rapids, MI: 49512, Munsell Color
Nylander JAA (2004) MrModeltest v2. Evolutionary biology center. Uppsala University, Uppsala
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818. https://doi.org/10.1093/bioinformatics/14.9.817
Qi LL, Li LY, Wu XJ, Zhao CG, Hu YQ, Liao GF, Liang N (2022) Study on species diversity of macrofungi under Castanopsis forest in Guangxi. Edible Fungi 44(03):12–15+21
Rehill PS, Bakshi BK (1966) Studies on Indian Thelephoraceae III. The genus Stereum. Indian Forest Bull Dehra Dun 250:1–20
Reid DA (1965) A monograph of the stipitate stereoid fungi. Beihefte zur Nova Hedwigia 18:1–382
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Ryvarden L (2023) Synopsis Fungorum 47 - Stereoid fungi - a world synopsis. Fungiflora Oslo 47:1–157
Si J, Zhang YZ, Liang JQ, Li HJ (2023) Morphology and phylogeny identify two new species and one new subspecies of Podoscypha from Yunnan Province, Southwest China. Front Microbiol 14:1151365. https://doi.org/10.3389/fmicb.2023.1151365
Sjökvist E, Larsson E, Eberhardt U, Ryvarden L, Larsson KH (2012) Stipitate stereoid basidiocarps have evolved multiple times. Mycologia 104:1046–1055. https://doi.org/10.3852/11-174
Spirin V, Vlasák J, Niemelä T, Miettinen O (2022) Studies in Spongipellis sensu stricto (Polyporales, Basidiomycota). Lilloa 59:341–358. https://doi.org/10.30550/j.lil/2022.59.S/2022.09
Stöver BC, Müller KF (2010) TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinformatics 11:7
Swofford DL (2002) PAUP*, phylogenetic analysis using parsimony, ver. 4.0b10. Sinauer Associates, Sunderland
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
Vu D, Groenewald M, de Vries M, Gehrmann T, Stielow B, Eberhardt U, Al-Hatmi A, Groenewald JZ, Cardinali G, Houbraken J, Boekhout T, Crous PW, Robert V, Verkley GJM (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud Mycol 92:135–154. https://doi.org/10.1016/j.simyco.2018.05.001
Wang YR, Wu YD, Vlasák J, Yuan Y, Dai YC (2021) Phylogenetic analysis demonstrating four new species in Megasporoporia sensu lato (Polyporales, Basidiomycota). Mycosphere 12(1):1012–1037. https://doi.org/10.5943/mycosphere/12/1/11
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic Press, San Diego
Wu SH (2003) Lignicolous homobasidiomycetes newly recorded from Taiwan. Mycotaxon 88:373–376
Wu YX, Shen S, Zhao CL (2019) Podoscypha yunnanensis sp. nov. (Polyporales, Basidiomycota) evidenced by morphological characters and phylogenetic analyses. Phytotaxa 387(3):210–218. https://doi.org/10.11646/phytotaxa.387.3.2
Zhao CL, Cui BK (2012) A new species of Perenniporia (Polyporales, Basidiomycota) described from southern China based on morphological and molecular characters. Mycol Progress 11:555–560. https://doi.org/10.1007/s11557-011-0770-1
Funding
Funding was provided to Fu-Chang Huang by the National Natural Science Foundation of China (Project No. 31960006).
Author information
Authors and Affiliations
Contributions
Specimens collecting: Fu-Chang Huang, Jian-Tian Lin, Hai-Fu Zheng, Yun-Long Liang, Miao-Miao Li, Cheng-Pei Ding.
DNA extraction, PCR: Jian-Tian Lin, Hai-Fu Zheng, Yun-Long Liang, Miao-Miao Li.
Morphological studies: Fu-Chang Huang.
The study conception, design, writing, review and editing: Fu-Chang Huang.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Yu-Cheng Dai.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, FC., Lin, JT., Zheng, HF. et al. Molecular phylogeny of Podoscypha (Basidiomycota, Podoscyphaceae) with descriptions of four new species, inferred from nuclear rDNA sequences. Mycol Progress 23, 31 (2024). https://doi.org/10.1007/s11557-024-01969-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-024-01969-x