Abstract
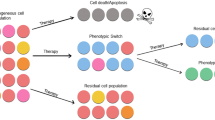
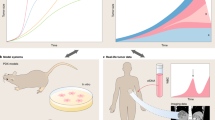
Evolutionary dynamics allows us to understand many changes happening in a broad variety of biological systems, ranging from individuals to complete ecosystems. It is also behind a number of remarkable organizational changes that happen during the natural history of cancers. These reflect tumour heterogeneity, which is present at all cellular levels, including the genome, proteome and phenome, shaping its development and interrelation with its environment. An intriguing observation in different cohorts of oncological patients is that tumours exhibit an increased proliferation as the disease progresses, while the timescales involved are apparently too short for the fixation of sufficient driver mutations to promote explosive growth. Here, we discuss how phenotypic plasticity, emerging from a single genotype, may play a key role and provide a ground for a continuous acceleration of the proliferation rate of clonal populations with time. We address this question by combining the analysis of real-time growth of non-small-cell lung carcinoma cells (N-H460) together with stochastic and deterministic mathematical models that capture proliferation trait heterogeneity in clonal populations to elucidate the contribution of phenotypic transitions on tumour growth dynamics.






Similar content being viewed by others
Code Availability
Simulations were conducted in MATLAB (version R2020a). Code files for the discrete model simulations and experimental data are publicly accessible at https://github.com/molabEvoDynamics/rep_StochasticFluctuationsDriveNonGeneticEvolution.
References
Aaronson S (1991) Growth factors and cancer. Science 254(5035):1146–1153. https://doi.org/10.1126/science.1659742
Altrock PM, Liu LL, Michor F (2015) The mathematics of cancer: integrating quantitative models. Nat Rev Cancer 15:730–745
Álvarez-Arenas A, Podolski-Renic A, Belmonte-Beitia J et al (2019) Interplay of Darwinian selection, Lamarckian Induction and microvesicle transfer on drug resistance in cancer. Sci Rep 9:9332. https://doi.org/10.1038/s41598-019-45863-z
Archetti M, Pienta KJ (2019) Cooperation among cancer cells: applying game theory to cancer. Nat Rev Cancer 19(2):110–117. https://doi.org/10.1038/s41568-018-0083-7
Ardaševa A, Anderson ARA, Gatenby RA et al (2020a) A comparative study between discrete and continuum models for the evolution of competing phenotype-structured cell populations in dynamical environments. Phys Rev E 102:042404. https://doi.org/10.1103/PhysRevE.102.042404
Ardaševa A, Gatenby RA, Anderson ARA et al (2020b) Evolutionary dynamics of competing phenotype-structured populations in periodically fluctuating environments. J Math Biol 80:775–807
Ardaševa A, Gatenby RA, Anderson ARA et al (2020c) A mathematical dissection of the adaptation of cell populations to fluctuating oxygen levels. Bull Math Biol 82(6):1–24. https://doi.org/10.1007/s11538-020-00754-7
Arim M, Abades SR, Neill PE, Lima M, Marquet PA (2006) Spread dynamics of invasive species. Proc Natl Acad Sci USA 103:374–378. https://doi.org/10.1073/pnas.0504272102
Batlle E, Clevers H (2017) Cancer stem cells revisited. Nat Med 23:1124–1134. https://doi.org/10.1038/nm.4409
Baylin SB, Jones PA (2016) Epigenetic determinants of cancer. Cold Spring Harb Perspect Biol 8:a019505. https://doi.org/10.1101/cshperspect.a019505
Benzekry S, Lamon C, Beheshti A et al (2014) Classical mathematical models for description and prediction of experimental tumor growth. PLoS Comput Biol 10:e1003800
Bertolaso M, Dieli AM (2017) Cancer and intercellular cooperation. J R Soc 4(10):170470. https://doi.org/10.1098/rsos.170470
Blum WF, Albertsson-Wikland K, Rosberg S, Ranke MB (1993) Serum levels of insulin-like growth factor I (IGF-1) and IGF binding protein 3 reflect spontaneous growth hormone secretion. J Clin Endocrinol Metab 76:1610–1616. https://doi.org/10.1210/jcem.76.6.7684744
Brandman O, Meyer T (2008) Feedback loops shape cellular signals in space and time. Science 322:390–395. https://doi.org/10.1126/science.1160617
Buehring GC, Williams RR (1976) Growth rate of normal and abnormal human mammary epithelia in cell culture. Cancer Res 36:3742–3747
Butler G, Keeton SJ, Johnson LJ, Dash PR (2020) A phenotypic switch in the dispersal strategy of breast cancer cells selected for metastatic colonization. Proc R Soc B 287:20202523. https://doi.org/10.1098/rspb.2020.2523
Byrne HM (2010) Dissecting cancer through mathematics: from the cell to the animal model. Nat Rev Cancer 10:221–230
Cairns J (1975) Mutation selection and the natural history of cancer. Nature 255:197–200. https://doi.org/10.1038/255197a0
Carmona-Fontaine C, Deforet M, Akkari L, Xavier J (2017) Metabolic origins of spatial organization in the tumor microenvironment. Proc Natl Acad Sci USA 114(11):2934–2939. https://doi.org/10.1073/pnas.1700600114
Chiodoni C, Di Martino MT, Zazzeroni F et al (2019) Cell communication and signaling: how to turn bad language into positive one. J Exp Clin Cancer Res 38(1):128. https://doi.org/10.1186/s13046-019-1122-2
Chiou SH, Cheng-Chia Y, Chi-Yang H et al (2008) Positive correlations of Oct-4 and Nanog in oral cancer stem-like cells and high-grade oral squamous cell carcinoma. Clin Cancer Res 14:4085–4095. https://doi.org/10.1158/1078-0432.CCR-07-4404
Codling EA, Plank MJ, Benhamou S (2008) Random walk models in biology. J R Soc Interface 5:813–834. https://doi.org/10.1098/rsif.2008.0014
Czarkowska-Paczek B, Bartlomiejczyk I, Przybylski J (2006) The serum levels of growth factors: PDGF, TGF-beta and VEGF are increased after strenuous physical exercise. J Physiol Pharmacol 57:189–97
Darwin C (1859) On the origin of species by means of natural selection, or, the preservation of favoured races in the struggle for life
Daughaday WH, Deuel TF (1991) Tumor secretion of growth factors. Endocrinol Metab Clin 20(3):539–63
Davis TW, Berry DL, Boyer GL, Gobler CH (2009) The effects of temperature and nutrients on the growth and dynamics of toxic and non-toxic strains of Microcystis during cyanobacteria blooms. Harmful Algae 8:715–725. https://doi.org/10.1016/j.hal.2009.02.004
Deforet M, Carmona-Fontaine C, Korolev KS, Xavier JB (2019) Evolution at the edge of expanding populations. Am Nat 194:291–305. https://doi.org/10.1086/704594
Easwaran H, Tsai HC, Baylin SB (2014) Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states and drug resistance. Mol Cell 54:716–727. https://doi.org/10.1016/j.molcel.2014.05.015
Eugenin EA (2019) Role of cell-to-cell communication in cancer: new features, insights, and directions. Cancer Rep 2:e1228. https://doi.org/10.1002/cnr2.1228
Feinberg AP, Irizarry RA (2010) Stochastic epigenetic variation as a driving force of development, evolutionary adaptation, and disease. Proc Natl Acad Sci USA 107:1757–1764. https://doi.org/10.1073/pnas.0906183107
Fiandaca G, Delitala M, Lorenzi T (2021) A mathematical study of the influence of hypoxia and acidity on the evolutionary dynamics of cancer. Bull Math Biol 83(7):1–29. https://doi.org/10.1007/s11538-021-00914-3
Flavahan WA, Gaskell E, Bernstein BE (2017) Epigenetic plasticity and the hallmarks of cancer. Science 357:eaal2380. https://doi.org/10.1126/science.aal2380
Frick P, Paudel B, Tyson D, Quaranta V (2015) Quantifying heterogeneity and dynamics of clonal fitness in response to perturbation. J Cell Physiol 230:1403–1412. https://doi.org/10.1002/jcp.24888
Geiler-Samerotte KA, Bauer CR, Li S, Ziv N, Gresham D, Siegal ML (2013) The details in the distributions: why and how to study phenotypic variability. Curr Opin Biotechnol 24:752–759. https://doi.org/10.1016/j.copbio.2013.03.010
Gerashchenko TS, Denisov EV, Litviakov NV et al (2013) Intratumor heterogeneity: nature and biological significance. Biochemistry 78(11):1201–1215. https://doi.org/10.1134/S0006297913110011
Gerlee P (2013) The model muddle: in search of tumor growth laws. Cancer Res 73:2407–2411
Goldberg AD, Allis CD, Bernstein E (2007) Epigenetics: a landscape takes shape. Cell 128:635–638. https://doi.org/10.1016/j.cell.2007.02.006
Greaves M (2015) Evolutionary determinants of cancer. Cancer Discov 5:806–821. https://doi.org/10.1158/2159-8290.CD-15-0439
Greene JM, Levy D, Herrada SP, Gottesman MM, Lavi O (2016) Mathematical modeling reveals that changes to local cell density dynamically modulate baseline variations in cell growth and drug response. Cancer Res 76:2882–2890. https://doi.org/10.1158/0008-5472.CAN-15-3232
Gupta PB, Fillmore CM, Jiang G et al (2011) Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell 146:633–644. https://doi.org/10.1016/j.cell.2011.07.026
Gupta PB, Pastushenko I, Skibinski A, Blanpain C, Kuperwasser C (2019) Phenotypic plasticity: driver of cancer initiation, progression, and therapy resistance. Cell Stem Cell 24:65–78. https://doi.org/10.1016/j.stem.2018.11.011
Hallastschek O, Fisher DS (2014) Acceleration of evolutionary spread by long-range dispersal. Proc Natl Acad Sci USA 111:E4911–E4919. https://doi.org/10.1073/pnas.1404663111
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674. https://doi.org/10.1016/j.cell.2011.02.013
Huang S (2013) Genetic and non-genetic instability in tumor progression: link between the fitness landscape and the epigenetic landscape of cancer cells. Cancer Metastasis Rev 32:423–448. https://doi.org/10.1007/s10555-013-9435-7
Huang S (2021) Reconciling non-genetic plasticity with somatic evolution in cancer. Trends Cancer 7:309–322. https://doi.org/10.1016/j.trecan.2020.12.007
Jiménez-Sánchez J, Bosque JJ, Jiménez-Londoño G et al (2021) Evolutionary dynamics at the tumor edge reveal metabolic imaging biomarkers. Proc Natl Acad Sci USA 118:e2018110118. https://doi.org/10.1073/pnas.2018110118
Kærn M, Elston T, Blake W (2005) Stochasticity in gene expression: from theories to phenotypes. Nat Rev Genet 6:451–464. https://doi.org/10.1038/nrg1615
Karki P, Sensenbach S, Angardi V et al (2021) BRAF-inhibitor-induced metabolic alterations in A375 melanoma cells. Metabolites 11(11):777. https://doi.org/10.3390/metabo11110777
Komarova NL (2014) Spatial interactions and cooperation can change the speed of evolution of complex phenotypes. Proc Natl Acad Sci USA 111:10789–10795. https://doi.org/10.1073/pnas.1400828111
Laland KN, Uller T, Feldman MW et al (2015) The extended evolutionary synthesis: its structure, assumptions and predictions. Proc R Soc Lond B 282:20151019. https://doi.org/10.1098/rspb.2015.1019
Leroi AM, Lenski RE, Bennett AF (1994) Evolutionary adaptation to temperature. III. Adaptation of Escherichia coli to a temporally varying environment. Evolution 48:1222–1229. https://doi.org/10.1111/j.1558-5646.1994.tb05307.x
Lipinski KA, Barber LJ, Davies MN et al (2016) Cancer evolution and the limits of predictability in precision cancer medicine. Trends Cancer 2:49–63. https://doi.org/10.1016/j.trecan.2015.11.003
Maheshri N, O’Shea EK (2007) Living with noisy genes: how cells function reliably with inherent variability in gene expression. Annu Rev Biophys Biomol Struct 36:413–34. https://doi.org/10.1146/annurev.biophys.36.040306.132705
Mansoori B, Baradaran B, Nazari A et al (2022) miRNAs in the cancer cell-to-cell communication: an insight into biological vehicles. Biomed Pharmacother 153:113449. https://doi.org/10.1016/j.biopha.2022.113449
Mardin BR et al (2013) EGF-induced centrosome separation promotes mitotic progression and cell survival. Dev Cell 25:229–40. https://doi.org/10.1016/j.devcel.2013.03.012
Marine JC, Dawson SJ, Dawson MA (2020) Non-genetic mechanisms of therapeutic resistance in cancer. Nat Rev Cancer 20:743–756. https://doi.org/10.1038/s41568-020-00302-4
McGranahan N, Swanton C (2017) Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168:613–628. https://doi.org/10.1016/j.cell.2017.01.018
Norberg J, Swaney DP, Dushoff J, Lin J, Casagrandi R (2001) Phenotypic diversity and ecosystem functioning in changing environments: a theoretical framework. Proc Natl Acad Sci USA 98:11376–11381. https://doi.org/10.1073/pnas.171315998
Nowell PC (1976) The clonal evolution of tumor cell populations. Science 194:23–28. https://doi.org/10.1126/science.959840
Oren Y, Tsabar M, Cuoco MS et al (2021) Cycling cancer persister cells arise from lineages with distinct programs. Nature. https://doi.org/10.1038/s41586-021-03796-6
Paczkowski M, Kretzschmar W, Markelc B et al (2021) Reciprocal interactions between tumour cell populations enhance growth and reduce radiation sensitivity in prostate cancer. Commun Biol 4:6. https://doi.org/10.1038/s42003-020-01529-5
Pérez-García VM et al (2020) Universal scaling laws rule explosive growth in human cancers. Nat Phys 16:1232–1237. https://doi.org/10.1038/s41567-020-0978-6
Powathil GG, Gordon KE, Hill LA, Chaplain MA (2012) Modelling the effects of cell-cycle heterogeneity on the response of a solid tumour to chemotherapy: biological insights from a hybrid multiscale cellular automaton model. J Theor Biol 308:1–19. https://doi.org/10.1016/j.jtbi.2012.05.015
Rabé M, Dumont S, Alvarez-Arenas A et al (2020) Identification of a transient state during the acquisition of temozolomide resistance in glioblastoma. Cell Death Dis. https://doi.org/10.1038/s41419-019-2200-2
Raj A, van Oudenaarden A (2008) Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135(2):216–26. https://doi.org/10.1016/j.cell.2008.09.050
Reboud X, Bell G (1997) Experimental evolution in Chlamydomonas. III. Evolution of specialist and generalist types in environments that vary in space and time. Heredity 78:507–514. https://doi.org/10.1038/hdy.1997.79
Rehman SK et al (2021) Colorectal cancer cells enter a diapause-like DTP state to survive chemotherapy. Cell 184:226–242. https://doi.org/10.1016/j.cell.2020.11.018
Reinius B, Sandberg R (2015) Random monoallelic expression of autosomal genes: stochastic transcription and allele-level regulation. Nat Rev Genet 16(11):653–64. https://doi.org/10.1038/nrg3888
Russo M, Pompei S, Sogari A et al (2022) A modified fluctuation-test framework characterizes the population dynamics and mutation rate of colorectal cancer persister cells. Nat Genet 54(7):976. https://doi.org/10.1038/s41588-022-01105-z
Schwager SC, Taufalele PV, Reinhart-King CA (2019) Cell–cell mechanical communication in cancer. Cell Mol Bioeng 12(1):1–14. https://doi.org/10.1007/s12195-018-00564-x
Shaffer SM, Dunagin MC, Torborg SR et al (2018) Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature. https://doi.org/10.1038/nature25162
Shen SS, Vagner S, Robert C (2020a) Persistent cancer cells: the deadly survivors. Cell 183(4):860–874. https://doi.org/10.1016/j.cell.2020.10.027
Shen SS, Faouzi S, Souquere S et al (2020b) Melanoma persister cells are tolerant to BRAF/MEK inhibitors via ACOX1-mediated fatty acid oxidation. Cell Rep 33(8):108421. https://doi.org/10.1016/j.celrep.2020.108421
Siezen RJ, Tzeneva VA, Castioni A et al (2010) Phenotypic and genomic diversity of Lactobacillus plantarum strains isolated from various environmental niches. Environ Microbiol 12:758–73. https://doi.org/10.1111/j.1462-2920.2009.02119.x
Smith HS, Lan S, Ceriani R, Hackett AJ, Stamper MR (1981) Clonal proliferation of cultured nonmalignant and malignant human-breast epithelia. Cancer Res 41:4637–4643
Soltani M, Vargas-Garcia CA, Antunes D, Singh A (2016) Intercellular variability in protein levels from stochastic expression and noisy cell cycle processes. PLoS Comput Biol 12:e1004972. https://doi.org/10.1371/journal.pJRSIi.1004972
Turner BM (2009) Epigenetic responses to environmental change and their evolutionary implications. Philos Trans R Soc Lond B Biol Sci 364:3403–3418. https://doi.org/10.1098/rstb.2009.0125
Tzamali E, Tzedakis G, Sakkalis V (2020) Modeling how heterogeneity in cell cycle length affects cancer cell growth dynamics in response to treatment. Front Oncol 10:1552. https://doi.org/10.3389/fonc.2020.01552
Vandel Verde R, Yoon N, Marusyk V et al (2020) Resistance to targeted therapies as a multifactorial, gradual adaptation to inhibitor specific selective pressures. Nat Commun 11:2393. https://doi.org/10.1038/s41467-020-16212-w
Vendramin R, Litchfield K, Swanton C (2021) Cancer evolution: Darwin and beyond. EMBO J 40(18):e108389. https://doi.org/10.15252/embj.2021108389
Viossat Y, Noble R (2021) A theoretical analysis of tumour containment. Nat Ecol Evol 5:826–835. https://doi.org/10.1038/s41559-021-01428-w
Vogelstein B, Papadopoulos N, Velculescu VE et al (2013) Cancer genome landscapes. Science 340:1546–1558. https://doi.org/10.1126/science.1235122
Waclaw B, Bozic I, Pittman M et al (2015) A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525:261–264. https://doi.org/10.1038/nature14971
Weinhold B (2006) Epigenetics: the science of change. Environ Health Perspect 114:A160–A167. https://doi.org/10.1289/ehp.114-a160
Acknowledgements
Authors thank Juan Jiménez Sánchez and Jesús Bosque Martínez (Universidad de Castilla-La Mancha, Spain) for their fruitful feedback on this work. We also thank Juan Antonio Delgado (Universidad de Murcia, Spain) for kindly sharing his broad knowledge on translational evolutionary theory with us through valuable discussions.
Funding
This work was supported by the James S. McDonnell Foundation (Collaborative award 220020450, doi: 10.37717/220020560), Junta de Comunidades de Castilla-La Mancha (Grants SBPLY/17/180501/000154 and SBPLY/19/180501/000211), Ministerio de Ciencia e Innovación MCIN/AEI/10.13039/501100011033 (grant PID2019-110895RB-I00) and Asociación Española Contra el Cáncer (Grant 2019-PRED-28372).
Author information
Authors and Affiliations
Contributions
GFC and VMP-G formulated the hypothesis and designed the research. JD, AP-R and MP conceived and performed the experiments. GFC, CO-S and VMP-G performed the numerical simulations and statistical analyses. GFC carried out the analytical study. CO-S, GFC and VMP-G wrote the paper. All authors gave final approval for publication.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ortega-Sabater, C., F. Calvo, G., Dinić, J. et al. Stochastic Fluctuations Drive Non-genetic Evolution of Proliferation in Clonal Cancer Cell Populations. Bull Math Biol 85, 8 (2023). https://doi.org/10.1007/s11538-022-01113-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11538-022-01113-4