Abstract
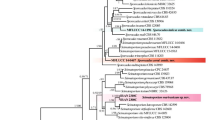
A new species of Ulocladium was discovered from diseased leaves of Lycopersicon esculentum and Duchesnea indica from Hunan Province of China. Morphologically, this species is very close to U. consortiale, U. cucurbitae, and U. subcucurbitae in producing narrow ellipsoid conidia at 1–3 days, but the conidial size range of this species at this stage could distinguish it from three well-known species. It also exhibits the multiplex conidium morphology at the different growth-stages (1–3 days and 4–7 days). The results of maximum parsimony (MP) and neighbor-joining (NJ) phylogenetic analyses of combined glyceraldehyde-3-phosphate dehydrogenase (gpd) gene and Alternaria alternata major allergen (Alt a 1) genes show that U. solani and U. subcucurbitae cluster in a unique and separate subclade with no clear affinities to a specific sistergroup, and demonstrate that the Ulocladium species group is monphyletic, but two clades of this section are recognized. Morphological features of this new species, the sequences of the Alt a 1 and gpd gene regions, and its comparison with related species in this genus are discussed.



Similar content being viewed by others
References
Barnes CS, Pacheco F, Landuyt J, Rosenthal D, Hu F, Portnoy J (1996) Production of a recombinant protein from Alternaria containing the reported N-terminal of the Alt a 1 allergen. In: Sehon A, Kraft D, Hay Glass KT (eds) Advances in experimental medicine and biology, vol. 409. Plenum Press, New York, pp 197–203
Berbee ML, Pirseyedi M, Hubbard S (1999) Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 91:964–977. doi:10.2307/3761627
Câmara MPS, O’Neill NR, van Berkum P (2002) Phylogeny of Stemphylium spp. based on ITS and glyceraldehydes-3-phosphate dehydrogenase gene sequences. Mycologia 94:660–672. doi:10.2307/3761717
Cramer RA, Lawrence CB (2003) Cloning of a gene encoding an Alt a 1 isoallergen differentially expressed by the necrotrophic fungus Alternaria brassicicola during Arabidopsis infection. Appl Environ Microbiol 69:2361–2364. doi:10.1128/AEM.69.4.2361–2364.2003
Cramer RA, Lawrence CB (2004) Identification of Alternaria brassicicola genes expressed in plant during pathogenesis of Arabidopsis thaliana. Fungal Genet Biol 41:115–128. doi:10.1016/j.fgb.2003.10.009
de Hoog GS, Horré R (2002) Molecular taxonomy of the Alternaria and Ulocladium species from humans and their identification in the routine laboratory. Mycoses 45:259–276. doi:10.1046/j.1439–0507.2002.00747.x
De Vouge MW, Thanker AM, Curran IHA, Zhang L, Muradia G, Rode H, Vijay HM (1996) Isolation and expression of a cDNA encoding on Alternaria alternata Alt a 1 subunit. Int Arch Allergy Immunol 111:385–395
Elmer PAG, Köhl J (1998) The survival and saprophytic competitive ability of the Botrytis spp. antagonist Ulocladium atrum in lily canopies. Eur J Plant Pathol 104(5):435–477. doi:10.1023/A:1008646418991
Farris JS, Kallersjo M, Kluge AG, Bult C (1995) Testing significance of incongruence. Cladistics 10:315–319. doi:10.1111/j.1096–0031.1994.tb00181.x
Grishkan I, Beharav A, Kirzhner V, Nevo E (2007) Adaptive spatiotemporal distribution of soil microfungi in ‘Evolution Canyon’ II, Nahal Shaharut, extreme southern Negev Desert, Israel. Biol J Linn Soc Lond 90:263–277. doi:10.1111/j.1095–8312.2007.00722.x
Hong SG, Cramer RA, Lawrence CB, Pryor BM (2005) Alt a 1 allergen homologs from Alternaria and related taxa: analysis of phylogenetic content and secondary structure. Fungal Genet Biol 42:119–129. doi:10.1016/j.fgb.2004.10.009
Huelsenbeck JP, Bull JJ, Cunningham CW (1996) Combining data in phylogenetic analysis. Trends in Ecol Evol 11:152–158. doi:10.1016/0169–5347(96)10006–9
Köhl J, Molhoek WWL, Goossen-van de Geijn HM, Lombaers van der Plas CH (2003) Potential of Ulocladium atrum for biocontrol of onion leaf spot through suppression of sporulation of Botrytis spp. Biocontrol 48(3):349–359. doi:10.1023/A:1023614622179
Leach CM, Aragaki M (1970) Effect of temperature on conidium characteristics of Ulocladium chartarum and Stemphylium floridanum. Mycologia 62:1071–1076. doi:10.2307/3757622
Preuss CGT (1851) Die Pilze Deutschlands. Heft. 30. In Jacob Sturm’s Deutschlands Flora, Abt III:73–96
Pryor BM, Bigelow DM (2003) Molecular characterization of Embellisia and Nimbya species and their relationship to Alternaria, Ulocladium, and Stemphylium. Mycologia 95:1139–1152. doi:10.2307/3761916
Pryor BM, Gilbertson RL (2000) Molecular phylogenetic relationships amongst Alternaria species and related fungi based on analysis of nuclear ITS and mtSSU rDNA sequences. Mycol Res 104:1312–1321. doi:10.1017/S0953756200003002
Sáenz-de-Santamaría M, Postigo I, Gutierrez-Rodríguez A, Cardona G, Guisantes JA, Asturias J, Martínez J (2005) The major allergen of Alternaria alternata (Alt a 1) is expressed in other members of the Pleosporaceae family. Mycoses 49:91–95. doi:10.1111/j.1439–0507.2006.01195.x
Saparrat MCN, Arambarri AM, Balatti PA (2007) Growth response and extracellular enzyme activity of Ulocladium botrytis LPSC 813 cultured on carboxy-methycellulose under a pH range. Biol Fertil Soils 44:383–386. doi:10.1007/s00374–007–0217–7
Simmons EG (1967) Typification of Alternaria, Stemphylium, and Ulocladium. Mycologia 59:67–92. doi:10.2307/3756943
Simmons EG (1982) Alternaria themes and variations (11–13). Mycotaxon 14:44–57
Simmons EG (1990) Alternaria themes and variations. Mycotaxon 70:79–119
Simmons EG (1998) Multiplex conidium morphology in species of the Ulocladium atrum group. Can J Bot 76:1553–1559. doi:10.1139/cjb-76–9–1533
Simmons EG, Roberts RG (1993) Alternaria themes and variations (73). Mycotaxon 48:109–140
Smith TL (1989) Disparate evolution of yeasts and filamentous fungi indicated by phylogenetic analysis of glyceraldehydes-3-phosphate dehydrogenase genes. Proc Natl Acad Sci USA 86:7063–7066. doi:10.1073/pnas.86.18.7063
Swofford DL (2000) PAUP*: Phylogenetic analysis using parsimony (* and other methods). Version 4.0b10. Sinauer Associates, Sunderland, Mass.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustsal X windows interface: flexible strategies for multiple sequence alignment aided by quality 1 analysis tools. Nucleic Acids Res 24:4876–4882. doi:10.1093/nar/25.24.4876
Wang Y, Bruno LC, Zhang XG (2008) Two new species of Ulocladium from Southwest China. Mycologia 100(3):455–459. doi:10.3852/07–019R3
Vannini A, Vettraino AM (2000) Ulocladium chartarum as the causal agent of a leaf necrosis on Quercus pubescens. For Pathol 30:297–303. doi:10.1046/j.1439–0329.2000.00210.x
Xue F, Zhang XG (2007) Ulocladium capsicuma, a new species identified by morphological and molecular phylogenetic data. Sydowia 59(1):161–178
Zhang G, Berbee ML (2001) Pyrenophora phylogenetics inferred from ITS and glyceraldehydes-3-phosphate dehydrogenase gene sequences. Mycologia 93:1048–1063. doi:10.2307/3761667
Zhang XG, Zhang TY (2002) Studies on the genus Ulocladium Preuss from China. Mycosystema 21(1):25–26
Zitter TA, Hsu LW (1990) A leaf spot of cucumber caused by Ulocladium cucurbitae in New York. Plant Dis 74:824–827. doi:10.1094/PD-74–0824
Acknowledgments
The authors would like to thank Dr. Jeffrey Stone for help with the English text. This work was supported by the National Natural Science Foundation of China (No. 30570006). We also would like to acknowledge the herbarium of the Centraalbureau voor Schimmelcultures for providing some valuable Ulocladium isolates in this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, Y., Pei, YF., Zhang, K. et al. Molecular and morphological description of a new species of Ulocladium from Southern China. Mycol Progress 8, 207–214 (2009). https://doi.org/10.1007/s11557-009-0592-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-009-0592-6