Abstract
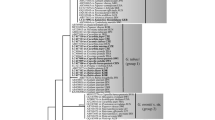
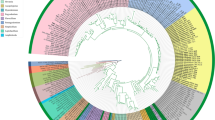
The family Cucurbitariaceae is rich in species diversity and has a wide host range and geographic distribution. In this study, we identified 12 Cucurbitariaceae isolates which were obtained from disease symptoms in two forest trees in Khuzestan province, Iran. In addition, this family is reassessed using phylogenetic analyses based on DNA sequences from five nuclear regions (ITS, LSU, TUB2, TEF1α, and RPB2). The phylogenetic analyses showed that the present isolates represent one new genus, Nothocucurbitaria, and three new species, Allocucurbitaria galinsogisoli, Nothocucurbitaria izehica, and Parafenestella quercicola, which are described and illustrated. Furthermore, the genus Allocucurbitaria is emended to accommodate Seltsamia ulmi that grouped with the type species of Allocucurbitaria. Parafenestella pittospori and A. prunicola are recombined into the genera Neocucurbitaria and Nothocucurbitaria, respectively. Comparative analysis of single-locus trees revealed that the TUB2 and TEF1α can distinguish most genera and species in Cucurbitariaceae, while the ITS and LSU phylogenies show low resolution at both generic and species level. The best single-locus marker, RPB2, was able to distinguish all generic and most species lineages in Cucurbitariaceae.




Similar content being viewed by others
Data availability
New sequences generated in the current study are deposited in NCBI GenBank and new species in MycoBank. Combined DNA dataset used for phylogenetic analyses is included in a supplementary file.
References
Ahmadpour SA, Mehrabi-Koushki M, Farokhinejad R, Asgari B (2022) New species of the family Didymellaceae in Iran. Mycol Prog 2:119–133
Alves A, Crous PW, Correia A, Phillips AJL (2008) Morphological and molecular data reveal cryptic speciation in Lasiodiplodia theobromae. Fungal Divers 28:1–13
Artand S, Mehrabi-Koushki M, Tabein S, Hyde KD, Jayawardena RS (2022) Revision of the Microsphaeropsis complex with addition of four new Paramicrosphaeropsis L.W.Hou, L. Cai & Crous species from Zagrosian forest trees in Iran. Cryptogam Mycol 43:159–175
Aveskamp MM, de Gruyter J, Woudenberg JHC, Verkley GJM, Crous PW (2010) Highlights of the Didymellaceae: a polyphasic approach to characterise Phoma and related pleosporalean genera. Stud Mycol 65:1–60
Barr ME (1987) Prodromus to class Loculoascomycetes. University of Massachusetts, Amherst, Massachusetts
Barr ME (1990) Some dictyosporous genera and species of Pleosporales in North America. Mem New York Bot Gard 62:1–92
Bubak F, Kabat JE (1912) Mykologische Beiträge. VII. Hedwigia 52:340–363
Chen Q, Jiang JR, Zhang GZ, Cai L, Crous PW (2015) Resolving the Phoma enigma. Stud Mycol 82:137–217
Crous PW, Schumacher RK, Akulov A, Thangavel R, Hernández-Restrepo M, Carnegie AJ, Cheewangkoon R, Wingfield MJ, Summerell BA, Quaedvlieg W, Coutinho TA, Roux J, Wood AR, Giraldo A, Groenewald JZ (2019) New and interesting fungi. 2. Fungal Syst Evol 3:57–134
Crous PW, Wingfield MJ, Lombard L, Roets F, Swart WJ, Alvarado P, Carnegie AJ, Moreno G, Luangsaard J, Thangavel R, Alexandrova AV, Baseia IG, Bellanger JM, Bessette AE, Bessette AR, De la Peña-Lastra S, García D, Gené J, Pham THG, Groenewald JZ (2019) Fungal Planet description sheets: 951–1041. Persoonia 43:223
Crous PW, Lombard L, Sandoval-Denis M, Seifert KA, Schroers HJ, Chaverri P, Gené J, Guarro J, Hirooka Y, Bensch K, Kema GHJ, Lamprecht SC, Cai L, Rossman AY, Stadler M, Summerbell RC, Taylor JW, Ploch S, Visagie CM, Thines M (2021) Fusarium: more than a node or a foot-shaped basal cell. Stud Mycol 98:100116
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772–772
de Gruyter J (2012) Revised taxonomy of Phoma and allied genera. Ph.D. dissertation, Wageningen University, The Netherlands
de Gruyter J, Woudenberg JH, Aveskamp MM, Verkley GJM, Groenewald JZ, Crous PW (2010) Systematic reappraisal of species in Phoma section Paraphoma, Pyrenochaeta and Pleurophoma. Mycologia 102:1066–1081
Doilom M, Liu JK, Jaklitsch WM, Ariyawansa H, Wijayawardene NN, Chukeatirote E, Zhang M, McKenzie EH, Geml J, Voglmayr H, Hyde KD (2013) An outline of the family Cucurbitariaceae. Sydowia 65:167–192
Edler D, Klein J, Antonelli A, Silvestro D (2021) raxmlGUI 2.0: a graphical interface and toolkit for phylogenetic analyses using RAxML. Methods Ecol Evol 12:373–377
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hill R, Levicky Q, Pitsillides F, Junnonen A, Arrigoni E, Bonnin JM, Kermode A, Mian S, Leitch IJ, Buddie AG, Buggs RJ, Gaya E (2023) Tapping culture collections for fungal endophytes: first genome assemblies for three genera and five species in the Ascomycota. Genome Biol Evol 15:evad038
Hong SM, Das K, Lim SK, Suh SJ, Lee SY, Jung HY (2023) Neocucurbitaria chlamydospora sp. nov.: a novel species of the family Cucurbitariaceae isolated from a stink bug in Korea. Mycobiology 51:115–121
Ilyukhin E, Markovskaja S, Elgorban AM, Al-Rejaie SS, Maharachchikumbura SS (2022) Genomic characteristics and comparative genomics analysis of Parafenestella ontariensis sp. nov. J Fungi 8:732
Jaklitsch WM, Voglmayr H (2020) Fenestelloid clades of the Cucurbitariaceae. Persoonia 44:1–40
Jaklitsch WM, Olariaga I, Voglmayr H (2016) Teichospora and the Teichosporaceae. Mycol Prog 15:1–20
Jaklitsch WM, Checa J, Blanco MN, Olariaga I, Tello S, Voglmayr H (2018) A preliminary account of the Cucurbitariaceae. Stud Mycol 90:71–118
Jankowiak R, Stępniewska H, Bilański P, Taerum SJ (2022) Fungi as potential factors limiting natural regeneration of pedunculate oak (Quercus robur) in mixed-species forest stands in Poland. Plant Pathol 71:805–817
Kornerup A, Wanscher JH (1967) Methuen handbook of colour, 2nd edn. Methuen & Co., London, England
Liu YJ, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Mol Biol Evol 16:1799–1808
Magaña-Dueñas V, Stchigel AM, Cano-Lira JF (2021) New coelomycetous fungi from freshwater in Spain. J. Fungi 7:368
Mehrabi-Koushki M, Khodadadi-Pourarpanahi S, Jounbozorgi S (2018) Fungal endophytes associated with some thermotolerant plants in salt-stress ecosystem. Mikol Fitopatol 52:187–195
Mehrabi-Koushki M, Artand S, Ahmadpour SA (2021) Botryosphaeria dothidea causes stem canker on Fatsia japonica in Iran. Australas Plant Dis Notes 16:1–4
O’Donnell K (1993) Fusarium and ITS near relatives. In: Reynolds DR, Taylor JW (eds) The fungal holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematics. CAB International, Wallingford, pp 225–233
O’Donnell K, Kistler HC, Cigelnik E, Ploetz RC (1998) Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proc Natl Acad Sci U S A 95:2044–2049
O’Donnell K, Cigelnik E (1997) Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 7:103–116
Punithalingam E, English MP (1975) Pyrenochaeta unguis-hominis sp. nov. on human toe-nails. Trans Br Mycol Soc 64:539–541
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Safi A, Mehrabi-Koushki M, Farokhinejad R (2021) Plenodomus dezfulensis sp. nov. causing leaf spot of Rapeseed in Iran. Phytotaxa 47:430–441
Su W, Xu R, Bhunjun CS, Tian S, Dai Y, Li Y, Phukhamsakda C (2022) Diversity of Ascomycota in Jilin: introducing novel woody litter taxa in Cucurbitariaceae. J Fungi 8:905
Sung GH, Sung JM, Hywel-Jones NL, Spatafora JW (2007) Amulti-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol 44:1204–1223
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Valenzuela-Lopez N, Sutton DA, Cano-Lira JF et al (2016) Coelomycetous fungi in the clinical setting: morphological convergence and cryptic diversity. J Clin Microbiol 55:552–567
Valenzuela-Lopez N, Sutton DA, Cano-Lira JF, Paredes K, Wiederhold N, Guarro J, Stchigel AM (2018) Coelomycetous Dothideomycetes with emphasis on the families Cucurbitariaceae and Didymellaceae. Stud Mycol 90:1–69
Verkley GJ, Gené J, Guarro J, Pérez-Santonja JJ, Rodríguez AE, Colom MF, Alio JL, Ferrer C (2010) Pyrenochaeta keratinophila sp. nov., isolated from an ocular infection in Spain. Rev Iberoam Micol 27:22–24
von Arx JA, Muller E (1950) A re-evaluation of the bitunicate ascomycetes with keys to families and genera. Trans Br Mycol Soc 9:1–159
Wanasinghe DN, Phookamsak R, Jeewon R, Li WJ, Hyde KD, Jones EG, Camporesi E, Promputtha I (2017) A family level rDNAbased phylogeny of Cucurbitariaceae and Fenestellaceae with descriptions of new Fenestella species and Neocucurbitaria gen. nov. Mycosphere 8:397–414
White TJ, Bruns T, Lee SJWT, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Wijayawardene NN, Hyde KD, Al-Ani LK, Tedersoo L, Haelewaters D, Rajeshkumar KC, Zhao RL, Aptroot A, Leontyev DV, Saxena RK, Tokarev YS, Dai DQ, Letcher PM, Stephenson SL, Ertz D, Lumbsch HT, Kukwa M, Issi IV, Madrid H, Suija A (2020) Outline of fungi and fungus-like taxa. Mycosphere Online: Fungal Biol 11:1060–1456
Winter HG (1885) Pilze. II. Abtheilung. Ascomyceten: Gymnoasceen und Pyrenomyceten. In: 2nd edn. In: Rabenhorst‘s Kryptogamen-Flora von Deutschland, Oesterreich und der Schweiz (Rabenhorst GL, ed), Kummer, Leipzig. 1(2):65–528
Woudenberg JHC, Aveskamp MM, De Gruyter J, Spiers AG, Crous PW (2009) Multiple Didymella teleomorphs are linked to the Phoma clematidina morphotype. Persoonia 22:56–62
Zhang TY, Wu YY, Zhang MY, Cheng J, Dube B, Yu HJ, Zhang YX (2019) New antimicrobial compounds produced by Seltsamia galinsogisoli sp. nov., isolated from Galinsoga parviflora as potential inhibitors of FtsZ. Sci Rep 9:8319
Funding
This work was financially supported by grant (SCU.AP1402.294) from the Research Council of Shahid Chamran University of Ahvaz.
Author information
Authors and Affiliations
Contributions
Payam Eisvand carried out sample preparation, fungal isolation and purification, morphometric and morphological determination, DNA isolation, PCR amplification, and contribution to describe the novel species. Mehdi Mehrabi-Koushki carried out the design and implementation of the research, DNA and phylogenetic analyses, and the writing of the manuscript. Pedro W. Crous contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
Not applicable.
Consent for publication
Informed consent was obtained from all individual participants included in the study.
Competing interests
The authors declare no competing interests.
Additional information
Section Editor: Roland Kirschner
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Eisvand, P., Mehrabi-Koushki, M. & Crous, P.W. A revision of the family Cucurbitariaceae with additional new taxa from forest trees in Iran. Mycol Progress 23, 14 (2024). https://doi.org/10.1007/s11557-024-01953-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-024-01953-5