Abstract
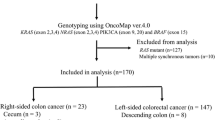
Recent data showed that metastatic colorectal (mCRC) tumors exhibiting extended RAS-BRAF mutations were resistant to anti-epidermal growth factor receptor (EGFR) monoclonal antibodies, making these drugs suitable for the so-called “super” wild-type (WT) patients only. This study aimed to compare the extended RAS-BRAF mutation frequency and characteristics according to location of tumor sampling. All consecutive mCRC specimens (N = 1659) referred to our institution from January 2008 till June 2014 were included in the analysis. Tumor genotyping (first for KRAS exon 2, then for BRAF exon 15, and later for KRAS exons 2, 3, and 4 and NRAS exons 2, 3, and 4) was performed with high-resolution melting analysis or allelic discrimination. The factors predicting for the presence of mutation were explored using multivariate binary logistic regression. Overall, the prevalence of KRAS exon 2 was 36.8 %, and it was lower in liver metastases (N = 138/490; 28.2 %) in comparison with primary tumors (N = 442/1086; 40.7 %), lung metastases (16/32; 50 %), or other metastatic sites (15/51; 29.4 %; P < 0.0001). Similarly, in the 1428 samples analyzed, BRAF mutations were less often found in liver metastases (N = 9/396; 2.3 %) as compared to primary tumors (N = 79/959; 8.2 %), lung metastases (N = 2/29; 6.9 %), or other metastatic locations (N = 2/44; 4.5 %; P < 0.0002). Overall occurrence of extended RAS mutation was 51.7 %. Of the 503 samples tested, the prevalence of extended RAS-BRAF mutations was twice as low in liver metastases (N = 53/151; 34.2 %) as compared to primary tumors (N = 191/322; 59.3 %, P < 0.0001). Univariate analysis identified age ≤65 years, male gender, and liver localization as predictors of super WT status. At multivariate analysis, only liver metastases were retained (RR 2.85 [95 % CI 1.91–4.30]). Colorectal liver metastases are twice as likely to exhibit a super WT genotype as compared to other tumor locations independently from other factors. This molecular feature has the potential to influence therapeutic strategy in mCRC patients.
Similar content being viewed by others
References
Cunningham D, Humblet Y, Siena S et al (2004) Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N Engl J Med 351:337–345. doi:10.1056/NEJMoa033025
Douillard J-Y, Siena S, Cassidy J et al (2010) Randomized, phase III trial of panitumumab with infusional fluorouracil, leucovorin, and oxaliplatin (FOLFOX4) versus FOLFOX4 alone as first-line treatment in patients with previously untreated metastatic colorectal cancer: the PRIME study. J Clin Oncol Off J Am Soc Clin Oncol 28:4697–4705. doi:10.1200/JCO.2009.27.4860
Van Cutsem E, Peeters M, Siena S et al (2007) Open-label phase III trial of panitumumab plus best supportive care compared with best supportive care alone in patients with chemotherapy-refractory metastatic colorectal cancer. J Clin Oncol Off J Am Soc Clin Oncol 25:1658–1664. doi:10.1200/JCO.2006.08.1620
Douillard J-Y, Oliner KS, Siena S et al (2013) Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 369:1023–1034. doi:10.1056/NEJMoa1305275
Lièvre A, Bachet J-B, Le Corre D et al (2006) KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res 66:3992–3995. doi:10.1158/0008-5472.CAN-06-0191
Peeters M, Price TJ, Cervantes A et al (2010) Randomized phase III study of panitumumab with fluorouracil, leucovorin, and irinotecan (FOLFIRI) compared with FOLFIRI alone as second-line treatment in patients with metastatic colorectal cancer. J Clin Oncol Off J Am Soc Clin Oncol 28:4706–4713. doi:10.1200/JCO.2009.27.6055
Mao C, Liao R-Y, Qiu L-X et al (2011) BRAF V600E mutation and resistance to anti-EGFR monoclonal antibodies in patients with metastatic colorectal cancer: a meta-analysis. Mol Biol Rep 38:2219–2223. doi:10.1007/s11033-010-0351-4
Douillard J-Y, Oliner KS, Siena S et al (2013) Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 369:1023–1034. doi:10.1056/NEJMoa1305275
De Roock W, Claes B, Bernasconi D et al (2010) Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol 11:753–762. doi:10.1016/S1470-2045(10)70130-3
Sorich MJ, Wiese MD, Rowland A, et al. (2014) Annals of oncology advance access published August 12, 2014. 1–27
Santini D, Loupakis F, Vincenzi B et al (2008) High concordance of KRAS status between primary colorectal tumors and related metastatic sites: implications for clinical practice. Oncologist 13:1270–1275. doi:10.1634/theoncologist. 2008-0181
Knijn N, Mekenkamp LJM, Klomp M et al (2011) KRAS mutation analysis: a comparison between primary tumours and matched liver metastases in 305 colorectal cancer patients. Br J Cancer 104:1020–1026. doi:10.1038/bjc.2011.26
Han C-B, Li F, Ma J-T, Zou H-W (2012) Concordant KRAS mutations in primary and metastatic colorectal cancer tissue specimens: a meta-analysis and systematic review. Cancer Investig 30:741–747. doi:10.3109/07357907.2012.732159
Tie J, Lipton L, Desai J et al (2011) KRAS mutation is associated with lung metastasis in patients with curatively resected colorectal cancer. Clin Cancer Res Off J Am Assoc Cancer Res 17:1122–1130. doi:10.1158/1078-0432.CCR-10-1720
Institut National du Cancer (2010) Bonnes pratiques pour la recherche à visée théranostique demutations somatiques dans les tumeurs solides. wwwe-cancerfr
Mancini I, Pinzani P, Pupilli C et al (2012) A high-resolution melting protocol for rapid and accurate differential diagnosis of thyroid nodules. J Molec Diagnos JMD 14:501–509. doi:10.1016/j.jmoldx.2012.03.003
Simi L, Pratesi N, Vignoli M et al (2008) High-resolution melting analysis for rapid detection of KRAS, BRAF, and PIK3CA gene mutations in colorectal cancer. Am J Clin Pathol 130:247–253. doi:10.1309/LWDY1AXHXUULNVHQ
Willmore-Payne C, Holden JA, Hirschowitz S, Layfield LJ (2006) BRAF and c-kit gene copy number in mutation-positive malignant melanoma. Hum Pathol 37:520–527. doi:10.1016/j.humpath.2006.01.003
Brink M, de Goeij AFPM, Weijenberg MP et al (2003) K-ras oncogene mutations in sporadic colorectal cancer in The Netherlands cohort study. Carcinogenesis 24:703–710
Samowitz WS, Curtin K, Schaffer D, et al. (2000) Relationship of Ki-ras mutations in colon cancers to tumor location, stage, and survival : a population-based study relationship of Ki-ras mutations in colon cancers to tumor location,. 1193–1197
Cejas P, López-Gómez M, Aguayo C et al (2009) KRAS mutations in primary colorectal cancer tumors and related metastases: a potential role in prediction of lung metastasis. PLoS One 4:e8199. doi:10.1371/journal.pone.0008199
Park JH, Han S-W, Oh D-Y et al (2011) Analysis of KRAS, BRAF, PTEN, IGF1R, EGFR intron 1 CA status in both primary tumors and paired metastases in determining benefit from cetuximab therapy in colon cancer. Cancer Chemother Pharmacol 68:1045–1055. doi:10.1007/s00280-011-1586-z
Losi L, Benhattar J, Costa J (1992) Stability of K-ras mutations throughout the natural history of human colorectal cancer. Euro J Cancer (Oxford, England: 1990) 28A:1115–1120
Weber J-C, Meyer N, Pencreach E et al (2007) Allelotyping analyses of synchronous primary and metastasis CIN colon cancers identified different subtypes. Int J Cancer J Int du Cancer 120:524–532. doi:10.1002/ijc.22343
Vauthey J-N, Zimmitti G, Kopetz SE et al (2013) RAS mutation status predicts survival and patterns of recurrence in patients undergoing hepatectomy for colorectal liver metastases. Ann Surg 258:619–626. doi:10.1097/SLA.0b013e3182a5025a, discussion 626–7
Gavin PG, Colangelo LH, Fumagalli D et al (2012) Mutation profiling and microsatellite instability in stage II and III colon cancer: an assessment of their prognostic and oxaliplatin predictive value. Clin Cancer Res Off J Am Assoc Cancer Res 18:6531–6541. doi:10.1158/1078-0432.CCR-12-0605
Venderbosch S, Nagtegaal ID, Maughan TS et al (2014) Mismatch repair status and BRAF mutation status in metastatic colorectal cancer patients: a pooled analysis of the CAIRO, CAIRO2, COIN and FOCUS studies. Clin Cancer Res. doi:10.1158/1078-0432.CCR-14-0332
Misale S, Yaeger R, Hobor S et al (2012) Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486:532–536. doi:10.1038/nature11156
Losi L, Baisse B, Bouzourene H, Benhattar J (2005) Evolution of intratumoral genetic heterogeneity during colorectal cancer progression. Carcinogenesis 26:916–922. doi:10.1093/carcin/bgi044
Giaretti W, Monaco R, Pujic N et al (1996) Intratumor heterogeneity of K-ras2 mutations in colorectal adenocarcinomas: association with degree of DNA aneuploidy. Am J Pathol 149:237–245
Baldus SE, Schaefer K-L, Engers R et al (2010) Prevalence and heterogeneity of KRAS, BRAF, and PIK3CA mutations in primary colorectal adenocarcinomas and their corresponding metastases. Clin Cancer Res Off J Am Assoc Cancer Res 16:790–799. doi:10.1158/1078-0432.CCR-09-2446
Poultsides G a, Bao F, Servais EL (2012) Pathologic response to preoperative chemotherapy in colorectal liver metastases: fibrosis, not necrosis, predicts outcome. Ann Surg Oncol 19:2797–2804. doi:10.1245/s10434-012-2335-1
Talmadge JE, Fidler IJ (2010) AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res 70:5649–5669. doi:10.1158/0008-5472.CAN-10-1040
Bernards R, Weinberg R a (2002) A progression puzzle. Nature 418:823. doi:10.1038/418823a
Fidler IJ, Kripke ML (1977) Metastasis results from preexisting variant cells within a malignant tumor. Science (New York, NY) 197:893–895
Wu X, Northcott P a, Dubuc A (2012) Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482:529–533. doi:10.1038/nature10825
Langley RR, Fidler IJ (2007) Tumor cell-organ microenvironment interactions in the pathogenesis of cancer metastasis. Endocr Rev 28:297–321. doi:10.1210/er.2006-0027
Kim M-J, Lee HS, Kim JH et al (2012) Different metastatic pattern according to the KRAS mutational status and site-specific discordance of KRAS status in patients with colorectal cancer. BMC Cancer 12:347. doi:10.1186/1471-2407-12-347
Lièvre A, Bachet J-B, Boige V et al (2008) KRAS mutations as an independent prognostic factor in patients with advanced colorectal cancer treated with cetuximab. J Clin Oncol Off J Am Soc Clin Oncol 26:374–379. doi:10.1200/JCO.2007.12.5906
Di Fiore F, Blanchard F, Charbonnier F et al (2007) Clinical relevance of KRAS mutation detection in metastatic colorectal cancer treated by Cetuximab plus chemotherapy. Br J Cancer 96:1166–1169. doi:10.1038/sj.bjc.6603685
Khambata-Ford S, Garrett CR, Meropol NJ et al (2007) Expression of epiregulin and amphiregulin and K-ras mutation status predict disease control in metastatic colorectal cancer patients treated with cetuximab. J Clin Oncol Off J Am Soc Clin Oncol 25:3230–3237. doi:10.1200/JCO.2006.10.5437
Heinemann V, von Weikersthal LF, Decker T et al (2014) FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): a randomised, open-label, phase 3 trial. Lancet Oncol 2045:1–11. doi:10.1016/S1470-2045(14)70330-4
Sclafani F, Cunningham D (2014) Cetuximab or bevacizumab in metastatic colorectal cancer? Lancet Oncol 2045:2–3. doi:10.1016/S1470-2045(14)70360-2
Conflict of interest
The authors have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table
Factors associated with BRAF mutation: univariate and multivariate analysis (DOCX 109 kb)
Rights and permissions
About this article
Cite this article
Allard, M.A., Saffroy, R., de la Maisonneuve, P.B. et al. Colorectal liver metastases are more often super wild type. Toward treatment based on metastatic site genotyping?. Targ Oncol 10, 415–421 (2015). https://doi.org/10.1007/s11523-014-0346-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-014-0346-5