Abstract
Purpose
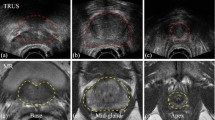
This paper addresses the detection of the clinical target volume (CTV) in transrectal ultrasound (TRUS) image-guided intraoperative for permanent prostate brachytherapy. Developing a robust and automatic method to detect the CTV on intraoperative TRUS images is clinically important to have faster and reproducible interventions that can benefit both the clinical workflow and patient health.
Methods
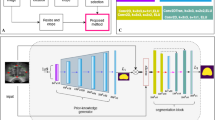
We present a multi-task deep learning method for an automatic prostate CTV boundary detection in intraoperative TRUS images by leveraging both the low-level and high-level (prior shape) information. Our method includes a channel-wise feature calibration strategy for low-level feature extraction and learning-based prior knowledge modeling for prostate CTV shape reconstruction. It employs CTV shape reconstruction from automatically sampled boundary surface coordinates (pseudo-landmarks) to detect the low-contrast and noisy regions across the prostate boundary, while being less biased from shadowing, inherent speckles, and artifact signals from the needle and implanted radioactive seeds.
Results
The proposed method was evaluated on a clinical database of 145 patients who underwent permanent prostate brachytherapy under TRUS guidance. Our method achieved a mean accuracy of \( 0.96 \pm 0.01\) and a mean surface distance error of \(0.10 \pm 0.06 \, \hbox {mm}\). Extensive ablation and comparison studies show that our method outperformed previous deep learning-based methods by more than 7% for the Dice similarity coefficient and 6.9 mm reduced 3D Hausdorff distance error.
Conclusion
Our study demonstrates the potential of shape model-based deep learning methods for an efficient and accurate CTV segmentation in an ultrasound-guided intervention. Moreover, learning both low-level features and prior shape knowledge with channel-wise feature calibration can significantly improve the performance of deep learning methods in medical image segmentation.

Source: modified image from Cancer Research UK



Similar content being viewed by others
References
Davis BJ, Horwitz EM, Lee WR, Crook JM, Stock RG, Merrick GS, Butler WM, Grimm PD, Stone NN, Potters L, Zietman AL (2012) American Brachytherapy Society consensus guidelines for transrectal ultrasound-guided permanent prostate brachytherapy. Brachytherapy 11:6–19. https://doi.org/10.1016/j.brachy.2011.07.005
Girum KB, Lalande A, Quivrin M, Bessières I, Pierrat N, Martin E, Cormier L, Petitfils A, Cosset JM, Créhange G (2018) Inferring postimplant dose distribution of salvage permanent prostate implant (PPI) after primary PPI on CT images. Brachytherapy 17:866–873. https://doi.org/10.1016/j.brachy.2018.07.017
Zelefsky MJ, Cohen GN, Taggar AS, Kollmeier M, McBride S, Mageras G, Zaider M (2017) Real-time intraoperative evaluation of implant quality and dose correction during prostate brachytherapy consistently improves target coverage using a novel image fusion and optimization program. Pract Radiat Oncol 7:319–324. https://doi.org/10.1016/j.prro.2017.01.009
Jaouen V, Bert J, Mountris KA, Boussion N, Schick U, Pradier O, Valeri A, Visvikis D (2019) Prostate volume segmentation in TRUS using hybrid edge-Bhattacharyya active surfaces. IEEE Trans Biomed Eng 66:920–933. https://doi.org/10.1109/TBME.2018.2865428
Karimi D, Zeng Q, Mathur P, Avinash A, Mahdavi S, Spadinger I, Abolmaesumi P, Salcudean SE (2019) Accurate and robust deep learning-based segmentation of the prostate clinical target volume in ultrasound images. Med Image Anal 57:186–196. https://doi.org/10.1016/j.media.2019.07.005
Wang Y, Dou H, Hu X, Zhu L, Yang X, Xu M, Qin J, Heng PA, Wang T, Ni D (2019) Deep attentive features for prostate segmentation in 3D transrectal ultrasound. IEEE Trans Med Imaging. https://doi.org/10.1109/TMI.2019.2913184
Ghavami N, Hu Y, Bonmati E, Rodell R, Gibson E, Moore C, Barratt D (2018) Integration of spatial information in convolutional neural networks for automatic segmentation of intraoperative transrectal ultrasound images. J Med Imaging 6:1. https://doi.org/10.1117/1.JMI.6.1.011003
Anas EMA, Mousavi P, Abolmaesumi P (2018) A deep learning approach for real time prostate segmentation in freehand ultrasound guided biopsy. Med Image Anal 48:107–116. https://doi.org/10.1016/j.media.2018.05.010
Ghose S, Oliver A, Martí R, Lladó X, Vilanova JC, Freixenet J, Mitra J, Sidibé D, Meriaudeau F (2012) A survey of prostate segmentation methodologies in ultrasound, magnetic resonance and computed tomography images. Comput Methods Programs Biomed 108(262):87. https://doi.org/10.1016/j.cmpb.2012.04.006
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation, pp 234–241. https://doi.org/10.1007/978-3-319-24574-4_28
Goceri E, Goceri N (2017) Deep learning in medical image analysis: recent advances and future trends. In: 2017 International conference on computer graphics, visualization, computer vision and image processing CGVCVIP, pp 305–310
Hu J, Shen L, Sun G (2018) Squeeze-and-excitation networks. In: CVPR, pp 7132–7141. https://doi.org/10.1109/CVPR.2018.00745
He K, Gkioxari G, Dollár P, Girshick R (2017) Mask r-cnn. In: ICCV, pp 2961–2969. https://doi.org/10.1109/TPAMI.2018.2844175
Goceri E (2019) Challenges and recent solutions for image segmentation in the era of deep learning. In: 2019 Ninth IPTA, pp 1–6. https://doi.org/10.1109/IPTA.2019.8936087
Radford A, Metz L, Chintala S (2015) Unsupervised representation learning with deep convolutional generative adversarial networks. In: ICLR, pp 1–16. arXiv:1511.06434
Lin M, Chen Q, Yan S (2013) Network in network, pp 1–10. arXiv:1312.4400
Zhou B, Khosla A, Lapedriza A, Oliva A, Torralba A (2016) Learning deep features for discriminative localization. In: CVPR. IEEE, pp 2921–2929. https://doi.org/10.1109/CVPR.2016.319
Girum KB, Créhange G, Hussain R, Walker PM, Lalande A (2019) Deep generative model-driven multimodal prostate segmentation in radiotherapy. In: AIRT, pp 119–127. https://doi.org/10.1007/978-3-030-32486-5_15
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization, pp 1–15. arXiv:1412.6980
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: ICVPR, pp 770–778
Goceri E (2019) Analysis of deep networks with residual blocks and different activation functions: classification of skin diseases. In: 2019 Ninth IPTA, pp 1–6. https://doi.org/10.1109/IPTA.2019.8936083
Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9:1735–1780. https://doi.org/10.1162/neco.1997.9.8.1735
Acknowledgements
The authors would like to thank NVIDIA for providing GPU (NVIDIA TITAN X, 12 GB) through their GPU Grant program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval and informed consent
Ethical approval and informed consent were not required for this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Girum, K.B., Lalande, A., Hussain, R. et al. A deep learning method for real-time intraoperative US image segmentation in prostate brachytherapy. Int J CARS 15, 1467–1476 (2020). https://doi.org/10.1007/s11548-020-02231-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-020-02231-x