Abstract
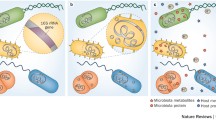
Several urinary disorders, including overactive bladder, urinary incontinence, and interstitial cystitis, are often characterized by negative urine cultures. The application of metagenomics (i.e., 16S rRNA microbial profiling or whole-genome shotgun sequencing) to urine samples has enabled the identification of previously undetected bacteria, contributing to the discovery and characterization of the urinary microbiome. The most frequent species isolated are Lactobacillus (15%), Corynebacterium (14.2%), Streptococcus (11.9%), Actinomyces (6.9%), and Staphylococcus (6.9%). Although several studies are emerging in this context, the role of urinary microbiota in the pathogenesis of infections and in tumor carcinogenesis remains unclear. Furthermore, data on the activity of gut microbiota in modulating sensitivity to immune checkpoint inhibitors in advanced cancer patients suggest that the influence of urinary microbiota on tumor response to anticancer therapy should also be investigated. Moreover, its possible relationship with tumor mutational burden, which is in turn correlated with response to immunotherapy, should be the focus of future studies. Of note, the effect of antibiotics on this complex scenario seems to deserve careful consideration.

Similar content being viewed by others
References
Siddiqui H, Nederbragt AJ, Lagesen K, Jeansson SL, Jakobsen KS. Assessing diversity of the female urine microbiota by high throughput sequencing of 16S rDNA amplicons. BMC Microbiol. 2011;11:244.
Wolfe AJ, Toh E, Shibata N, Rong R, Kenton K, Fitzgerald M, Mueller ER, Schreckenberger P, Dong Q, Nelson DE, Brubaker L. Evidence of uncultivated bacteria in the adult female bladder. J Clin Microbiol. 2012;50:1376–83.
Hilt EE, McKinley K, Pearce MM, Rosenfeld AB, Zilliox MJ, Mueller ER, Brubaker L, Gai X, Wolfe AJ, Schreckenberger PC. Urine is not sterile: use of enhanced urine culture techniques to detect resident bacterial flora in the adult female bladder. J Clin Microbiol. 2014;52:871–6.
Ticinesi A, Nouvenne A, Tana C, Prati B, Cerundolo N, Miraglia C, de Angelis GL, Di Mario F, Meschi T. The impact of intestinal microbiota on bio-medical research: definitions, techniques and physiology of a “new frontier”. Acta Biomed. 2018;89(9S):52–9.
Fouts DE, Pieper R, Szpakowski S, Pohl H, Knoblach S, Suh MJ, Huang ST, Ljungberg I, Sprague BM, Lucas SK, Torralba M, Nelson KE, Groah SL. Integrated next-generation sequencing of 16S rDNA and metaproteomics differentiate the healthy urine microbiome from asymptomatic bacteriuria in neuropathic bladder associated with spinal cord injury. J Transl Med. 2012;10:174.
Lewis DA, Brown R, Williams J, White P, Jacobson SK, Marchesi JR, Drake MJ. The human urinary microbiome; bacterial DNA in voided urine of asymptomatic adults. Front Cell Infect Microbiol. 2013;3:41.
Sheflin AM, Whitney AK, Weir TL. Cancer-promoting effects of microbial dysbiosis. Curr Oncol Rep. 2014;16(10):406.
Dave M, Higgins PDR, Middha S, Rioux K. The human gut microbiome: current knowledge, challenges, and future directions. Transl Res. 2012;160:246–57.
Thomas AM, Manghi P, Asnicar F, Pasolli E, Armanini F, Zolfo M, et al. Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation. Nat Med. 2019;25(4):667–78.
Shrestha E, White JR, Yu SH, Kulac I, Ertunc O, De Marzo AM, Yegnasubramanian S, Mangold LA, Partin AW, Sfanos KS. Profiling the urinary microbiome in men with positive versus negative biopsies for prostate cancer. J Urol. 2018;199:161–71.
Puhr M, De Marzo A, Isaacs W, Lucia MS, Sfanos K, Yegnasubramanian S, Culig Z. Inflammation, microbiota, and prostate cancer. Eur Urol Focus. 2016;2:374–82.
Alanee S, El-Zawahry A, Dynda D, Dabaja A, McVary K, Karr M, Braundmeier-Fleming A. A prospective study to examine the association of the urinary and fecal microbiota with prostate cancer diagnosis after transrectal biopsy of the prostate using 16sRNA gene analysis. Prostate. 2019;79:81–7.
Markowski MC, Boorjian SA, Burton JP, Hahn NM, Ingersoll MA, Vareki SM, Pal SK, Sfanos KS. The microbiome and genitourinary cancer: a collaborative review. Eur Urol. 2019;18:31051.
Bajic P, Wolfe AJ, Gupta GN. The urinary microbiome: implications in bladder cancer pathogenesis and therapeutics. Urology. 2019;126:10–5.
Gopalakrishnan V, Spencer CN, Nezi L, et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science. 2018;359:97–103.
Matson V, Fessler J, Bao R, Chongsuwat T, Zha Y, Alegre ML, Luke JJ, Gajewski TF. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science. 2018;359:104–8.
Routy B, Le Chatelier E, Derosa L, et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359:91–7.
Santoni M, Piva F, Conti A, Santoni A, Cimadamore A, Scarpelli M, Battelli N, Montironi R. Re: gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Eur Urol. 2018;74:521–2.
Park J, Kim M, Kang SG, et al. Short-chain fatty acids induce both effector and regulatory T cells by suppression of histone deacetylases and regulation of the mTOR-S6 K pathway. Mucosal Immunol. 2015;8:80–93.
Katayama Y, Yamada T, Tanimura K, Yoshimura A, Takeda T, Chihara Y, et al. Impact of bowel movement condition on immune checkpoint inhibitor efficacy in patients with advanced non-small cell lung cancer. Thorac Cancer. 2019;10(3):526–32.
Vandeputte D, Falony G, Vieira-Silva S, Tito RY, Joossens M, Raes J. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut. 2016;65(1):57–62.
Mancabelli L, Milani C, Lugli GA, et al. Unveiling the gut microbiota composition and functionality associated with constipation through metagenomic analyses. Sci Rep. 2017;7(1):9879.
Belkaid Y, Segre JA. Dialogue between skin microbiota and immunity. Science. 2014;346:954–9.
Thio H. The microbiome in psoriasis and psoriatic arthritis: the skin perspective. J Rheumatol Suppl. 2018;94:30–1.
Yu Y, Champer J, Beynet D, Kim J, Friedman AJ. The role of the cutaneous microbiome in skin cancer: lessons learned from the gut. J Drugs Dermatol. 2015;14(5):461–5.
Alexandrov LB, Nik-Zainal S, Wedge DC, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, et al. PD-1 Blockade in tumors with mismatch-repair deficiency. N Engl J Med. 2015;372:2509–20.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. 2015;348:124–8.
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371:2189–99.
Lalani AA, Sonpavde GP. Systemic treatments for metastatic urothelial carcinoma. Expert Opin Pharmacother. 2019;20:201–8.
Atkins MB, Clark JI, Quinn DI. Immune checkpoint inhibitors in advanced renal cell carcinoma: experience to date and future directions. Ann Oncol. 2017;28:1484–94.
Derosa L, Hellmann MD, Spaziano M, et al. Negative association of antibiotics on clinical activity of immune checkpoint inhibitors in patients with advanced renal cell and non-small cell lung cancer. Ann Oncol. 2018; https://doi.org/10.1093/annonc/mdy103.
Elkrief A, El Raichani L, Richard C, Messaoudene M, Belkaid W, Malo J, et al. Antibiotics are associated with decreased progression-free survival of advanced melanoma patients treated with immune checkpoint inhibitors. Oncoimmunology. 2019;8(4):e1568812.
Zilberman-Schapira G, Zmora N, Itav S, Bashiardes S, Elinav H, Elinav E. The gut microbiome in human immunodeficiency virus infection. BMC Med. 2016;14:83.
Vazquez-Castellano JF, Serrano-Villar S, Jimenez-Hernandez N, Soto del Rio MD, Gayo S, Rojo D, et al. Interplay between gut microbiota metabolism and inflammation in HIV infection. ISME J 2018;12:1964–76.
Fadlallah J, El Kafsi H, Sterlin D, Juste C, Parizot C, Dorgham K, et al. Microbial ecology perturbation in human IgA deficiency. Sci Transl Med 2018. https://doi.org/10.1126/scitranslmed.aan1217.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
No external funding was used in the preparation of this manuscript.
Conflict of interest
Melissa Bersanelli received honoraria for advisory roles and as speaker at scientific events from Bristol-Myers Squibb (BMS), Novartis, and Pfizer. Andrea Ticinesi received a congress grant from Novo Nordisk. Sebastiano Buti received honoraria for advisory roles and as speaker at scientific events from Pfizer, BMS, IPSEN, Pierre-Fabre, Merck Sharp & Dohme (MSD), and AstraZeneca. Matteo Santoni declares that he has no conflicts of interest that might be relevant to the contents of this manuscript.
Rights and permissions
About this article
Cite this article
Bersanelli, M., Santoni, M., Ticinesi, A. et al. The Urinary Microbiome and Anticancer Immunotherapy: The Potentially Hidden Role of Unculturable Microbes. Targ Oncol 14, 247–252 (2019). https://doi.org/10.1007/s11523-019-00643-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-019-00643-7