Abstract
Purpose
Age-matched average 3D models facilitate both surgical planning and intraoperative guidance of cranial birth defects such as craniosynostosis. We aimed to develop an algorithm that accepts any number of CT scans as input and generates highly accurate, average models with minimal user input that are ready for 3D printing and clinical use.
Methods
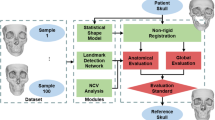
Using a compiled database of ‘normal’ pediatric computed tomography (CT) scans, we report Normscan, an open-source platform built in Python that allows users to generate normative models of CT scans through user-defined landmarks. We use the basion, nasion, and left and right porions as anatomical landmarks for initial correspondence and then register the models using the iterative closest points algorithm before downstream averaging.
Results
Normscan is fast and easy to use via our user interface and also creates highly accurate average models of any number of input models. Additionally, it is highly repeatable, with coefficients of variance for the surface area and volume of the average model being less than 3% across ten independent trials. Average models can then be 3D printed and/or visualized in augmented reality.
Conclusions
Normscan provides an end-to-end pipeline for the creation of average models of skulls. These models can be used for the generation of databases of specific demographic anatomical models as well as for intraoperative guidance and surgical planning. While Normscan was designed for craniosynostosis repair, due to the modular nature of the algorithm, Normscan has many applications in other areas of surgical planning and research.





Similar content being viewed by others
References
Goodwin AF, Kim R, Bush JO, Klein OD (2015) From bench to bedside and back: improving diagnosis and treatment of craniofacial malformations utilizing animal models. Curr Top Dev Biol 115:459–492. https://doi.org/10.1016/bs.ctdb.2015.07.003
Kajdic N, Spazzapan P, Velnar T (2018) Craniosynostosis - recognition, clinical characteristics, and treatment. Bosn J Basic Med Sci 18:110–116. https://doi.org/10.17305/bjbms.2017.2083
García-Mato D, Ochandiano S, García-Sevilla M, Navarro-Cuéllar C, Darriba-Allés JV, García-Leal R, Calvo-Haro JA, Pérez-Mañanes R, Salmerón JI, Pascau J (2019) Craniosynostosis surgery: workflow based on virtual surgical planning, intraoperative navigation and 3D printed patient-specific guides and templates. Sci Rep 9:17691. https://doi.org/10.1038/s41598-019-54148-4
Bhalodia R, Dvoracek LA, Ayyash AM, Kavan L, Whitaker R, Goldstein JA (2020) Quantifying the severity of metopic craniosynostosis: a pilot study application of machine learning in craniofacial surgery. J Craniofac Surg 31:697. https://doi.org/10.1097/SCS.0000000000006215
Marcus JR, Domeshek LF, Das R, Marshall S, Nightingale R, Stokes TH, Mukundan S (2008) Objective three-dimensional analysis of cranial morphology. Eplasty 8:e20
Teshima TL, Patel V, Mainprize JG, Edwards G, Antonyshyn OM (2015) A three-dimensional statistical average skull: application of biometric morphing in generating missing anatomy. J Craniofac Surg 26:1634. https://doi.org/10.1097/SCS.0000000000001869
Saber NR, Phillips J, Looi T, Usmani Z, Burge J, Drake J, Kim PCW (2012) Generation of normative pediatric skull models for use in cranial vault remodeling procedures. Childs Nerv Syst 28:405–410. https://doi.org/10.1007/s00381-011-1630-7
Weinberg SM, Raffensperger ZD, Kesterke MJ, Heike CL, Cunningham ML, Hecht JT, Kau CH, Murray JC, Wehby GL, Moreno LM, Marazita ML (2016) The 3D facial norms database: part 1. a web-based craniofacial anthropometric and image repository for the clinical and research community. Cleft Palate-Craniofacial J Off Publ Am Cleft Palate-Craniofacial Assoc 53:e185–e197. https://doi.org/10.1597/15-199
Zhou Q-Y, Park J, Koltun V (2018) Open3D: A Modern Library for 3D Data Processing
Materialise Mimics | 3D Medical Image Processing Software. https://www.materialise.com/en/healthcare/mimics-innovation-suite/mimics. Accessed 2 Sep 2023
Besl PJ, McKay ND (1992) A method for registration of 3-D shapes. IEEE Trans Pattern Anal Mach Intell 14:239–256. https://doi.org/10.1109/34.121791
Kazhdan MM, Bolitho M, Hoppe H (2006) Poisson Surface Reconstruction. In: Sheffer A, Polthier K (eds) In: Proceedings of the Fourth Eurographics Symposium on Geometry Processing. Eurographics Association, Cagliari, Sardinia, Italy, pp 61–70
Mirtich B (1996) Fast and accurate computation of polyhedral mass properties. J Graph Tools 1:31–50. https://doi.org/10.1080/10867651.1996.10487458
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin J-C, Pujol S, Bauer C, Jennings D, Fennessy F, Sonka M, Buatti J, Aylward S, Miller JV, Pieper S, Kikinis R (2012) 3D slicer as an image computing platform for the quantitative imaging network. Magn Reson Imaging 30:1323–1341. https://doi.org/10.1016/j.mri.2012.05.001
Ho OA, Saber N, Stephens D, Clausen A, Drake J, Forrest C, Phillips J (2017) Comparing the use of 3D photogrammetry and computed tomography in assessing the severity of single-suture nonsyndromic craniosynostosis. Plast Surg 25:78–83. https://doi.org/10.1177/2292550317694845
Lu J, Wang Z, Hua B, Chen K (2020) Automatic point cloud registration algorithm based on the feature histogram of local surface. PLoS ONE 15:e0238802. https://doi.org/10.1371/journal.pone.0238802
Acknowledgements
We gratefully acknowledge Robert Finedor for thoughtful discussions on techniques to optimize the algorithm. We also thank the Alkureishi lab for their support, thoughtful academic guidance on this project, and for creating a wonderful working environment.
Author information
Authors and Affiliations
Contributions
GRN and LA designed the study; GRN performed the experiments and analyzed the data; MAM and NBZ performed statistical analysis, LA supervised the experiments; GRN, NBZ, and LA wrote the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors have no conflicts of interest.
Previous presentations
This work was presented as a poster at the The Midwest Association of Plastic Surgeons 61st Annual Meeting in July 2023 in Lake Geneva, WI.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nahass, G.R., Marques, M.A., Bou Zeid, N. et al. Normscan: open-source Python software to create average models from CT scans. Int J CARS (2024). https://doi.org/10.1007/s11548-024-03185-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11548-024-03185-0